Abstract
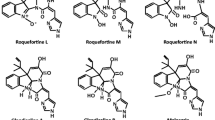
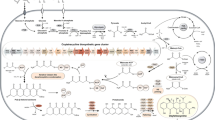
The production of mycotoxins and other secondary metabolites in Penicillium roqueforti is of great interest because of its long history of use in blue-veined cheese manufacture. In this article, we report the cloning and characterization of the roquefortine gene cluster in three different P. roqueforti strains isolated from blue cheese in the USA (the type strain), France, and the UK (Cheshire cheese). All three strains showed an identical roquefortine gene cluster organization and almost identical (98–99 %) gene nucleotide sequences in the entire 16.6-kb cluster region. When compared with the Penicillium chrysogenum roquefortine/meleagrin seven-gene cluster, the P. roqueforti roquefortine cluster contains only four genes (rds, rdh, rpt, and gmt) encoding the roquefortine dipeptide synthetase, roquefortine D dehydrogenase, roquefortine prenyltransferase, and a methyltransferase, respectively. Silencing of the rds or rpt genes by the RNAi strategy reduced roquefortine C production by 50 % confirming the involvement of these two key genes in roquefortine biosynthesis. An additional putative gene, orthologous of the MFS transporter roqT, is rearranged in all three strains as a pseudogene. The same four genes and a complete (not rearranged) roqT, encoding a MFS transporter containing 12 TMS domains, occur in the seven-gene cluster in P. chrysogenum although organized differently. Interestingly, the two “late” genes of the P. chrysogenum roquefortine/meleagrin gene cluster that convert roquefortine C to glandicoline B and meleagrin are absent in the P. roqueforti four-gene cluster. No meleagrin production was detected in P. roqueforti cultures grown in YES medium, while P. chrysogenum produces meleagrin in these conditions. No orthologous genes of the two missing meleagrin synthesizing genes were found elsewhere in the recently released P. roqueforti genome. Our data suggest that during evolution, the seven-gene cluster present in P. chrysogenum, and probably also in other glandicoline/meleagrin producing fungi, has been trimmed down to a short cluster in P. roqueforti leading to the synthesis of roquefortine C rather than meleagrin as a final product.






Similar content being viewed by others
References
Ali H, Ries MI, Nijland JG, Lankhorst PP, Hankemeier T, Bovenberg RA, Vreeken RJ, Driessen AJ (2013) A branched biosynthetic pathway is involved in production of roquefortine and related compounds in Penicillium chrysogenum. PLoS One 8(6):e65328
Aninat C, Hayashi Y, Andre F, Delaforge M (2001) Molecular requirements for inhibition of cytochrome p450 activities by roquefortine. Chem Res Toxicol 14:1259–1265
Casqueiro J, Bañuelos O, Gutiérrez S, Hijarrubia MJ, Martín JF (1999) Intrachromosomal recombination between direct repeats in Penicillium chrysogenum: gene conversión and deletion events. Mol Gen Genet 261:994–1000
Cepeda-García C, Domínguez-Santos R, García-Rico RO, García-Estrada C, Cajiao A, Fierro F, Martín JF (2014) Direct involvement of the CreA transcription factor in penicillin biosynthesis and expression of the pcbAB gene in Penicillium chrysogenum. Appl Microbiol Biotechnol 98:7113–7124
Chang SC, Lu CY, Li SY, Wei YH (1991a) Potentiation effect of corn extract on the production of eremofortin C, EC oxidase, and PR toxin by Penicillium roqueforti. Proc Natl Sci Counc Repub China B 15:153–159
Chang SC, Wei YH, Wei DL, Chen YY, Jong SC (1991b) Factors affecting the production of eremofortin C and PR toxin in Penicillium roqueforti. Appl Environ Microbiol 57:2581–2585
Cheeseman K, Ropars J, Renault P, Dupont J, Gouzy J, Branca A, Abraham AL, Ceppi M, Conseiller E, Debuchy R, Malagnac F, Goarin A, Silar P, Lacoste S, Sallet E, Bensimon A, Giraud T, Brygoo Y (2014) Multiple recent horizontal transfers of a large genomic region in cheese making fungi. Nat Commun 5:2876
Díez B, Barredo JL, Alvarez E, Cantoral JM, van Solingen P, Groenen MAM, Veenstra AE, Martín JF (1989) Two genes involved in penicillin biosynthesis are linked in a 5.1 kb SalI fragment in the genome of Penicillium chrysogenum. Mol Gen Genet 218:572–576
Domínguez-Santos R, Martín JF, Kosalková K, Prieto C, Ullán RV, García-Estrada C (2012) The regulatory factor PcRFX1 controls the expression of the three genes of β-lactam biosynthesis in Penicillium chrysogenum. Fungal Genet Biol 49:866–881
Fernández-Aguado M, Ullán RV, Teijeira F, Rodríguez-Castro R, Martín JF (2013) The transport of phenylacetic acid across the peroxisomal membrane is mediated by the PaaT protein in Penicillium chrysogenum. Appl Microbiol Biotechnol 97:3073–3084
Fernández-Bodega MA, Mauriz E, Gómez A, Martín JF (2009) Proteolytic activity, mycotoxins and andrastin A in Penicillium roqueforti strains isolated from Cabrales, Valdeón and Bejes-Tresviso local varieties of blue-veined cheeses. Intern J Food Microbiol 136:18–25
Fierro F, García-Estrada C, Castillo NI, Rodríguez R, Velasco-Conde T, Martín JF (2006) Transcriptional and bioinformatic analysis of the 56.8 kb DNA region amplified in tandem repeats containing the penicillin gene cluster in Penicillium chrysogenum. Fungal Genet Biol 43:618–629
Fontaine K, Passeró E, Vallone L, Hymery N, Coton M, Jany JL, Mounier J, Coton E (2015) Occurrence of roquefortine C, mycohenolic acid and aflatoxin M1 mycotoxins in blue-veined cheeses. Food Control 47:634–640
Frisvad F, Smedsgaard J, Larsen T, Samson R (2004) Mycotoxins, drugs and other extrolites produced by species in Penicillium subgenus Penicillium. Stud Mycol 49:201–242
García-Estrada C, Ullán RV, Albillos SM, Fernández-Bodega MÁ, Durek P, von Döhren H, Martín JF (2011) A single cluster of coregulated genes encodes the biosynthesis of the mycotoxins roquefortine C and meleagrin in Penicillium chrysogenum. Chem Biol 18:1499–1512
García-Rico RO, Fierro F, Mauriz E, Gómez A, Fernández-Bodega MA, Martín JF (2008) The heterotrimeric Gα protein Pga1 regulates biosynthesis of penicillin, chrysogenin and roquefortine in Penicillium chrysogenum. Microbiology 154:3567–3578
Hidalgo PI, Ullán RV, Albillos SM, Montero O, Fernández-Bodega MÁ, García-Estrada C, Fernández-Aguado M, Martín JF (2014) Molecular characterization of the PR-toxin gene cluster in Penicillium roqueforti and Penicillium chrysogenum: cross talk of secondary metabolite pathways. Fungal Genet Biol 62:11–24
Houbraken J, Frisvad JC, Samson RA (2011) Fleming’s penicillin production strain is not Penicillium chrysogenum but P. rubens. Fungus 2:87–95
Kato N, Suzuki H, Takagi H, Asami Y, Kakeya H, Uramoto M, Usui T, Takahashi S, Sugimoto Y, Osada H (2009) Identification of cytochrome P450s required for fumitremorgin biosynthesis in Aspergillus fumigatus. Chembiochem 10(5):920–928
Kopp B, Rehm HJ (1981) Studies on the inhibition of bacterial macromolecule synthesis by roquefortine, a mycotoxin from penicillium roqueforti. Eur J Appl Microbiol Biotechnol 13:232–235
Kopp-Holtwiesche B, Rehm HJ (1990) Antimicrobial action of roquefortine. J Environ Pathol Toxicol Oncol 10:41–44
Kozlovsky AG, Vinokurova NG, Reshetilova TA, Sakharovsky VG, Baskunov BP, Seleznev SG (1994) New metabolites of Penicillium glandicola var. glandicola: glandicoline A and glandicoline B. Microbiology 30:334–337
Li SM (2009) Evolution of aromatic prenyltransferases in the biosynthesis of indole derivatives. Phytochemistry 70:1746–1757
Martín JF (2000) Molecular control of expression of penicillin biosynthesis genes in fungi: regulatory proteins interact with a bidirectional promoter region. J Bacteriol 182:2355–2362
Martín JF (2015) Fungal transformation: from protoplasts to targeted recombination systems. In: van den Berg M, Maruthachalam K (eds) Genetic transformation systems in fungi, Vol. 1, Fungal Biology. Springer Int Publ Switzerland. In press.
Martín JF, Coton M (2015) Blue cheese: microbiota and fungal metabolites. In: Frias J, Martínez-Villaluenga C, Peñas, E (eds) Fermented Foods in Health and Disease Prevention. Elsevier, New York. In press.
Martín JF, Casqueiro J, Liras P (2005) Secretion systems for secondary metabolites: how producer cells send out messages of intercellular communication. Curr Opin Microbiol 8:282–293
Martín JF, Liras P, García-Estrada C (2014) Roquefortine and Prenylated Indole Alkaloids. In: Martín JF, García-Estrada C, Zeilinger S (eds) Biosynthesis and Molecular Genetics of Fungal Secondary Metabolites. Springer, New York, pp 111–128
Nielsen KF, Sumarah MW, Frisvad JC, Niller JD (2006) Production of Metabolites from the Penicillium roqueforti complex. J Agric Food Chem 54:3756–3763
Ohmomo S, Oguma K, Ohashi T, Abe M (1978) Isolation of a new indole alkaloid, roquefortine D, from the cultures of Penicillium roqueforti. Agric Biol Chem 42:2387–2389
Overy DP, Nielsen KF, Smedsgaard J (2005) Roquefortine/oxaline biosynthesis pathway metabolites in Penicillium ser. Corymbifera: in planta production and implications for competitive fitness. J Chem Ecol 31:2373–2390
Polonsky J, Merrien MA, Scott PM (1977) Roquefortine and isofumigaclavine A, alkaloids from Penicillium roqueforti. Ann Nutr Aliment 31:963–968
Reshetilova TA, Vinokurova NG, Khmelenina VN, Kozlovsky AG (1995) The role of roquefortine in the synthesis of alkaloids meleagrin, glandicolines A and B and oxaline in fungi Penicillium glandicola and P. atramentosum. Mikrobiology 64:27–29
Ries MI, Ali H, Lankhorst PP, Hankemeier T, Bovenberg RA, Driessen AJ, Vreeken RJ (2013) Novel key metabolites reveal further branching of the roquefortine/meleagrin biosynthetic pathway. J Biol Chem 288:37289–37295
Tudzynski P, Neubauer L (2014) Ergot Alkaloids. In: Martín JF, García-Estrada C, Zeilinger S (eds) Biosynthesis and Molecular Genetics of Fungal Secondary Metabolites. Springer, New York, pp 303–316
Ullán RV, Godio RP, Teijeira F, Vaca I, García-Estrada C, Feltrer R, Kosalkova K, Martín JF (2008) RNA-silencing in Penicillium chrysogenum and Acremonium chrysogenum: validation studies using beta-lactam genes expression. J Microbiol Methods 75:209–218
Vaiman D (2002) Agar plug/serial dilution approach for rapid PCR screening of phage libraries. Biotechniques 33:764–766
Wagener RE, Davis ND (1980) Diener UL (1980) Penitrem A and Roquefortine Production by Penicillium commune. Appl Environ Microbiol 39:882–887
Ware GM, Thorpe CW, Pohland AE (1980) Determination of roquefortine in blue cheese and blue cheese dressing by high pressure liquid chromatography with ultraviolet and electrochemical detectors. J Assoc Off Anal Chem 63:637–641
Acknowledgments
This work was supported by a Grant of the European Union (EUROFUNGBASE LSSG-CT-2005-018964) to J.F.M. R. Domínguez-Santos received a fellowship (EDU1204/2010) from the Junta de Castilla y León. We thank D. Miller (Ottawa, Canada) for providing a sample of meleagrin.
Ethical statement
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
All the authors declare that they have no conflict of interest in this article.
Author information
Authors and Affiliations
Corresponding author
Additional information
K. Kosalková and R. Domínguez-Santos contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 383 kb)
Rights and permissions
About this article
Cite this article
Kosalková, K., Domínguez-Santos, R., Coton, M. et al. A natural short pathway synthesizes roquefortine C but not meleagrin in three different Penicillium roqueforti strains. Appl Microbiol Biotechnol 99, 7601–7612 (2015). https://doi.org/10.1007/s00253-015-6676-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-6676-0