Abstract
Activity cliffs have large impact in drug discovery; therefore, their detection and quantification are of major importance. This work introduces the metric activity cliff enrichment factor and expands the previously reported activity cliff generator concept by adding chemotype information to representations of the activity landscape. To exemplify these concepts, three molecular databases with multiple biological activities were characterized. Compounds in each database were grouped into chemotype classes. Then, pairwise comparisons of structure similarities and activity differences were calculated for each compound and used to construct chemotype-based structure–activity similarity (SAS) maps. Different landscape distributions among four major regions of the SAS maps were observed for different subsets of molecules grouped in chemotypes. Based on this observation, the activity cliff enrichment factor was calculated to numerically detect chemotypes enriched in activity cliffs. Several chemotype classes were detected having major proportion of activity cliffs than the entire database. In addition, some chemotype classes comprising compounds with smooth structure activity relationships (SAR) were detected. Finally, the activity cliff generator concept was applied to compounds grouped in chemotypes to extract valuable SAR information.
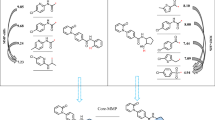
Graphic abstract









Similar content being viewed by others
Abbreviations
- ACEF:
-
Activity cliff enrichment factor
- COX:
-
Cyclooxygenase
- DAT:
-
Dopamine transporter
- ECFP:
-
Extended connectivity fingerprint
- EstateIndices:
-
Electrotopological state indices
- MACCS:
-
Molecular ACCess System
- MATs:
-
Monoamine transporters
- MEQI:
-
Molecular Equivalence Indices
- MEQNUM:
-
Molecular equivalence number
- NAC/CF:
-
Number of activity cliffs / chemotype frequency
- NET:
-
Norepinephrine transporter
- NSGs:
-
Network-like similarity graphs
- PPAR:
-
Peroxisome proliferator-activated receptor
- QSAR:
-
Quantitative structure–activity relationships
- ROCS:
-
Rapid overlay of chemical structures
- SALI:
-
Structure–activity landscape index
- SARI:
-
SAR index
- SAS:
-
Structure–activity similarity
- SERT:
-
Serotonin transporter
- SAR:
-
Structure–activity relationships
- TopAtomPairs:
-
Topological atom pairs
- TopPh4AtomPairs:
-
Topological pharmacophore atom pairs
- TopAtomTorsions:
-
Topological atom torsions
- TopAtomTriplets:
-
Topological atom triplets
- TopPh4AtomTriplets:
-
Topological pharmacophore atom triplets
References
Maggiora GM (2006) On outliers and activity cliffs: why QSAR often disappoints. J Chem Inf Model 46:1535. doi:10.1021/ci060117s
Cruz-Monteagudo M, Medina-Franco JL, Pérez-Castillo Y, Nicolotti O, Cordeiro MNDS, Borges F (2014) Activity cliffs in drug discovery: Dr Jekyll or Mr Hyde? Drug Discov Today 19:1069–1080. doi:10.1016/j.drudis.2014.02.003
Stumpfe D, Bajorath J (2012) Exploring activity cliffs in medicinal chemistry. J Med Chem 55:2932–2942. doi:10.1021/jm201706b
Pérez-Villanueva J, Medina-Franco JL, Caulfield TR, Hernández-Campos A, Hernández-Luis F, Yépez-Mulia L, Castillo R (2011) Comparative molecular field analysis (CoMFA) and comparative molecular similarity indices analysis (CoMSIA) of some benzimidazole derivatives with trichomonicidal activity. Eur J Med Chem 46:3499–3508. doi:10.1016/j.ejmech.2011.05.016
Hernández-Vázquez E, Méndez-Lucio O, Hernández-Luis F (2013) Activity landscape analysis, CoMFA and CoMSIA studies of pyrazole CB1 antagonists. Med Chem Res 22:4133–4145. doi:10.1007/s00044-012-0418-y
Iyer P, Wawer M, Bajorath J (2011) Comparison of two- and three-dimensional activity landscape representations for different compound data sets. Med Chem Commun 2:113–118. doi:10.1039/c0md00188k
Shanmugasundaram V, Maggiora GM (2001) Characterizing property and activity landscapes using an information-theoretic approach. Cinf-032. In: 222nd ACS national meeting, Chicago. American Chemical Society, Washington, D. C
Wawer M, Peltason L, Weskamp N, Teckentrup A, Bajorath J (2008) Structure–activity relationship anatomy by network-like similarity graphs and local structure–activity relationship indices. J Med Chem 51:6075–6084. doi:10.1021/jm800867g
Guha R, Van Drie JH (2008) Structure–activity landscape index: identifying and quantifying activity cliffs. J Chem Inf Model 48:646–658. doi:10.1021/ci7004093
Peltason L, Bajorath J (2007) SAR index: quantifying the nature of structure–activity relationships. J Med Chem 50:5571–5578. doi:10.1021/jm0705713
Bajorath J, Peltason L, Wawer M, Guha R, Lajiness MS, Van Drie JH (2009) Navigating structure–activity landscapes. Drug Discov Today 14:698–705. doi:10.1016/j.drudis.2009.04.003
Kayastha S, Dimova D, Iyer P, Vogt M, Bajorath J (2014) Large-scale assessment of activity landscape feature probabilities of bioactive compounds. J Chem Inf Model 54:442–450. doi:10.1021/ci400677b
Méndez-Lucio O, Pérez-Villanueva J, Castillo R, Medina-Franco JL (2012) Identifying activity cliff generators of PPAR ligands using SAS maps. Mol Inf 31:837–846. doi:10.1002/minf.201200078
Hu Y, Bajorath J (2012) Extending the activity cliff concept: structural categorization of activity cliffs and systematic identification of different types of cliffs in the ChEMBL database. J Chem Inf Model 52:1806–1811. doi:10.1021/ci300274c
Jayanthi LD, Ramamoorthy S (2005) Regulation of monoamine transporters: influence of psychostimulants and therapeutic antidepressants. AAPS J 7:E728–E738. doi:10.1208/aapsj070373
Torres GE, Gainetdinov RR, Caron MG (2003) Plasma membrane monoamine transporters: structure, regulation and function. Nat Rev Neurosci 4:13–25. doi:10.1038/nrn1008
Schneider C, Pozzi A (2011) Cyclooxygenases and lipoxygenases in cancer. Cancer Metastasis Rev 30:277–294. doi:10.1007/s10555-011-9310-3
Kirane A, Toombs JE, Ostapoff K, Carbon JG, Zaknoen S, Braunfeld J, Schwarz RE, Burrows FJ, Brekken RA (2012) Apricoxib, a novel inhibitor of COX-2, markedly improves standard therapy response in molecularly defined models of pancreatic cancer. Clin Cancer Res 18:5031–5042. doi:10.1158/1078-0432.CCR-12-0453
Moller DE (2001) New drug targets for type 2 diabetes and the metabolic syndrome. Nature 414:821–827. doi:10.1038/414821a
Balakumar P, Rose M, Ganti SS, Krishan P, Singh M (2007) PPAR dual agonists: are they opening pandora’s box? Pharmacol Res 56:91–98. doi:10.1016/j.phrs.2007.03.002
Méndez-Lucio O, Pérez-Villanueva J, Castillo R, Medina-Franco JL (2012) Activity landscape modeling of PPAR ligands with dual-activity difference maps. Bioorg Med Chem 20:3523–3532. doi:10.1016/j.bmc.2012.04.005
Dimova D, Wawer M, Wassermann AM, Bajorath J (2011) Design of multitarget activity landscapes that capture hierarchical activity cliff distributions. J Chem Inf Model 51:258–266. doi:10.1021/ci100477m
Pérez-Villanueva J, Medina-Franco JL, Méndez-Lucio O, Yoo J, Soria-Arteche O, Izquierdo T, Lozada MC, Castillo R (2012) CASE plots for the chemotype-based activity and selectivity analysis: a CASE study of cyclooxygenase inhibitors. Chem Biol Drug Des 80:752–762. doi:10.1111/cbdd.12019
Medina-Franco JL, Yongye AB, Pérez-Villanueva J, Houghten RA, Martínez-Mayorga K (2011) Multitarget structure–activity relationships characterized by activity-difference maps and consensus similarity measure. J Chem Inf Model 51:2427–2439. doi:10.1021/ci200281v
Chen X, Lin Y, Gilson MK (2001) The binding database: overview and user’s guide. Biopolymers 61:127–141. doi:10.1002/1097-0282(2002)61:2lt127:AID-BIP10076>3.0.CO;2-N
Chen X, Lin Y, Liu M, Gilson MK (2002) The binding database: data management and interface design. Bioinformatics 18:130–139. doi:10.1093/bioinformatics/18.1.130
Liu T, Lin Y, Wen X, Jorissen RN, Gilson MK (2007) BindingDB: a web-accessible database of experimentally determined protein–ligand binding affinities. Nucleic Acids Res 35:D198–D201. doi:10.1093/nar/gkl999
Xu YJ, Johnson M (2001) Algorithm for naming molecular equivalence classes represented by labeled pseudographs. J Chem Inf Comput Sci 41:181–185. doi:10.1021/ci0003911
Xu YJ, Johnson M (2002) Using molecular equivalence numbers to visually explore structural features that distinguish chemical libraries. J Chem Inf Comput Sci 42:912–926. doi:10.1021/ci025535l
Xu J (2002) A new approach to finding natural chemical structure classes. J Med Chem 45:5311–5320. doi:10.1021/jm010520k
Xu J, Gu Q, Liu H, Zhou J, Bu X, Huang Z, Lu G, Li D, Wei D, Wang L, Gu L (2013) Chemomics and drug innovation. Sci China Chem 56:71–85. doi:10.1007/s11426-012-4761-0
Gu Q, Yan X, Xu J (2013) Drug discovery inspired by mother nature: seeking natural biochemotypes and the natural assembly rules of the biochemome. J Pharm Pharm Sci 16:331–341
Sud M (2012) MayaChemTools: an open source package for computational discovery. Comp-306, In 243nd ACS National Meeting, San Diego. American Chemical Society, Washington, D. C
ROCS 3.1.0. OpenEye Scientific Software, Santa Fe. http://www.eyesopen.com
Jaccard P (1901) Étude comparative de la distribution florale dans une portion des Alpes et des Jura. Bull Soc Vaudoise Sci Nat 37:547–549
Willett P, Barnard JM, Downs GM (1998) Chemical similarity searching. J Chem Inf Comput Sci 38:983–996. doi:10.1021/ci9800211
Filimonov D, Poroikov V, Borodina Y, Gloriozova T (1999) Chemical similarity assessment through multilevel neighborhoods of atoms: definition and comparison with the other descriptors. J Chem Inf Comput Sci 39:666–670. doi:10.1021/ci980335o
Hall LH, Kier LB (1995) Electrotopological state indices for atom types: a novel combination of electronic, topological, and valence state information. J Chem Inf Comput Sci 35:1039–1045. doi:10.1021/ci00028a014
Rogers D, Hahn M (2010) Extended-connectivity fingerprints. J Chem Inf Model 50:742–754. doi:10.1021/ci100050t
Durant JL, Leland BA, Henry DR, Nourse JG (2002) Reoptimization of MDL keys for use in drug discovery. J Chem Inf Comput Sci 42:1273–1280. doi:10.1021/ci010132r
Carhart RE, Smith DH, Venkataraghavan R (1985) Atom pairs as molecular features in structure–activity studies: definition and applications. J Chem Inf Comput Sci 25:64–73. doi:10.1021/ci00046a002
Nilakantan R, Bauman N, Dixon JS, Venkataraghavan R (1987) Topological torsion: a new molecular descriptor for SAR applications. Comparison with other descriptors. J Chem Inf Comput Sci 27:82–85. doi:10.1021/ci00054a008
Renner S, Fechner U, Schneider G (2006) In pharmacophores and pharmacophore searches, vol 32. Wiley-VCH, Weinheim
Bonachéra F, Parent B, Barbosa F, Froloff N, Horvath D (2006) Fuzzy tricentric pharmacophore fingerprints. 1. Topological fuzzy pharmacophore triplets and adapted molecular similarity scoring schemes. J Chem Inf Model 46:2457–2477. doi:10.1021/ci6002416
Chang CE, Gilson MK (2003) Tork: conformational analysis method for molecules and complexes. J Comput Chem 24:1987–1998. doi:10.1002/jcc.10325
Vconf v2.0. VeraChem LLC, Germantown 2004. http://www.verachem.com
Yongye AB, Byler K, Santos R, Martínez-Mayorga K, Maggiora GM, Medina-Franco JL (2011) Consensus models of activity landscapes with multiple chemical, conformer, and property representations. J Chem Inf Model 51:1259–1270. doi:10.1021/ci200081k
Rush TS III, Grant JA, Mosyak L, Nicholls A (2005) A shape-based 3D scaffold hopping method and its application to a bacterial protein–protein interaction. J Med Chem 48:1489–1495. doi:10.1021/jm040163o
Sykes MJ, Sorich MJ, Miners JO (2006) Molecular modeling approaches for the prediction of the nonspecific binding of drugs to hepatic microsomes. J Chem Inf Model 46:2661–2673. doi:10.1021/ci600221h
Medina-Franco JL, Martínez-Mayorga K, Bender A, Marín RM, Giulianotti MA, Pinilla C, Houghtent RA (2009) Characterization of activity landscapes using 2D and 3D similarity methods: consensus activity cliffs. J Chem Inf Model 49:477–491. doi:10.1021/ci800379q
Chen B, Mueller C, Willett P (2010) Combination rules for group fusion in similarity-based virtual screening. Mol Inf 29:533–541. doi:10.1002/minf.201000050
Medina-Franco JL (2012) Scanning structure–activity relationships with structure–activity similarity and related maps: from consensus activity cliffs to selectivity switches. J Chem Inf Model 52:2485–2493. doi:10.1021/ci300362x
Pérez-Villanueva J, Santos R, Hernández-Campos A, Giulianotti MA, Castillo R, Medina-Franco JL (2011) Structure–activity relationships of benzimidazole derivatives as antiparasitic agents: dual activity-difference (DAD) maps. Med Chem Commun 2:44–49. doi:10.1039/c0md00159g
Medina-Franco JL, Petit J, Maggiora GM (2006) Hierarchical strategy for identifying active chemotype classes in compound databases. Chem Biol Drug Des 67:395–408. doi:10.1111/j.1747-0285.2006.00397.x
Pérez-Villanueva J, Santos R, Hernández-Campos A, Giulianotti MA, Castillo R, Medina-Franco JL (2010) Towards a systematic characterization of the antiprotozoal activity landscape of benzimidazole derivatives. Bioorg Med Chem 18:7380–7391. doi:10.1016/j.bmc.2010.09.019
Acknowledgments
The authors would like to express their sincere gratitude to the BindingDB team for providing the studied structure and activity data; to Dr. Mark Johnson for providing the program MEQI; to MayaChemTools for providing the scripts for fingerprint calculations; to VeraChem LLC for providing VConf; to OpenEye Scientific Software, Inc., for providing ROCS (UAM); and to Tableau Software for providing Tableau Public. O. M-L is very grateful to CONACyT (No. 217442/312933) and the Cambridge Overseas Trust for funding. JL. M-F thanks the National Autonomous University of Mexico (UNAM), grant PAIP 5000-9163, for funding.
Author information
Authors and Affiliations
Corresponding author
Additional information
This paper is dedicated to the Memory of Dra. Maria Concepción Lozada García.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pérez-Villanueva, J., Méndez-Lucio, O., Soria-Arteche, O. et al. Activity cliffs and activity cliff generators based on chemotype-related activity landscapes. Mol Divers 19, 1021–1035 (2015). https://doi.org/10.1007/s11030-015-9609-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11030-015-9609-z