Abstract
Background
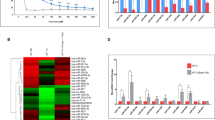
Drug resistance in colorectal cancers is assumed to be mediated by changes in the expression of microRNAs, but the specific identities and roles of microRNAs are largely unclear. We examined the effect of 5-fluorouracil (5-FU) resistance on microRNA expression.
Methods
Two types of 5-FU-resistant colon cancer cells were derived from the DLD-1 and KM12C cell lines. The expressions of microRNAs were profiled with a microarray containing 723 microRNAs and validated by quantitative real-time polymerase chain reaction (qRT-PCR). To survey the downstream mediators of microRNA, we used a microRNA:mRNA immunoprecipitation (RIP)-Chip and pathway analysis tool to identify potential direct targets of microRNA.
Results
In response to 5-FU, miR-19b and miR-21 were over-expressed in 5-FU-resistant cells. Of note, miR-19b was up-regulated 3.47-fold in the DLD-1 resistant cells, which exhibited no alteration in cell cycle profiles despite exposure to 5-FU. After transfection of miR-19b, specific mRNAs were recruited to microRNA:mRNA complexes isolated with Ago2 antibody and subjected to whole-genome transcriptional analysis. In this analysis, 66 target mRNAs were enriched by at least 5.0-fold in the microRNA:mRNA complexes from DLD-1 resistant cells. Ingenuity pathway analysis of mRNA targets significantly (P < 0.05) indicated the category “Cell Cycle” as a probable area of the molecular and cellular function related with 5-FU resistance. Among candidate mRNA targets, SFPQ and MYBL2 have been linked to cell cycle functions.
Conclusions
We revealed up-regulation of miR-19b in response to 5-FU and potential targets of miR-19b mediating the cell cycle under treatment with 5-FU. Our study provides an important insight into the mechanism of 5-FU resistance in colorectal cancers.




Similar content being viewed by others
References
Jemal A, Siegel R, Xu J, et al. Cancer statistics, 2010. CA Cancer J Clin. 2010;60:277–300.
Efficacy of adjuvant fluorouracil and folinic acid in colon cancer. International Multicentre Pooled Analysis of Colon Cancer Trials (IMPACT) investigators. Lancet. 1995; 345:939–44.
Douillard JY, Cunningham D, Roth AD, et al. Irinotecan combined with fluorouracil compared with fluorouracil alone as first-line treatment for metastatic colorectal cancer: a multicentre randomised trial. Lancet. 2000;355:1041–7.
Dean M, Fojo T, Bates S. Tumour stem cells and drug resistance. Nat Rev Cancer. 2005;5:275–84.
Iorns E, Lord CJ, Turner N, et al. Utilizing RNA interference to enhance cancer drug discovery. Nat Rev Drug Discov. 2007;6:556–68.
Mariadason JM, Arango D, Shi Q, et al. Gene expression profiling-based prediction of response of colon carcinoma cells to 5-fluorouracil and camptothecin. Cancer Res. 2003;63:8791–812.
Boyer J, Allen WL, McLean EG, et al. Pharmacogenomic identification of novel determinants of response to chemotherapy in colon cancer. Cancer Res. 2006;66:2765–77.
Ooyama A, Takechi T, Toda E, et al. Gene expression analysis using human cancer xenografts to identify novel predictive marker genes for the efficacy of 5-fluorouracil-based drugs. Cancer Sci. 2006;97:510–22.
Karasawa H, Miura K, Fujibuchi W, et al. Down-regulation of cIAP2 enhances 5-FU sensitivity through the apoptotic pathway in human colon cancer cells. Cancer Sci. 2009;100:903–13.
Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 2004;116:281–97.
He L, Hannon GJ. MicroRNAs: small RNAs with a big role in gene regulation. Nat Rev Genet. 2004;5:522–31.
Nilsen TW. Mechanisms of microRNA-mediated gene regulation in animal cells. Trends Genet. 2007;23:243–9.
Pillai RS, Bhattacharyya SN, Filipowicz W. Repression of protein synthesis by miRNAs: how many mechanisms? Trends Cell Biol. 2007;17:118–26.
Miranda KC, Huynh T, Tay Y, et al. A pattern-based method for the identification of MicroRNA binding sites and their corresponding heteroduplexes. Cell. 2006;126:1203–17.
Kloosterman WP, Plasterk RH. The diverse functions of microRNAs in animal development and disease. Dev Cell. 2006;11:441–50.
Rosenfeld N, Aharonov R, Meiri E, et al. MicroRNAs accurately identify cancer tissue origin. Nat Biotechnol. 2008;26:462–9.
Calin GA, Croce CM. MicroRNA signatures in human cancers. Nat Rev Cancer. 2006;6:857–66.
Si ML, Zhu S, Wu H, et al. miR-21-mediated tumor growth. Oncogene. 2007;26:2799–803.
Yang H, Kong W, He L, et al. MicroRNA expression profiling in human ovarian cancer: miR-214 induces cell survival and cisplatin resistance by targeting PTEN. Cancer Res. 2008;68:425–33.
Murakami Y, Kazuno H, Emura T, et al. Different mechanisms of acquired resistance to fluorinated pyrimidines in human colorectal cancer cells. Int J Oncol. 2000;17:277–83.
Fukushima M, Fujioka A, Uchida J, et al. Thymidylate synthase (TS) and ribonucleotide reductase (RNR) may be involved in acquired resistance to 5-fluorouracil (5-FU) in human cancer xenografts in vivo. Eur J Cancer. 2001;37:1681–7.
Boyer J, McLean EG, Aroori S, et al. Characterization of p53 wild-type and null isogenic colorectal cancer cell lines resistant to 5-fluorouracil, oxaliplatin, and irinotecan. Clin Cancer Res. 2004;10:2158–67.
Schroeder A, Mueller O, Stocker S, et al. The RIN: an RNA integrity number for assigning integrity values to RNA measurements. BMC Mol Biol. 2006;7:3.
Allison DB, Cui X, Page GP, et al. Microarray data analysis: from disarray to consolidation and consensus. Nat Rev Genet. 2006;7:55–65.
Zhang X, Chen J, Radcliffe T, et al. An array-based analysis of microRNA expression comparing matched frozen and formalin-fixed paraffin-embedded human tissue samples. J Mol Diagn. 2008;10:513–9.
Wang WX, Wilfred BR, Hu Y, et al. Anti-Argonaute RIP-Chip shows that miRNA transfections alter global patterns of mRNA recruitment to microribonucleoprotein complexes. RNA. 2010;16:394–404.
Calvano SE, Xiao W, Richards DR, et al. A network-based analysis of systemic inflammation in humans. Nature. 2005;437:1032–7.
Mu P, Han YC, Betel D, et al. Genetic dissection of the miR-17 ~ 92 cluster of microRNAs in Myc-induced B-cell lymphomas. Genes Dev. 2009;23:2806–11.
Olive V, Bennett MJ, Walker JC, et al. miR-19 is a key oncogenic component of mir-17-92. Genes Dev. 2009;23:2839–49.
Giovannetti E, Funel N, Peters GJ, et al. MicroRNA-21 in pancreatic cancer: correlation with clinical outcome and pharmacologic aspects underlying its role in the modulation of gemcitabine activity. Cancer Res. 2010;70:4528–38.
Wang H, Ach RA, Curry B. Direct and sensitive miRNA profiling from low-input total RNA. RNA. 2007;13:151–9.
Tan LP, Seinen E, Duns G, et al. A high throughput experimental approach to identify miRNA targets in human cells. Nucleic Acids Res. 2009;37:e137.
Hayashida Y, Nishibu T, Inoue K, et al. A useful approach to total analysis of RISC-associated RNA. BMC Res Notes. 2009;2:169.
Schetter AJ, Leung SY, Sohn JJ, et al. MicroRNA expression profiles associated with prognosis and therapeutic outcome in colon adenocarcinoma. JAMA. 2008;299:425–36.
Schepeler T, Reinert JT, Ostenfeld MS, et al. Diagnostic and prognostic microRNAs in stage II colon cancer. Cancer Res. 2008;68:6416–24.
Lanza G, Ferracin M, Gafa R, et al. mRNA/microRNA gene expression profile in microsatellite unstable colorectal cancer. Mol Cancer. 2007;6:54.
Murakami Y, Yasuda T, Saigo K, et al. Comprehensive analysis of microRNA expression patterns in hepatocellular carcinoma and non-tumorous tissues. Oncogene. 2006;25:2537–45.
Blower PE, Chung JH, Verducci JS, et al. MicroRNAs modulate the chemosensitivity of tumor cells. Mol Cancer Ther. 2008;7:1–9.
Meng F, Henson R, Lang M, et al. Involvement of human micro-RNA in growth and response to chemotherapy in human cholangiocarcinoma cell lines. Gastroenterology. 2006;130:2113–29.
Kovalchuk O, Filkowski J, Meservy J, et al. Involvement of microRNA-451 in resistance of the MCF-7 breast cancer cells to chemotherapeutic drug doxorubicin. Mol Cancer Ther. 2008;7:2152–9.
Xia L, Zhang D, Du R, et al. miR-15b and miR-16 modulate multidrug resistance by targeting BCL2 in human gastric cancer cells. Int J Cancer. 2008;123:372–9.
Rossi L, Bonmassar E, Faraoni I. Modification of miR gene expression pattern in human colon cancer cells following exposure to 5-fluorouracil in vitro. Pharmacol Res. 2007;56:248–53.
O’Donnell KA, Wentzel EA, Zeller KI, et al. c-Myc-regulated microRNAs modulate E2F1 expression. Nature. 2005;435:839–43.
Mendell JT. miRiad roles for the miR-17-92 cluster in development and disease. Cell. 2008;133:217–22.
Gozani O, Patton JG, Reed R. A novel set of spliceosome-associated proteins and the essential splicing factor PSF bind stably to pre-mRNA prior to catalytic step II of the splicing reaction. EMBO J. 1994;13:3356–67.
Rajesh C, Baker DK, Pierce AJ, et al. The splicing-factor related protein SFPQ/PSF interacts with RAD51D and is necessary for homology-directed repair and sister chromatid cohesion. Nucleic Acids Res. 2011;39:132–45.
Salton M, Lerenthal Y, Wang SY, et al. Involvement of Matrin 3 and SFPQ/NONO in the DNA damage response. Cell Cycle. 2010;9:1568–76.
Oh IH, Reddy EP. The myb gene family in cell growth, differentiation and apoptosis. Oncogene. 1999;18:3017–33.
Joaquin M, Watson RJ. Cell cycle regulation by the B-Myb transcription factor. Cell Mol Life Sci. 2003;60:2389–401.
Acknowledgments
This study was supported by Grants for project research (Development of fundamental technology for analysis and evaluation of functional agricultural products and functional foods). This study was also supported by Grants-in-Aid (#23590943) from the Ministry of Education, Culture, Sports, Science and Technology, Japan (T.T.).
Conflict of interest
There is no financial interest with regard to the submitted manuscript that might be construed as a conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kurokawa, K., Tanahashi, T., Iima, T. et al. Role of miR-19b and its target mRNAs in 5-fluorouracil resistance in colon cancer cells. J Gastroenterol 47, 883–895 (2012). https://doi.org/10.1007/s00535-012-0547-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00535-012-0547-6