Abstract
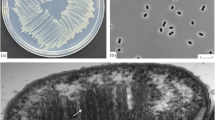
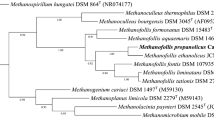
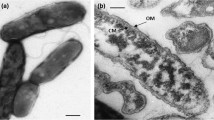
A new genus of methanogenic bacteria is described, which was isolated from a mesophilic sewage digester. It is most probably the filamentous bacterium, earlier referred to asMethanobacterium soehngenii, “fat rod” or “acetate organism”. The single non-motile, non-sporeforming cells are rod-shaped (0.8×2 μm) and are normally combined end to end in long filaments, surrounded by a sheath-like structure. The filaments form characteristic bundles.Methanothrix soehngenii decarboxylates acetate, yielding methane and carbon dioxide. Other methanogenic substrates (H2−CO2, formate, methanol, methylamines) are not used for growth or methane formation. Formate is split into hydrogen and carbon dioxide. The temperature optimum for growth and methane formation is 37°C and the optimal pH range is 7.4–7.8. Sulfide and ammonia serve as sulfur and nitrogen source respectively. Oxygen completely inhibits growth and methane formation, but the bacteria do not loose their viability when exposed to high oxygen concentrations. 100 mg/l vancomycin showed no inhibition of growth and methanogenesis. No growth and methane formation was observed in the presence of: 2-bromoethanesulfonic acid, viologen dyes, chloroform, and KCN. The bacterium has a growth yield on acetate of 1.1–1.4 g biomass per mol acetate. The apparent “K S ” of the acetate conversion system to methane and carbon dioxide is 0.7 mmol/l. The DNA base composition is 51.9 mol% guanine plus cytosine. The nameMethanothrix is proposed for this new genus of filamentous methane bacterium. The type species,Methanothrix soehngenii sp. nov., is named in honor of N. L. Söhngen.
Similar content being viewed by others
References
Balch WE, Wolfe RS (1976) New approach to the cultivation of methanogenic bacteria: 2-mercaptoethanesulfonic acid (HS-CoM)-dependent growth ofMethanobacterium ruminantium in a pressurized atmosphere. Appl Environ Microbiol 32:781–791
Balch WE, Wolfe RS (1979) Specifity and biological distribution of coenzyme M (2-mercaptoethanesulfonic acid) J Bacteriol 137:256–263
Balch WE, Fox GE, Magrum LJ, Woese CR, Wolfe RS (1979) Methanogens: Reevaluation of a unique biological group. Microbiol Rev 43:260–296
Baresi L, Wolfe RS (1981) Levels of coenzyme F420, coenzyme M, hydrogenase, and methylcoenzyme M methylreductase in acetategrownMethanosarcina. Appl Environ Microbiol 41:388–391
Barker HA (1936) Studies upon the methane-producing bacteria. Arch Mikrobiol 7:420–438
Belyaev SS, Finkel'shtein ZI, Ivanov MV (1975) Intensity of bacterial methane formation in ooze deposits of certain lakes. Microbiology 44:272–275
Boone DR, Bryant MP (1980) Propionate-degrading bacterium,Synthrophobacter wolinii, sp. nov. gen. nov., from methanogenic ecosystems. Appl Environ Microbiol 40:626–632
Bryant MP (1972) Commentary on the Hungate technique, for culture of anaerobic bacteria. Am J Clin Nutr 25:1324–1328
Buswell AM, Hatfield WD (1936) Anaerobic fermentations. Ill State Wat Survey Bull 32:59–65
Cappenberg TE (1976) Methanogenesis in the bottom deposits of a small stratified lake, in HG Schlegel, G Gottschalk, N Pfennig (eds) Microbial formation and utilization of gases (H2, CH4, CO). Verlag Goltze KG. Göttingen, pp 125–134
Cappenberg TE, Prins H (1974) Interrelations between sulfate-reducing and methane-producing bacteria in bottom deposits of a fresh water lake. III. Experiments with14C-labelled substrates. Antonie van Leeuwenhoek J Microbiol Serol 40:457–469
Colvin JR, Sowden LC, van den Berg L (1979) The ultrastructure of the major species of an enriched methanogenic culture utilizing acetic acid. Can J Microbiol 25:826–832
Ferry JG, Wolfe RS (1976) Anaerobic degradation of benzoate to methane by a microbial consortium. Arch Microbiol 107:33–40
Godsy EM (1980) Isolation ofMethanobacterium bryantii from a deep aquifier by using a novel broth-antibiotic disc method. Appl Environ Microbiol 39:1074–1075
Groenewege J (1920) Bakteriologische Untersuchungen über biologische Reinigung. Med Burg Geneesk Dienst 1:67–125
Gunsalus RP (1977) The methyl-coenzyme M reductase system inMethanobacterium thermoautotrophicum. Thesis, University of Illinois, Urbana-Champaign
Gusalus RP, Wolfe RS (1980) Methyl coenzyme M reductase fromMethanobacterium thermoautotrophicum. J Biol Chem 255:1891–1895
Hammes WP, Winter J, Kandler O (1979) The sensitivity of the pseudomurein-containing genusMethanobacterium to inhibitors of murein synthesis. Arch Microbiol 123:275–279
Healy JB, Young LY, Reinhard M (1980) Methanogenic decomposition of ferulic acid, a model lignin derivative. Appl Environ Microbiol 39:436–444
Hilpert R, Winter J, Hammes WP, Kandler O (1981) Sensitivity of archaebacteria to autibiotics. Zbl Bakt 1. Abt Orig C 2:11–21
Holdemann LV, Moore WEC (ed) (1972) Anaerobe laboratory manual. Virginia Polytechnic Institute and State University, Blacksburg
Hoppe-Seyler F (1876) Über die Processe der Gährungen und ihre Beziehung zum Leben der Organismen. Pflügers Arch ges Physiol 12:1–17
Hungate RE (1950) The anaerobic mesophilic cellulolytic bacteria. Bacteriol Rev 14:1–49
Huser BA (1981) Methanbildung aus Acetat: Isolierung eines neuen Archaebakteriums. Thesis Nr. 6750, Swiss Federal Institute of Technology, Zürich
Hutten TJ, de Jong MH, Peeters BPH, van der Drift Ch, Vogels GD (1981) Coenzyme M derivatives and their effects on methane formation from carbon dioxide and methanol by cell extracts ofMethanosarcina barkeri. J Bacteriol 145:27–34
Jeris JS, McCarty PL (1965) The biochemistry of methane fermentation using C14 tracers. J Wat Poll Contr Fed 37:178–192
Kaspar HF, Wuhrmann K (1978) Kinetic parameters and relative turnovers of some important catabolic reactions in digesting sludge. Appl Environ Microbiol 36:1–7
Khan AW, Mes-Hartree M (1981) Metabolism of acetate and hydrogen by a mixed population of anaerobes capable of converting cellulose to methane. J Appl Bacteriol 50:283–288
Lawrence AW, McCarty PL (1969) Kinetics of methane fermentation in anaerobic treatment. J Wat Poll Cont Fed 41:1–17
Mackie RI, Bryant MP (1981) Metabolic activity of fatty acid-oxidizing bacteria and the contribution of acetate, propionate, butyrate, and CO2 to methanogenesis in cattle waste at 40 and 60°C. Appl Environ Microbiol 41:1363–1373
Mah RA, Hungate RE, Ohwaki K (1976) Acetate, a key intermediate in methanogenesis, in: HG Schlegel, J Barnea (eds) Microbial, energy conversion. E. Goltze KG, Göttingen, pp 97–106
Mah RA, Smith MR, Baresi L (1978) Studies on an acetate-fermenting strain ofMethanosarcina. Appl. Environ Microbiol 35:1174–1184
McBride BC, Wolfe RS (1971) A new coenzyme of methyl-transfer, coenzyme M. Biochem 10:2317–2324
McInerney MJ, Bryant MP, Pfennig N (1979) Anaerobic bacterium that degrades fatty acids in syntrophic, association with methanogens. Arch Microbiol 122:129–135
McInerney MJ, Mackie RI, Bryant MP (1981) Syntrophic association of a butyrate-degrading bacterium andMethanosarcina enriched from bovine rumen fluid. Appl Environ Microbiol 41:826–828
Mountfort DO, Asher RA (1978) Changes in proportions of acetate and carbon dioxide used as methane precursors during the anaerobic digestion of bovine waste. Appl Environ Microbiol 35:648–654
Mylroie RL, Hungate RE (1954) Experiments on the methane bacteria in sludge. Can J Microbiol 1:55–64
Pretorius WA (1972) The effect of formate on the growth of acetate utilizing methanogenic bacteria. Wat Res 6:1213–1217
Roberton AM, Wolfe RS (1970) Adenosine triphosphate pools inMethanobacterium. J Bacteriol 102:43–51
Schnellen CGTP (1947) Onderzoekingen over de Methaangisting. Proefschrift, Technische Hoogeschool, Delft
Smit J (1930) Die Gärungssarcinen. Eine Monographie. Pflanzenforschung 14:1–59
Smith PH (1966) The microbial ecology of sludge methanogenesis. Devel Ind Microbiol 7:156–161
Smith PH, Mah RA (1966) Kinetics of acetate metabolism during sludge digestion. Appl Microbiol 14:368–371
Smith MR, Mah RA (1978) Growth and methanogenesis byMethanosarcina strain 227 on acetate and methanol. Appl Environ Microbiol 36:870–879
Smith MR, Zinder SH, Mah RA (1980) Microbial methanogenesis from acetate. Process Biochem 15:34–39
Söhngen NL (1906) Het ontstaan en verdwijnen van waterstof en methaan onder den invloed van het organische leven. Proefschrift. Technische Hoogeschool, Delft
Stackebrandt E, Kandler O (1979) Taxonomy of the genusCellulomonas based on phenotypic characters and deoxyribonucleic acid-deoxyribonucleic acid homology, and proposal of seven neotype strains. Int J Syst Bacteriol 29:273–282
Stackebrandt E, Seewaldt E, Ludwig W, Schleifer K-H, Huser BA (1982) Classification ofMethanothrix soehngenii, a new methanogenic archaebacterium. Zbl Bakt Hyg 1. Abt Orig C, in press
Takai Y (1970) The mechanism of methane fermentation in flooded paddy soil. Soil Sci Plant Nutr 16:238–244
Takai Y, Koyama T, Kamura T (1963) Microbial metabolism in reduction process of paddy soils. Soil Sci and Plant Nutr 9:1–5
Takamiya A (1942) Über die Formicodehydrase vonEscherichia coli. Acta Phytochim 13:1–9
Toerien DF, Thiel PG, Pretorius WA (1971) Substrate flow in anaerobic digestion. Adv Wat Poll Res 11:II 29/1–29/7
van den Berg L, Patel GB, Clark DS, Lentz CP (1976) Factors affecting rate of methane formation from acetic acid by enriched methanogenic cultures. Can J Microbiol 22:1312–1319
Weimer PJ, Zeikus JG (1978) Acetate metabolism inMethanosarcina barkeri. Arch Microbiol 119:175–182
Winfrey MR, Zeikus JG (1979) Anaerobic metabolism of immediate methane precursors in Lake Mendota. Appl Environ Microbiol 37:244–253
Woese CR (1981) Archaebacteria. Scient Am 244:94–106
Wolfe RS (1971) Microbial formation of methane, in: AH Rose, JF Wilkinson (eds) Advances in microbiological physiology. Academic Press Inc., New York, vol 6, pp 107–146
Wolin EA, Wolin MJ, Wolfe RS (1963) Formation of methane by bacterial extracts J Biol Chem 238:2882–2886
Zehnder AJB, Brock TD (1979) Methane formation and methane oxidation by methanogenic bacteria. J Bacteriol 137:420–432
Zehnder AJB, Huser B, Brock TD (1979) Measuring radioactive methane with the liquid scintillation counter. Appl Environ Microbiol 37:897–899
Zehnder AJB, Huser BA, Brock, TD, Wuhrmann K (1980) Characterization of an acetate-decarboxylating, non-hydrogenoxidizing methane bacterium. Arch Microbiol 124:1–11
Zeikus JG, Weimer PJ, Nelson DR, Daniels L (1975) Bacterial methanogenesis: Acetate as a methane precursor in pure culture. Arch Microbiol 104:129–134
Zinder SH, Mah RA (1979) Isolation and characterization of a thermophilic strain ofMethanosarcina unable to use H2−CO2 for methanogenesis. Appl Environ Microbiol 38:996–1008
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Huser, B.A., Wuhrmann, K. & Zehnder, A.J.B. Methanothrix soehngenii gen. nov. sp. nov., a new acetotrophic non-hydrogen-oxidizing methane bacterium. Arch. Microbiol. 132, 1–9 (1982). https://doi.org/10.1007/BF00690808
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00690808