Abstract
Main conclusion
ITS2+ trnH - psbA was the best combination of DNA barcode to resolve the Dalbergia wood species studied. We demonstrate the feasibility of building a DNA barcode reference database using xylarium wood specimens.
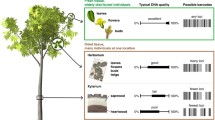
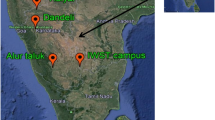
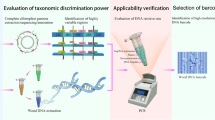
The increase in illegal logging and timber trade of CITES-listed tropical species necessitates the development of unambiguous identification methods at the species level. For these methods to be fully functional and deployable for law enforcement, they must work using wood or wood products. DNA barcoding of wood has been promoted as a promising tool for species identification; however, the main barrier to extensive application of DNA barcoding to wood is the lack of a comprehensive and reliable DNA reference library of barcodes from wood. In this study, xylarium wood specimens of nine Dalbergia species were selected from the Wood Collection of the Chinese Academy of Forestry and DNA was then extracted from them for further PCR amplification of eight potential DNA barcode sequences (ITS2, matK, trnL, trnH-psbA, trnV-trnM1, trnV-trnM2, trnC-petN, and trnS-trnG). The barcodes were tested singly and in combination for species-level discrimination ability by tree-based [neighbor-joining (NJ)] and distance-based (TaxonDNA) methods. We found that the discrimination ability of DNA barcodes in combination was higher than any single DNA marker among the Dalbergia species studied, with the best two-marker combination of ITS2+trnH-psbA analyzed with NJ trees performing the best (100% accuracy). These barcodes are relatively short regions (<350 bp) and amplification reactions were performed with high success (≥90%) using wood as the source material, a necessary factor to apply DNA barcoding to timber trade. The present results demonstrate the feasibility of using vouchered xylarium specimens to build DNA barcoding reference databases.


Similar content being viewed by others
References
Alvarez I, Wendel JF (2003) Ribosomal ITS sequences and plant phylogenetic inference. Mol Phylogenet Evol 29(3):417–434
Alves TLS, Chauveau O, Eggers L, Souza-Chies TTD (2014) Species discrimination in Sisyrinchium (Iridaceae): assessment of DNA barcodes in a taxonomically challenging genus. Mol Ecol Resour 14(2):324–335
Austerlitz F, David O, Schaeffer B, Bleakley K, Olteanu M, Leblois R, Veuille M, Laredo C (2009) DNA barcode analysis: a comparison of phylogenetic and statistical classification methods. BMC Bioinformatics 10(14):S10
Baker CS (2008) A truer measure of the market: the molecular ecology of fisheries and wildlife trade. Mol Ecol 17(18):3985–3998
Bergo MCJ, Pastore TCM, Coradin VTR, Wiedenhoeft AC, Braga JWB (2016) NIRS identification of Swietenia macrophylla is robust across specimens from 27 countries. IAWA J 37(3):420–430
Besnard G, Rubio RRd, Vargas P (2007) Plastid and nuclear DNA polymorphism reveals historical processes of isolation and reticulation in the olive tree complex (Olea europaea). J Biogeogr 34(4):736–752
Bhagwat RM, Dholakia BB, Kadoo NY, Balasundaran M, Gupta VS (2015) Two new potential barcodes to discriminate Dalbergia species. PLoS One 10(11):e0142965
Bolson M, Smidt EDC, Brotto ML, Silvapereira V (2015) ITS and trnH-psbA as efficient barcodes to identify threatened commercial woody angiosperms from southern Brazilian Atlantic rainforests. PLoS One 10(12):e0143049
Cai ZM, Zhang YX, Zhang LN, Gao LM, Li DZ (2012) Testing four candidate barcoding markers in temperate woody bamboos (Poaceae: Bambusoideae). J Syst Evol 50(6):527–539
Casiraghi M, Labra M, Ferri E, Galimberti A, De MF (2010) DNA barcoding: a six-question tour to improve users’ awareness about the method. Brief Bioinform 11(4):440–453
CBOL Plant Working Group (2009) A DNA barcode for land plants. Proc Natl Acad Sci USA 106(31):12794–12797
Chase MW, Cowan RS, Hollingsworth PM, Berg CVD, Madriñán S, Petersen G, Seberg O, Jørgsensen T, Cameron KM, Carine M (2007) A proposal for a standardised protocol to barcode all land plants. Taxon 56(2):295–299
Chen S, Hui Y, Han J, Chang L, Song J, Shi L, Zhu Y, Ma X, Gao T, Pang X (2010) Validation of the region as a novel DNA barcode for identifying medicinal plant species. PLoS One 5(1):e8613
Cody RB, Dane AJ, Dawson-Andoh B, Adedipe EO, Nkansah K (2012) Rapid classification of White Oak (Quercus alba) and Northern Red Oak (Quercus rubra) by using pyrolysis direct analysis in real time (DART™) and time-of-flight mass spectrometry. J Anal Appl Pyrol 95:134–137
Cowan RS, Chase MW, Kress WJ, Savolainen V (2006) 300,000 species to identify: problems, progress, and prospects in DNA barcoding of land plants. Taxon 55(3):611–616
Degen B, Ward SE, Lemes MR, Navarro C, Cavers S, Sebbenn AM (2013) Verifying the geographic origin of mahogany (Swietenia macrophylla King) with DNA-fingerprints. Forensic Sci Int-Gen 7(1):55–62
Deguilloux FM, Petit R (2004) DNA-based control of oak wood geographic origin in the context of the cooperage industry. Ann Forest Sci 61(1):97–104
Eurlings MCM, Lens F, Pakusza C, Peelen T, Wieringa JJ, Gravendeel B (2013) Forensic identification of indian snakeroot (Rauvolfia serpentina Benth. ex Kurz) using DNA barcoding. J Forensic Sci 58(3):822–830
Fazekas AJ, Burgess KS, Kesanakurti PR, Graham SW, Newmaster SG, Husband BC, Percy DM, Hajibabaei M, Barrett SCH (2007) Multiple multilocus DNA barcodes from the plastid genome discriminate plant species equally well. PLoS One 3(7):e2802
Feliner GN, Rossello JA (2007) Better the devil you know? Guidelines for insightful utilization of nrDNA ITS in species-level evolutionary studies in plants. Mol Phylogenet Evol 44(2):911–919
Frézal L, Leblois R (2008) Four years of DNA barcoding: current advances and prospects. Infect Genet Evol 8(5):727–736
Gao LM, Yan LI, Phan LK, Yan LJ, Thomas P, Long KP, Möller M, Li DZ (2017) DNA barcoding of East Asian Amentotaxus (Taxaceae): potential new species and implications for conservation. J Syst Evol 55(1):16–24
Gathier G, van der Niet T, Peelen T, van Vugt RR, Eurlings MC, Gravendeel B (2013) Forensic identification of CITES protected slimming cactus (Hoodia) using DNA barcoding. J Forensic Sci 58(6):1467–1471
Gismondi A, Rolfo MF, Leonardi D, Rickards O, Canini A (2012) Identification of ancient Olea europaea L. and Cornus mas L. seeds by DNA barcoding. CR Biol 335(7):472–479
Gismondi A, Marco GD, Delorenzo M, Canini A (2015) Upgrade of Castanea sativa (Mill.) genetic resources by sequencing of barcode markers. J Genet 94(3):519–524
Gismondi A, Marco GD, Martini F, Sarti L, Crespan M, Martínez-Labarga C, Rickards O, Canini A (2016) Grapevine carpological remains revealed the existence of a Neolithic domesticated Vitis vinifera L. specimen containing ancient DNA partially preserved in modern ecotypes. J Archaeol Sci 69:75–84
Gould BA, León B, Buffen AM, Thompson LG (2010) Evidence of a high-Andean, mid-Holocene plant community: an ancient DNA analysis of glacially preserved remains. Am J Bot 97(9):1579–1584
Hajibabaei M, Singer GAC, Hebert PD, Hickey DA (2007) DNA barcoding: how it complements taxonomy, molecular phylogenetics and population genetics. Trends Genet 23(4):167–172
Han J, Zhu Y, Chen X, Liao B, Yao H, Song J, Chen S, Meng F (2013) The short ITS2 sequence serves as an efficient taxonomic sequence tag in comparison with the full-length ITS. Biomed Res Int 1:741476
Hartvig I, Czako M, Kjær ED, Nielsen LR, Theilade I (2015) The use of DNA barcoding in identification and conservation of rosewood (Dalbergia spp.). PLoS One 10(9):e0138231
Hassold S, Nd LP, Bauert MR, Razafintsalama A, Ramamonjisoa L, Widmer A (2016) DNA barcoding of malagasy rosewoods: towards a molecular identification of CITES-listed Dalbergia species. PLoS One 11(6):e0157881
Hebert PD, Cywinska A, Ball SL (2003) Biological identifications through DNA barcodes. Proc R Soc Lond Ser B Biol Sci 270(1512):313–321
Hollingsworth PM, Forrest LL, Spouge JL, Hajibabaei M, Ratnasingham S, Bank MVD, Chase MW, Cowan RS, Erickson DL (2009) From the cover: a DNA barcode for land plants. Proc Natl Acad Sci USA 106(31):12794–12797
Hollingsworth PM, Graham SW, Little DP (2011) Choosing and using a plant DNA barcode. PLoS One 6(5):e19254
Horacek M, Jakusch M, Krehan H (2009) Control of origin of larch wood: discrimination between European (Austrian) and Siberian origin by stable isotope analysis. Rapid Commun Mass Spectrom 23(23):3688–3692
Huang X, Ci X, Conran JG, Li J (2015) Application of DNA barcodes in Asian tropical trees—a case study from Xishuangbanna nature reserve, southwest China. PLoS One 10(6):e0129295
Ji Y, Ashton L, Pedley SM, Edwards DP, Tang Y, Nakamura A, Kitching R, Dolman PM, Woodcock P, Edwards FA (2013) Reliable, verifiable and efficient monitoring of biodiversity via metabarcoding. Ecol Lett 16(10):1245–1257
Jiao L, Yin Y, Xiao F, Sun Q, Song K, Jiang X (2012) Comparative analysis of two DNA extraction protocols from fresh and dried wood of Cunninghamia lanceolata (Taxodiaceae). IAWA J 33(4):441–456
Jiao L, Yin Y, Cheng Y, Jiang X (2014) DNA barcoding for identification of the endangered species Aquilaria sinensis: comparison of data from heated or aged wood samples. Holzforschung 68(4):487–494
Jiao L, Liu X, Jiang X, Yin Y (2015) Extraction and amplification of DNA from aged and archaeological Populus euphratica wood for species identification. Holzforschung 69(8):925–931
Jörger KM, Schrödl M (2014) How to use CAOS software for taxonomy? A quick guide to extract diagnostic nucleotides or amino acids for species descriptions. Spixiana 37(1):21–26
Kagawa A, Leavitt SW (2010) Stable carbon isotopes of tree rings as a tool to pinpoint the geographic origin of timber. J Wood Sci 56(3):175–183
Kress WJ, Wurdack KJ, Zimmer EA, Weigt LA, Janzen DH (2005) Use of DNA barcodes to identify flowering plants. Proc Natl Acad Sci USA 102(23):8369–8374
Krüger I, Muhr J, Hartl-Meier C, Schulz C, Borken W (2014) Age determination of coarse woody debris with radiocarbon analysis and dendrochronological cross-dating. Eur J For Res 133(5):931–939
Kumar S, Nei M, Dudley J, Tamura K (2008) MEGA: a biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief Bioinform 9(4):299–306
Lahaye R, van der Bank M, Bogarin D, Warner J, Pupulin F, Gigot G, Maurin O, Duthoit S, Barraclough TG, Savolainen V (2008) DNA barcoding the floras of biodiversity hotspots. Proc Natl Acad Sci USA 105(8):2923–2928
Lee SY, Ng WL, Mahat MN, Nazre M, Mohamed R (2016) DNA barcoding of the endangered Aquilaria (Thymelaeaceae) and its application in species authentication of agarwood products traded in the market. PLoS One 4:e0154631
Little DP, Stevenson DW (2007) A comparison of algorithms for the identification of specimens using DNA barcodes: examples from gymnosperms. Cladistics 23(1):1–21
Lowe AJ, Cross HB (2011) The application of DNA methods to timber tracking and origin verification. IAWA J 32(2):251–262
Lowe AJ, Dormontt EE, Bowie MJ, Degen B, Gardner S, Thomas D, Clarke C, Rimbawanto A, Wiedenhoeft A, Yin Y (2016) Opportunities for improved transparency in the timber trade through scientific verification. Bioscience 66(11):990–998
Lv T, Teng R, Shao Q, Wang H, Zhang W, Li M, Zhang L (2015) DNA barcodes for the identification of Anoectochilus roxburghii and its adulterants. Planta 242(5):1167–1174
Mcclure PJ, Chavarria GD, Espinoza E (2015) Metabolic chemotypes of CITES protected Dalbergia timbers from Africa, Madagascar, and Asia. Rapid Commun Mass Spectrom 29(9):783–788
Meier R, Shiyang K, Vaidya G, Ng PK (2006) DNA barcoding and taxonomy in Diptera: a tale of high intraspecific variability and low identification success. Syst Biol 55(5):715–728
Newmaster SG, Subramanyam R (2009) Testing plant barcoding in a sister species complex of pantropical Acacia (Mimosoideae, Fabaceae). Mol Ecol Resour 9(Suppl. 1):172–180
Nithaniyal S, Newmaster SG, Ragupathy S, Krishnamoorthy D, Vassou SL, Parani M (2014) DNA barcode authentication of wood samples of threatened and commercial timber trees within the tropical dry evergreen forest of India. PLoS One 9(9):e107669
Ogata T, Yahara S, Hisatsune R, Konishi R, Nohara T (1990) Isoflavan and related compounds from Dalbergia odorifera. Chem Pharma Bull 38:2750–2755
Pang X, Liu C, Shi L, Liu R, Liang D, Li H, Cherny SS, Chen S (2012) Utility of the trnH-psbA intergenic spacer region and its combinations as plant DNA barcodes: a meta-analysis. PLoS One 7(11):e48833
Pastore TCM, Braga JWB, Coradin VTR, Okino EYA, Camargos JAA, De Muñiz GIB, Bressan OA, Davrieux F (2011) Near infrared spectroscopy (NIRS) as a potential tool for monitoring trade of similar woods: discrimination of true mahogany, cedar, andiroba, and curupixá. Holzforschung 65(1):73–80
Phong DT, Tang DV, Hien VTT, Ton ND, Hai NV (2014) Nucleotide diversity of a nuclear and four chloroplast DNA regions in rare tropical wood species of Dalbergia in Vietnam: a DNA barcode identifying utility. Asian J Appl Sci 2(2):116–125
Poczai P, Hyvönen J (2010) Nuclear ribosomal spacer regions in plant phylogenetics: problems and prospects. Mol Biol Rep 37(4):1897–1912
Puillandre N, Bouchet P, Boisselier-Dubayle MC, Brisset J, Buge B, Castelin M, Chagnoux S, Christophe T, Corbari L, Lambourdière J (2012) New taxonomy and old collections: integrating DNA barcoding into the collection curation process. Mol Ecol Resour 12(3):396–402
Ragupathy S, Newmaster SG, Murugesan M, Balasubramaniam V (2009) DNA barcoding discriminates a new cryptic grass species revealed in an ethnobotany study by the hill tribes of the Western Ghats in southern India. Mol Ecol Resour 9(Suppl. 1):164–171
Reboredo F (2013) Socio-economic, environmental, and governance impacts of illegal logging. Environ Syst Decis 33(2):295–304
Ren H, Lu LM, Soejima A, Luke Q, Zhang DX, Chen ZD, Wen J (2011) Phylogenetic analysis of the grape family (Vitaceae) based on the noncoding plastid trnC-petN, trnH-psbA, and trnL-F sequences. Taxon 60(3):629–637
Ribeiro R, Lemos-Filho J, Ramos A, Lovato M (2011) Phylogeography of the endangered rosewood Dalbergia nigra (Fabaceae): insights into the evolutionary history and conservation of the Brazilian Atlantic Forest. Heredity 106(1):46–57
Santos GU, Callado CH, da Costa Souza M, Costa CG (2013) Wood Anatomy of Myrciaria, Neomitranthes, Plinia and Siphoneugena Species (Myrteae, Myrtaceae). IAWA J 34(3):313–323
Sarkar IN, Planet PJ, Desalle R (2008) CAOS Software for use in character based DNA barcoding. Mol Ecol Resour 8(6):1256–1259
Schroeder H, Cronn R, Yanbaev Y, Jennings T, Mader M, Degen B, Kersten B (2016) Development of molecular markers for determining continental origin of wood from white oaks (Quercus L. sect. Quercus). PLoS One 11(6):e0158221
Taberlet P, Coissac E, Pompanon F, Gielly L, Miquel C, Valentini A, Vermat T, Corthier G, Brochmann C, Willerslev E (2007) Power and limitations of the chloroplast trnL (UAA) intron for plant DNA barcoding. Nucleic Acids Res 35(3):e14
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739
Tang X, Zhao G, Ping L (2011) Wood identification with PCR targeting noncoding chloroplast DNA. Plant Mol Biol 77(6):609–617
Tao Y, Wang Y (2010) Bioactive sesquiterpenes isolated from the essential oil of Dalbergia odorifera T. Chen. Fitoterapia 81(5):393–396
The IUCN Red List of Threatened Species Version (2016). https://www.iucnredlist.org/. Accessed 5 Apr 2017
The Plant List Version 1.1 (2013). http://www.theplantlist.org/. Accessed 5 Apr 2017
Wang W, Weng X, Cheng D (2000) Antioxidant activities of natural phenolic components from Dalbergia odorifera T. Chen. Food Chem 71(1):45–49
Wiedenhoeft A (2014) Curating xylaria. In: Salick J, Konchar K, Nesbitt M (eds) Curating biocultural collections: a handbook. Kew Publishing in Association with Missouri Botanical Garden, UK, pp 127–134
Xu C, Dong W, Shi S, Cheng T, Li C, Liu Y, Wu P, Wu H, Gao P, Zhou S (2015) Accelerating plant DNA barcode reference library construction using herbarium specimens: improved experimental techniques. Mol Ecol Resour 15(6):1366–1374
Yan LJ, Liu J, Möller M, Zhang L, Zhang XM, Li DZ, Gao LM (2015) DNA barcoding of Rhododendron (Ericaceae), the largest Chinese plant genus in biodiversity hotspots of the Himalaya-Hengduan Mountains. Mol Ecol Resour 15(4):932–944
Yang J, Vázquez L, Chen X, Li H, Zhang H, Liu Z, Zhao G (2017) Development of chloroplast and nuclear DNA markers for Chinese Oaks (Quercus Subgenus Quercus) and assessment of their utility as DNA barcodes. Fronti Plant Sci 8:816
Yao H, Song J, Liu C, Luo K, Han J, Li Y, Pang X, Xu H, Zhu Y, Xiao P (2010) Use of ITS2 region as the universal DNA barcode for plants and animals. PLoS One 5(10):e13102
Yu M, Liu K, Zhou L, Zhao L, Liu S (2016) Testing three proposed DNA barcodes for the wood identification of Dalbergia odorifera T. Chen and Dalbergia tonkinensis Prain. Holzforschung 70(2):127–136
Zemlak TS, Ward RD, Connell AD, Holmes BH, Hebert PD (2009) DNA barcoding reveals overlooked marine fishes. Mol Ecol Resour 9(s1):237–242
Zhang M, Fritsch PW, Cruz BC (2009) Phylogeny of Caragana (Fabaceae) based on DNA sequence data from rbcL, trnS–trnG, and ITS. Mol Phylogenet Evol 50(3):547–559
Zhang ZL, Song MF, Guan YH, Li HT, Niu YF, Zhang LX, Ma XJ (2015) DNA barcoding in medicinal plants: testing the potential of a proposed barcoding marker for identification of Uncaria species from China. Biochem Syst Ecol 60:8–14
Acknowledgements
This work was supported by the China Postdoctoral Science Foundation (Grant no. 2016M590152), the National Natural Science Foundation of China (Grant no. 31600451), the Fundamental Research Funds of Chinese Academy of Forestry (Grant no. CAFYBB2017ZE003), the China Scholarship Council (Grant no. 2016-3035), and U.S. State Department Interagency Agreement (19318814Y0010-140001-0001/P00001).We thank Sarah Friedrich, Department of Botany, University of Wisconsin-Madison for her technical assistance with the phylogram figure.
Author information
Authors and Affiliations
Corresponding author
Additional information
Min Yu and Lichao Jiao contributed to the work equally and should be regarded as co-first authors.
Electronic supplementary material
Below is the link to the electronic supplementary material.
425_2017_2758_MOESM1_ESM.pdf
Supplementary material 1 (PDF 2918 kb) Fig. S1 Relative distributions of interspecific and intraspecific Kimura two-parameter (K2P) distances for the eight DNA markers and their combinations
425_2017_2758_MOESM2_ESM.pdf
Supplementary material 2 (PDF 1372 kb) Fig. S2 Taxon identification trees constructed using neighbor-joining analysis of Kimura 2-parameter (K2P) distances showing patterns of the eight DNA markers and their combinations
Rights and permissions
About this article
Cite this article
Yu, M., Jiao, L., Guo, J. et al. DNA barcoding of vouchered xylarium wood specimens of nine endangered Dalbergia species. Planta 246, 1165–1176 (2017). https://doi.org/10.1007/s00425-017-2758-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-017-2758-9