Abstract
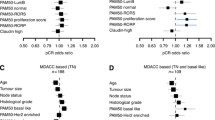
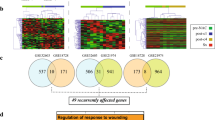
We aimed to investigate the association between gene co-expression modules and responses to neoadjuvant chemotherapy in breast cancer by using a systematic biological approach. The gene expression profiles and clinico-pathological data of 508 (discovery set) and 740 (validation set) patients with breast cancer who received neoadjuvant chemotherapy were analyzed. Weighted gene co-expression network analysis was performed and identified seven co-regulated gene modules. Each module and gene signature were evaluated with logistic regression models for pathological complete response (pCR). The association between modules and pCR in each intrinsic molecular subtype was also investigated. Two transcriptional modules were correlated with tumor grade, estrogen receptor status, progesterone receptor status, and chemotherapy response in breast cancer. One module that constitutes upregulated cell proliferation genes was associated with a high probability for pCR in the whole (odds ratio (OR) = 5.20 and 3.45 in the discovery and validation datasets, respectively), luminal B, and basal-like subtypes. The prognostic potentials of novel genes, such as MELK, and pCR-related genes, such as ESR1 and TOP2A, were identified. The upregulation of another gene co-expression module was associated with weak chemotherapy responses (OR = 0.19 and 0.33 in the discovery and validation datasets, respectively). The novel gene CA12 was identified as a potential prognostic indicator in this module. A systems biology network-based approach may facilitate the discovery of biomarkers for predicting chemotherapy responses in breast cancer and contribute in developing personalized medicines.



Similar content being viewed by others
Abbreviations
- WGCNA:
-
Weighted gene co-expression network analysis
- TOM:
-
Topological overlap measure
- k.total:
-
Network connectivity
- k.in:
-
Intramodular connectivity
- ME:
-
Module eigengene
- ER:
-
Estrogen receptor
- PR:
-
Progesterone receptor
- HER2:
-
Human epidermal growth factor 2
- PCC:
-
Pearson’s correlation coefficient
- CI:
-
Confidence interval
- GS:
-
Gene significance
- AUC:
-
Area under the receiver operating characteristic curve
- OR:
-
Odds ratio
- pCR:
-
Pathological complete response
- RD:
-
Residual disease
- GGI:
-
Gene expression grade index
- GEO:
-
Gene expression omnibus
- LN:
-
Lymph node
References
von Minckwitz G et al (2012) Neoadjuvant treatments for triple-negative breast cancer (TNBC). Ann Oncol 23(6):35–39
Hatzis C et al (2011) A genomic predictor of response and survival following taxane-anthracycline chemotherapy for invasive breast cancer. JAMA 305(18):1873–1881
Tabchy A et al (2010) Evaluation of a 30-gene paclitaxel, fluorouracil, doxorubicin, and cyclophosphamide chemotherapy response predictor in a multicenter randomized trial in breast cancer. Clin Cancer Res 16(21):5351–5361
Bonadonna G et al (1998) Primary chemotherapy in operable breast cancer: eight-year experience at the Milan Cancer Institute. J Clin Oncol 16(1):93–100
Rastogi P et al (2008) Preoperative chemotherapy: updates of national surgical adjuvant breast and bowel project protocols B-18 and B-27. J Clin Oncol 26(5):778–785
Symmans WF et al (2007) Measurement of residual breast cancer burden to predict survival after neoadjuvant chemotherapy. J Clin Oncol 25(28):4414–4422
Rouzier R et al (2005) Nomograms to predict pathologic complete response and metastasis-free survival after preoperative chemotherapy for breast cancer. J Clin Oncol 23(33):8331–8339
Huober J et al (2010) Effect of neoadjuvant anthracycline-taxane-based chemotherapy in different biological breast cancer phenotypes: overall results from the GeparTrio study. Breast Cancer Res Treat 124(1):133–140
Silver DP et al (2010) Efficacy of neoadjuvant Cisplatin in triple-negative breast cancer. J Clin Oncol 28(7):1145–1153
Tordai A et al (2008) Evaluation of biological pathways involved in chemotherapy response in breast cancer. Breast Cancer Res 10(2):29
Witkiewicz AK et al (2014) Systematically defining single-gene determinants of response to neoadjuvant chemotherapy reveals specific biomarkers. Clin Cancer Res 20(18):4837–4848
Sotiriou C et al (2006) Gene expression profiling in breast cancer: understanding the molecular basis of histologic grade to improve prognosis. J Natl Cancer Inst 98(4):262–272
van’t Veer LJ et al (2002) Gene expression profiling predicts clinical outcome of breast cancer. Nature 415(6871):530–536
Paik S et al (2004) A multigene assay to predict recurrence of tamoxifen-treated, node-negative breast cancer. N Engl J Med 351(27):2817–2826
Liedtke C et al (2009) Genomic grade index is associated with response to chemotherapy in patients with breast cancer. J Clin Oncol 27(19):3185–3191
Gianni L et al (2005) Gene expression profiles in paraffin-embedded core biopsy tissue predict response to chemotherapy in women with locally advanced breast cancer. J Clin Oncol 23(29):7265–7277
Straver ME et al (2010) The 70-gene signature as a response predictor for neoadjuvant chemotherapy in breast cancer. Breast Cancer Res Treat 119(3):551–558
Perou CM et al (2000) Molecular portraits of human breast tumours. Nature 406(6797):747–752
Sotiriou C et al (2009) Gene-expression signatures in breast cancer. N Engl J Med 360(8):790–800
Parker JS et al (2009) Supervised risk predictor of breast cancer based on intrinsic subtypes. J Clin Oncol 27(8):1160–1167
Rouzier R et al (2005) Breast cancer molecular subtypes respond differently to preoperative chemotherapy. Clin Cancer Res 11(16):5678–5685
Clarke C et al (2013) Correlating transcriptional networks to breast cancer survival: a large-scale coexpression analysis. Carcinogenesis 34(10):2300–2308 Epub 2013/06/07
Wirapati P et al (2008) Meta-analysis of gene expression profiles in breast cancer: toward a unified understanding of breast cancer subtyping and prognosis signatures. Breast Cancer Res 10(4):28
Ivliev AE et al (2010) Coexpression network analysis identifies transcriptional modules related to proastrocytic differentiation and sprouty signaling in glioma. Cancer Res 70(24):10060–10070
Liu R et al (2015) Identification and validation of gene module associated with lung cancer through coexpression network analysis. Gene 563(1):56–62
Udyavar AR et al (2013) Co-expression network analysis identifies Spleen Tyrosine Kinase (SYK) as a candidate oncogenic driver in a subset of small-cell lung cancer. BMC Syst Biol 7(5):1752
Liu R et al (2015) Network-based approach to identify prognostic biomarkers for estrogen receptor-positive breast cancer treatment with tamoxifen. Cancer Biol Ther 16(2):317–324
Presson AP et al (2011) Protein expression based multimarker analysis of breast cancer samples. BMC Cancer 11(230):1471–2407
RCTR (2014) A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL http://www.R-projectorg/
Iwamoto T et al (2011) Gene pathways associated with prognosis and chemotherapy sensitivity in molecular subtypes of breast cancer. J Natl Cancer Inst 103(3):264–272
Shi L et al (2010) The microarray quality control (MAQC)-II study of common practices for the development and validation of microarray-based predictive models. Nat Biotechnol 28(8):827–838
Desmedt C et al (2011) Multifactorial approach to predicting resistance to anthracyclines. J Clin Oncol 29(12):1578–1586
Johnson WE et al (2007) Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics. 8(1):118–127
Miller J et al (2011) Strategies for aggregating gene expression data: the collapseRows R function. BMC Bioinform 12(1):322
Zhang B et al (2005) A general framework for weighted gene co-expression network analysis. Stat Appl Genet Mol Biol 4:17
Langfelder P et al (2008) WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics 9(559):1471–2105
Nakayama S et al (2009) Prediction of paclitaxel sensitivity by CDK1 and CDK2 activity in human breast cancer cells. Breast Cancer Res 11(1):24
Torikoshi Y et al (2013) Novel functional assay for spindle-assembly checkpoint by cyclin-dependent kinase activity to predict taxane chemosensitivity in breast tumor patient. J Cancer 4(9):697–702
Kim SJ et al (2012) Recurrence risk score based on the specific activity of CDK1 and CDK2 predicts response to neoadjuvant paclitaxel followed by 5-fluorouracil, epirubicin and cyclophosphamide in breast cancers. Ann Oncol 23(4):891–897
Pritchard KI et al (2006) HER2 and responsiveness of breast cancer to adjuvant chemotherapy. N Engl J Med 354(20):2103–2111
Paik S et al (1998) erbB-2 and response to doxorubicin in patients with axillary lymph node-positive, hormone receptor-negative breast cancer. J Natl Cancer Inst 90(18):1361–1370
Knoop AS et al (2005) retrospective analysis of topoisomerase IIa amplifications and deletions as predictive markers in primary breast cancer patients randomly assigned to cyclophosphamide, methotrexate, and fluorouracil or cyclophosphamide, epirubicin, and fluorouracil: Danish breast cancer cooperative group. J Clin Oncol 23(30):7483–7490
O’Malley FP et al (2009) Topoisomerase II alpha and responsiveness of breast cancer to adjuvant chemotherapy. J Natl Cancer Inst 101(9):644–650
Warsow G et al (2013) Differential network analysis applied to preoperative breast cancer chemotherapy response. PLoS One 8(12):e81784
Ross-Innes CS et al (2012) Differential oestrogen receptor binding is associated with clinical outcome in breast cancer. Nature 481(7381):389–393
Xu C et al (2015) FOXA1 expression significantly predict response to chemotherapy in estrogen receptor-positive breast cancer patients. Ann Surg Oncol 22(6):2034–2039
Horimoto Y et al (2015) Low FOXA1 expression predicts good response to neo-adjuvant chemotherapy resulting in good outcomes for luminal HER2-negative breast cancer cases. Br J Cancer 112(2):345–351
Denkert C et al (2013) RNA-based determination of ESR1 and HER2 expression and response to neoadjuvant chemotherapy. Ann Oncol 24(3):632–639
Tominaga N et al (2012) Clinicopathological analysis of GATA3-positive breast cancers with special reference to response to neoadjuvant chemotherapy. Ann Oncol 23(12):3051–3057
Chen Y et al (2011) Estrogen receptor-related genes as an important panel of predictors for breast cancer response to neoadjuvant chemotherapy. Cancer Lett 302(1):63–68
Salmans ML et al (2013) The estrogen-regulated anterior gradient 2 (AGR2) protein in breast cancer: a potential drug target and biomarker. Breast Cancer Res 15(2):204
Rody A et al (2009) T-cell metagene predicts a favorable prognosis in estrogen receptor-negative and HER2-positive breast cancers. Breast Cancer Res 11(2):9
Ignatiadis M et al (2012) Gene modules and response to neoadjuvant chemotherapy in breast cancer subtypes: a pooled analysis. J Clin Oncol 30(16):1996–2004
Conflict of interest
None of the authors has any conflict of interest regarding this study.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Liu, R., Lv, QL., Yu, J. et al. Correlating transcriptional networks with pathological complete response following neoadjuvant chemotherapy for breast cancer. Breast Cancer Res Treat 151, 607–618 (2015). https://doi.org/10.1007/s10549-015-3428-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-015-3428-x