Abstract
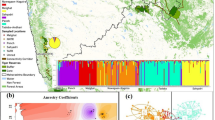
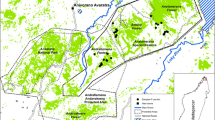
Population decline and fragmentation often lead to reduced genetic diversity and population differentiation. Habitat destruction throughout Madagascar has caused population decline and extinction of many endemic species. Lemur populations, including those of the largest extant lemur, Indri indri, have been fragmented into remaining forest patches. We assessed the level of genetic diversity in indri populations in three protected reserves by genotyping a total of 43 individuals at 17 microsatellite loci. Genetic diversity in terms of heterozygosity was high in all three reserves, with no differences between reserves. Population structure and F ST analyses revealed Analamazaotra Forest Station and the Torotorofotsy Conservation Area, which are separated by ca. 18 km to be genetically differentiated from each other with some admixture. Betampona Strict Nature Reserve, which is separated from the other reserves by ca. 130 km, exhibited clear population genetic differentiation, with no signs of admixture with the other reserves. Our genetic diversity estimates are similar to those for other Indridae in similar habitats and may reflect past rather than current population processes, given that populations have declined recently. Our results suggest that Betampona may be genetically isolated and that it is important to maintain gene flow between remaining populations to prevent loss of genetic diversity for the future conservation of Indri indri.



Similar content being viewed by others
References
Allendorf, F. W., Hohenlohe, P. A., & Luikart, G. (2010). Genomics and the future of conservation genetics. Nature Reviews Genetics, 11(10), 697–709.
Andriaholinirina, N., Baden, A., Blanco, M., Chikhi, L., Cooke, A., et al. (2014). Indri indri. The IUCN Red List of Threatened Species. Version 2014.3. www.iucnredlist.org. Accessed 24 Nov 2014.
Belkhir, K., Borsa, P., Chikhi, L., Raufaste, N., & Bonhomme, F. (1996). GENETIX 4.05, logiciel sous Windows® pour la génétique des populations. Laboratoire génome, populations, interactions, CNRS UMR, 5000, 1996–2004.
Bellemain, E., Nawaz, M. A., Valentini, A., Swenson, J. E., & Taberlet, P. (2007). Genetic tracking of the brown bear in northern Pakistan and implications for conservation. Biological Conservation, 134(4), 537–547.
Britt, A., Randriamandratonirina, N. J., Glasscock, K. D., & Iambana, B. R. (2003). Diet and feeding behaviour of Indri indri in a low-altitude rain forest. Folia Primatologica, 73(5), 225–239.
Chapuis, M. P., & Estoup, A. (2007). Microsatellite null alleles and estimation of population differentiation. Molecular Biology and Evolution, 24(3), 621–631.
Chapuis, M. P., Lecoq, M., Michalakis, Y., Loiseau, A., Sword, G. A., et al. (2008). Do outbreaks affect genetic population structure? A worldwide survey in Locusta migratoria, a pest plagued by microsatellite null alleles. Molecular Ecology, 17(16), 3640–3653.
Chikhi, L., Sousa, V. C., Luisi, P., Goossens, B., & Beaumont, M. A. (2010). The confounding effects of population structure, genetic diversity and the sampling scheme on the detection and quantification of population size changes. Genetics, 186(3), 983–995.
Cornuet, J. M., & Luikart, G. (1996). Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics, 144(4), 2001–2014.
Craul, M., Chikhi, L., Sousa, V., Olivieri, G. L., Rabesandratana, A., et al. (2009). Influence of forest fragmentation on an endangered large-bodied lemur in northwestern Madagascar. Biological Conservation, 142(12), 2862–2871.
Curtis, J. M., & Taylor, E. B. (2004). The genetic structure of coastal giant salamanders (Dicamptodon tenebrosus) in a managed forest. Biological Conservation, 115(1), 45–54.
Cushman, S. A., McKelvey, K. S., & Schwartz, M. K. (2009). Use of empirically derived source-destination models to map regional conservation corridors. Conservation Biology, 23, 368–376.
Di Rienzo, A., Peterson, A. C., Garza, J. C., Valdes, A. M., Slatkin, M., & Freimer, N. B. (1994). Mutational processes of simple-sequence repeat loci in human populations. Proceedings of the National Academy of Sciences of the United States of America, 91(8), 3166–3170.
Evanno, G., Regnaut, S., & Goudet, J. (2005). Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molecular Ecology, 14(8), 2611–2620.
Excoffier, L., & Lischer, H. E. (2010). Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Molecular Ecology Resources, 10(3), 564–567.
Frankham, R., Briscoe, D. A., & Ballou, J. D. (2002). Introduction to conservation genetics. Cambridge: Cambridge University Press.
Gelman, A., & Hill, J. (2006). Data analysis using regression and multilevel/hierarchical models. Cambridge: Cambridge University Press.
Gelman, A., & Rubin, D. B. (1992). Inference from iterative simulation using multiple sequences. Statistical Science 7(4), 457–472.
Glessner, K. D., & Britt, A. (2005). Population density and home range size of Indri indri in a protected low altitude rain forest. International Journal of Primatology, 26(4), 855–872.
Goodman, S. M., & Ganzhorn, J. U. (2004). Elevational ranges of lemurs in the humid forests of Madagascar. International Journal of Primatology, 25(2), 331–350.
Goossens, B., Chikhi, L., Ancrenaz, M., Lackman-Ancrenaz, I., Andau, P., & Bruford, M. W. (2006). Genetic signature of anthropogenic population collapse in orang-utans. PLoS Biology, 4(2), e25.
Goudet, J. (1995). FSTAT (version 1.2): a computer program to calculate F-statistics. Journal of Heredity, 86(6), 485–486.
Groves, C. P. (2001). Primate taxonomy. Washington, DC: Smithsonian Institution Press.
Harper, G. J., Steininger, M. K., Tucker, C. J., Juhn, D., & Hawkins, F. (2007). Fifty years of deforestation and forest fragmentation in Madagascar. Environmental Conservation, 34(4), 325–333.
Holmes, S. M., Baden, A. L., Brenneman, R. A., Engberg, S. E., Louis, E. E., Jr., & Johnson, S. E. (2013). Patch size and isolation influence genetic patterns in black-and-white ruffed lemur (Varecia variegata) populations. Conservation Genetics, 14(3), 615–624.
Junge, R. E., Barrett, M. A., & Yoder, A. D. (2011). Effects of anthropogenic disturbance on indri (Indri indri) health in Madagascar. American Journal of Primatology, 73(7), 632–642.
Jungers, W. L., Godfrey, L. R., Simons, E. L., & Chatrath, P. S. (1995). Subfossil Indri indri from the Ankarana Massif of northern Madagascar. American Journal of Physical Anthropology, 97(4), 357–366.
Keyghobadi, N., Roland, J., Matter, S. F., & Strobeck, C. (2005). Among-and within-patch components of genetic diversity respond at different rates to habitat fragmentation: an empirical demonstration. Proceedings of the Royal Society B: Biological Sciences, 272(1562), 553–560.
Kindlmann, P., & Burel, F. (2008). Connectivity measures: a review. Landscape Ecology, 23(8), 879–890.
Lawler, R. R. (2011). Historical demography of a wild lemur population (Propithecus verreauxi) in southwest Madagascar. Population Ecology, 53(1), 229–240.
Lawler, R. R., Richard, A. F., & Riley, M. A. (2001). Characterization and screening of microsatellite loci in a wild lemur population (Propithecus verreauxi verreauxi). American Journal of Primatology, 55(4), 253–259.
Lippe, C., Dumont, P., & Bernatchez, L. (2006). High genetic diversity and no inbreeding in the endangered copper redhorse, Moxostoma hubbsi (Catostomidae, Pisces): the positive sides of a long generation time. Molecular Ecology, 15(7), 1769–1780.
Mittermeier, R. A., Louis, E., Hawkins, F., Langrand, O., Ganzhorn, J., et al. (2010). Lemurs of Madagascar (3rd ed.). Arlington, VA: Conservation International.
Nei, M., Maruyama, T., & Chakraborty, R. (1975). The bottleneck effect and genetic variability in populations. Evolution, 29, 1–10.
Olivieri, G. L., Sousa, V., Chikhi, L., & Radespiel, U. (2008). From genetic diversity and structure to conservation: Genetic signature of recent population declines in three mouse lemur species (Microcebus spp.). Biological Conservation, 141, 1257–1271.
Peakall, R. O. D., & Smouse, P. E. (2006). GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Molecular Ecology Notes, 6(1), 288–295.
Piry, S., Luikart, G., & Cornuet, J. M. (1999). BOTTLENECK: a computer program for detecting recent reductions in the effective population size using allele frequency data. The Journal of Heredity, 90(4), 502–503.
Pollock, J. (1977). The ecology and sociology of feeding in Indri indri. In T. Clutton-Brock (Ed.), Primate ecology: Studies of feeding and ranging behaviour in lemurs, monkeys and apes (pp. 37–69). London: Academic.
Pollock, J.I. (1975). Field observations on Indri indri: A preliminary report. In Lemur biology (pp. 287–311). New York: Springer-Verlag.
Pollock, J. I. (1979). Spatial distribution and ranging behavior in lemurs. In G. A. Doyle (Ed.), The study of prosimian behavior (pp. 359–409). New York: Academic.
Powzyk, J. A. (1997). The socio-ecology of two sympatric indrids, Propithecus diadema diadema and Indri indri: A comparison of feeding strategies and their possible repercussions on species-specific behaviors. Ph. D. dissertation, Duke University.
Powzyk, J., & Thalmann, U. (2003). Indri indri, indri. In S. M. Goodman & J. P. Benstead (Eds.), The natural history of Madagascar (pp. 1342–1345). Chicago: University of Chicago Press.
Pritchard, J. K., Stephens, M., & Donnelly, P. (2000). Inference of population structure using multilocus genotype data. Genetics, 155(2), 945–959.
Quéméré, E., Amelot, X., Pierson, J., Crouau-Roy, B., & Chikhi, L. (2012). Genetic data suggest a natural prehuman origin of open habitats in northern Madagascar and question the deforestation narrative in this region. Proceedings of the National Academy of Sciences of the United States of America, 109(32), 13028–13033.
Quéméré, E., Louis, E. E., Jr., Ribéron, A., Chikhi, L., & Crouau-Roy, B. (2010). Non-invasive conservation genetics of the critically endangered golden-crowned sifaka (Propithecus tattersalli): high diversity and significant genetic differentiation over a small range. Conservation Genetics, 11(3), 675–687.
R Core Team (2015). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL http://www.R-project.org/.
Rakotoarisoa, G., Shore, G., Mcguire, S., Engberg, S., Louis, E., & Brenneman, R. (2006). Characterization of 20 microsatellite marker loci in Coquerel’s sifaka (Propithecus coquereli). Molecular Ecology Notes, 6(4), 1119–1121.
Raymond, M., & Rousset, F. (1995). GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. Journal of Heredity, 86(3), 248–249.
Razakamaharavo, V. R., McGuire, S. M., Vasey, N., Louis, E. E., Jr., & Brenneman, R. A. (2010). Genetic architecture of two red ruffed lemur (Varecia rubra) populations of Masoala National Park. Primates, 51(1), 53–61.
Schuelke, M. (2000). An economic method for the fluorescent labeling of PCR fragments. Nature Biotechnology, 18(2), 233–234.
Städler, T., Haubold, B., Merino, C., Stephan, W., & Pfaffelhuber, P. (2009). The impact of sampling schemes on the site frequency spectrum in nonequilibrium subdivided populations. Genetics, 182(1), 205–216.
Storz, J. F., & Beaumont, M. A. (2002). Testing for genetic evidence of population expansion and contraction: an empirical analysis of microsatellite DNA variation using a hierarchical Bayesian model. Evolution, 56(1), 154–166.
Templeton, A. R., Shaw, K., Routman, E., & Davis, S. K. (1990). The genetic consequences of habitat fragmentation. Annals of the Missouri Botanical Garden, 77(1), 13–27.
Van Oosterhout, C., Hutchinson, W. F., Wills, D. P., & Shipley, P. (2004). MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Molecular Ecology Notes, 4(3), 535–538.
Weir, B.S., & Cockerham, C.C. (1984). Estimating F-statistics for the analysis of population structure. Evolution, 38(6), 1358–1370.
Whitlock, M. C., & McCauley, D. E. (1999). Indirect measures of gene flow and migration: FST ≠ 1/(4Nm + 1). Heredity, 82, 117–125.
Zaonarivelo, J. R., Sommer, J. A., Shore, G. E., McGuire, S. M., Engberg, S. E., et al. (2007). Isolation and characterization of 20 microsatellite marker loci from the Indri (Indri indri) genome. Molecular Ecology Notes, 7(1), 25–28.
Acknowledgments
We thank two anonymous reviewers for their constructive suggestions. We thank Madagascar National Parks (MNP) for permission to conduct this research, the Madagascar Institute for the Conservation of Tropical Ecosystems (MICET) and Madagascar Fauna Group (MFG) for logistical assistance in Madagascar, the Madagascar Biodiversity Project Field Team for field support, and the Duke Lemur Center for planning assistance. We thank Dr. Charles Faulkner (College of Veterinary and Comparative Medicine, Lincoln Memorial University) who has completed much of the parasitological identification over several years. We thank the team at Betampona Strict Nature Reserve, R. Dolch and the Mitsinjo Association, C. Williams, C. Welch, A. Katz, S. Zehr, F. Rasambainarivo, K. Freeman, G. Kett, B. Iambana, A. Junge, A. Greven, T. Rakotonanahary, H. Rafalinirina, and B. Allen for assistance in the field and in preparation. We also thank G. Stillings for help with ArcGIS. This work was supported by the St. Louis Zoo Field Research for Conservation Fund, the Duke University Center for International Studies, the Duke University Graduate School, the Nicholas School of the Environment, the Commonwealth of Kentucky, and the National Science Foundation (DEB-0949532). M. A. Barrett was a Robert Wood Johnson Foundation Health & Society Scholar at the University of California (UC) San Francisco and UC Berkeley; she thanks the program for its financial support.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling Editor: Joanna M. Setchell
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
Supporting information with priors used for MSVAR analysis (Table SI), and pairwise F ST 95% confidence interval values calculated from the full data set and a data set that excluded potential null alleles (Table SII) are available online. (DOCX 89 kb)
Rights and permissions
About this article
Cite this article
Nunziata, S.O., Wallenhorst, P., Barrett, M.A. et al. Population and Conservation Genetics in an Endangered Lemur, Indri indri, Across Three Forest Reserves in Madagascar. Int J Primatol 37, 688–702 (2016). https://doi.org/10.1007/s10764-016-9932-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10764-016-9932-y