Abstract
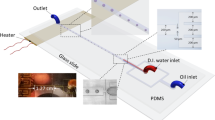
A two-temperature continuous-flow polymerase chain reaction (PCR) polymer chip has been constructed that takes advantage of droplet technology to avoid sample contamination and adsorption at the surface. Samples contained in aqueous droplets are continuously moved by an oil carrier-fluid through various temperature zones, introducing the possibility of real-time quantitative PCR. In the present paper, we investigate many of the factors affecting droplet-based PCR chip design, including thermal mass, flow rate, and thermal resistance. The study focuses particularly on the fluid and substrate temperature distribution within the PCR chip and the droplet residence times in critical temperature zones. The simulations demonstrate that the flow rate strongly affects the temperature field within the carrier-fluid. Above a critical flow rate, the carrier-fluid fails to achieve the required temperatures for DNA amplification. In addition, the thermal resistances of the different layers in the chip are shown to have a major impact on the temperature profile in the channel.

















Similar content being viewed by others
Abbreviations
- c p :
-
specific heat capacity, J/(kg K)
- d :
-
droplet diameter, m
- d 1 :
-
thickness of cellulose acetate layer, m
- d 2 :
-
thickness of polycarbonate layer, m
- f b :
-
buoyancy force, N
- f Saff :
-
Saffman lift force, N
- g :
-
acceleration due to gravity, m/s2
- k :
-
thermal conductivity, W/(m K)
- p :
-
pressure, N/m2
- Q :
-
flow rate, μl/min
- R 1 :
-
thermal resistance of acetate layer, m2 K/W
- R 2 :
-
thermal resistance of polycarbonate layer, m2 K/W
- R 3 :
-
thermal resistance of natural convection, m2 K/W
- Re :
-
Reynolds number, -
- T :
-
temperature, K
- t :
-
time, s
- V :
-
velocity vector, m/s
- V r :
-
relative velocity, m/s
- γ :
-
DNA amplification efficiency, -
- μ :
-
dynamic viscosity, N s/m2
- ν :
-
kinematic viscosity, m2/s
- ρ :
-
density, kg/m3
- τ :
-
shear stress, N/m2
- 1,2,3:
-
layers 1, 2, and 3
References
Auroux PA, Iossifidis D, Reyes DR, Manz A (2002) Micro total analysis systems 2: analytical standard operations and applications. Anal Chem 74:2637–2652
Auroux PA, Koc Y, de Mello A, Manz A, Day PJR (2004) Miniaturised nucleic acid analysis. Lab Chip 4:534–546
Auroux PA, Day PJR, Manz A (2005) Quantitative study of the adsorption of PCR reagents during on-chip bi-directional shunting PCR. In: Proceedings of the 9th International conference on miniaturized systems for chemistry and life sciences, Boston, MA, USA
Belgrader P, Benett W, Hadley D, Richards J, Stratton P, Mariella R, Milanovich F (1999) Infectious disease: PCR detection of bacteria in seven minutes. Science 284:449–450
Bu MQ, Tracy M, Ensell G, Wilkinson JS, Evans AGR (2003) Design and theoretical evaluation of a novel microfluidic device to be used for PCR. J Micromech Microeng 13:S125–S130
CFD-ACE+ User Manual Version (2006) ESI CFD Inc., Huntsville, AL 35806, USA
Chen ZY, Qian SZ, Abrams WR, Malamud D, Bau HH (2004) Thermosiphon-based PCR reactor: experiment and modeling. Anal Chem 76:3707–3715
Chiou J, Matsudaira P, Sonin A, Ehrlich D (2001) A closed-cycle capillary polymerase chain reaction machine. Anal Chem 73:2018–2021
Gulliksen A, Solli L, Karlsen F, Rogne H, Hovig E, Nordstrom T, Sirevag R (2004) Real-time nucleic acid sequence-based amplification in nanoliter volumes. Anal Chem 76:9–14
Hardt S, Dadic D, Doffing F, Drese KS, Münchow G, Sörensen O (2004) Development of a slug-flow PCR chip with minimum heating cycle times. Nanotechnology 1:55–58
Kopp MU, de Mello AJ, Manz A (1998) Chemical amplification: continuous-flow PCR on a chip. Science 280:1046–1048
Lee JY, Rohlman CE, Molony LA, Engelke DR (1991) Characterization of RPR1, an essential gene encoding the RNA component of saccharomyces-cerevisiae nuclear RNASE-P. Mol Cell Biol 11(2):721–730
Liu J, Enzelberger M, Quake S (2002) A nanoliter rotary device for polymerase chain reaction. Electrophoresis 23:1531–1536
Nakano H, Matsuda K, Yohda M, Nagamune T, Endo I, Yamane T (1994) High-speed polymerase chain-reaction in constant flow. Biosci Biotechnol Biochem58:349–352
Nisisako T, Torii T, Higuchi T (2002) Droplet formation in a microchannel network. Lab Chip 2:24–26
Northrup MA, Ching MT, White RM, Watson RT (1993) DNA amplification with a microfabricated reaction chamber. Transducers 93:924–926
Park N, Kim S, Hahn JH (2003) Cylindrical compact thermal-cycling device for continuous-flow polymerase chain reaction. Anal Chem 75:6029–6033
Sadler DJ, Changrani R, Roberts P, Chou CF, Zenhausern F (2003) Thermal management of bioMEMS: temperature control for ceramic-based PCR and DNA detection devices. IEEE Trans Compon Packag Technol 26:309–316
Saffman PG (1965) The lift on a small sphere in a slow shear flow. J Fluid Mech 22:385–400; Corrigendum (1968) 31:624
Saiki RK, Scharf S, Faloona F, Mullis KB, Horn GT, Erlich HA, Arnheim N (1985) Enzymatic amplification of betaglobin genomic sequences and restriction site analysis for diagnosis of sickle-cell anemia. Science 230:1350–1354
Vilkner T, Janasek D, Manz A (2004) Micro total analysis systems: recent developments. Anal Chem 76:3373–3385
West J, Karamata B, Lillis B, Gleeson JP, Alderman J, Collins JK, Lane W, Mathewson A, Berney H (2002) Application of magnetohydrodynamic actuation to continuous flow chemistry. Lab Chip 2:224–230
Wilding P, Shoffner MA, Kricka LJ (1994) PCR in a silicon microstructure. Clin Chem 40:1815–1818
Wittwer CT, Garling DJ (1991) Rapid cycle DNA amplification: time and temperature optimisation. Biotechniques 10:76–83
Wittwer CT, Herrmann MG, Gundry CN, Elenitoba-Johnson KSJ (2001) Real-time multiplex PCR assays. Methods 25:430–442
Zhang Q, Wang W, Zhang H, Wang Y (2002) Temperature analysis of continuous-flow micro-PCR based on FEA. Sens Actuators B 82:75–81
Acknowledgments
The authors are grateful to the UK Engineering and Physical Sciences Research Council (EPSRC) for supporting this research under Grant No. GR/S82978/01. Additional support was provided by EPSRC under the auspices of Collaborative Computational Project 12 (CCP12). Thermographic Measurement Ltd., Flintshire, UK is also acknowledged for supplying samples of the thermochromic dye.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mohr, S., Zhang, YH., Macaskill, A. et al. Numerical and experimental study of a droplet-based PCR chip. Microfluid Nanofluid 3, 611–621 (2007). https://doi.org/10.1007/s10404-007-0153-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10404-007-0153-8