Abstract
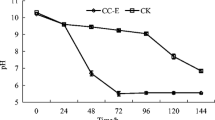
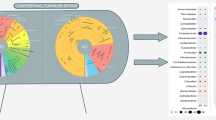
Bacterial community succession in the start-up of a large-scale, completely-mixed composting reactor was analyzed by 16S rRNA gene (16S rDNA) clone analysis and denaturing gradient gel electrophoresis (DGGE) combined with measurements of temperature, pH, moisture contents, and decomposing rate. DGGE analysis and physicochemical parameters showed that bacterial community succession occurred in four phases; (1) at the start of operation and pH decreasing period (day 0–3), (2) pH decreased and increased period (day 4–11), (3) middle term, moisture content decreasing and maximum temperature increased period (day 12–16) and (4) latter term, temperature decreasing period (day 17–24). Lactobacillus spp. and Bacillus coagulans were detected from the initial phase and middle term, respectively. 16S rDNA clone analysis showed that the dominant bacteria shifted from the order “Lactobacillales” to Bacillales and Actinomycetales. The order “Lactobacillales” was unique which may be caused by using the plastic bottle flakes (polyethylene terephthalate, PET) as bulking agent.



Similar content being viewed by others
Abbreviations
- BLAST:
-
Basic local alignment search tool
- DGGE:
-
Denaturing gradient gel electrophoresis
- rDNA:
-
rRNA gene
- r.p.m.:
-
Revolutions per minute
- VS:
-
Volatile substrate
References
Altschul SF, Gish W, Miller W, Meyers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Beffa T, Blank M, Lyon PF, Vogt G, Marchiani M, Fischer JL, Aragano M (1996) Isolation of Thermus strains from hot composts (60 to 80°C). Appl Environ Microbiol 62:1723–1727
Blanc M, Marilley L, Beffa T, Aragno M (1999) Thermophilic bacterial communities in hot composts as revealed by most probable number counts and molecular (16S rDNA) methods. FEMS Microbiol Ecol 28:141–149. doi:10.1111/j.1574-6941.1999.tb00569.x
Danon M, Franke-Whittle IH, Insam H, Chen Y, Hadar Y (2008) Molecular analysis of bacterial community succession during prolonged compost curing. FEMS Microbiol Ecol 65:133–144. doi:10.1111/j.1574-6941.2008.00506.x
Dees P, Ghiorse W (2001) Microbial diversity in hot synthetic compost as revealed by PCR-amplified rRNA sequences from cultivated isolates and extracted DNA. FEMS Microbiol Ecol 35:207–216. doi:10.1111/j.1574-6941.2001.tb00805.x
Ellis RJ, Morgan P, Weightman AJ, Fry JC (2003) Cultivation-dependent and -independent approaches for determining bacterial diversity in heavy-metal-contaminated soil. Appl Environ Microbiol 69:3223–3230. doi:10.1128/AEM.69.6.3223-3230.2003
Franke-Whittle IH, Klammer SH, Insam H (2005) Design and application of an oligonucleotide microarray for the investigation of compost microbial communities. J Microbiol Methods 62:37–56. doi:10.1016/j.mimet.2005.01.008
Green SJ, Michel JFC, Hader Y, Minz D (2004) Similarity of bacterial communities in sawdust- and straw-amended cow manure composts. FEMS Microbiol Lett 233:115–123. doi:10.1016/j.femsle.2004.01.049
Haruta S, Kondo M, Nakamura K, Aiba H, Ueno S, Ishii M, Igarashi Y (2002) Microbial community changes during organic solid waste treatment analyzed by double gradient-denaturing gradient gel electrophoresis and fluorescence in situ hybridization. Appl Microbiol Biotechnol 60:224–231. doi:10.1007/s00253-002-1074-9
Hemmi H, Shimoyama T, Nakayama T, Hoshi K, Nishino T (2004) Molecular biological analysis of microflora in a garbage treatment process under thermoacidophilic conditions. J Biosci Bioeng 97:119–126
Ishii K, Takii S (2003) Comparison of microbial communities in four different composting processes as evaluated by denaturing gradient gel electrophoresis analysis. J Appl Microbiol 95:109–119. doi:10.1046/j.1365-2672.2003.01949.x
Ishii K, Fukui M, Takii S (2000) Microbial succession during a composting process as evaluated by denaturing gradient gel electrophoresis analysis. J Appl Microbiol 89:768–777. doi:10.1046/j.1365-2672.2000.01177.x
Ivors K, Beyer D, Wuest P, Kang S (2002) Survey of fungal diversity in mushroom compost using sequences of PCR-amplified genes encoding 18S ribosomal RNA. In: Insam H, Riddech K, Klammer S (eds) Microbiology of composting. Springer, Berlin, pp 17–24
Kurosawa N, Itoh YH, Itoh T (2005) Thermus kawarayensis sp. nov., a new member of the genus Thermus, isolated from Japanese hot springs. Extremophiles 9:81–84. doi:10.1007/s00792-004-0419-y
Muyzer G, De Waal EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
Nagao N, Osa S, Matsuyama T, Namiki I, Yamamoto H, Toda T (2005) Optimization of washing rate in a hybrid recycling system of solid state and submerged fermentation. Process Biochem 40:3321–3326. doi:10.1016/j.procbio.2005.03.030
Nagao N, Watanabe K, Osa S, Matsuyama T, Kurosawa N, Toda T (2008) Bacterial community and decomposition rate in long term fed-batch composting using woodchip and polyethylene terephthalate (PET) as bulking agents. World J Microbiol Biotechnol 24:1417–1424. doi:10.1007/s11274-007-9625-y
Nakamura K, Haruta S, Nguyen HL, Ishii M, Igarashi Y (2004) Enzyme production-based approach for determinutesing the functions of microorganisms within a community. Appl Environ Microbiol 70:3329–3337. doi:10.1128/AEM.70.6.3329-3337.2004
Narihiro T, Abe T, Yamanaka Y, Hiraishi A (2004a) Microbial population dynamics during fed-batch operation of commercially available garbage composters. Appl Microbiol Biotechnol 65:488–495. doi:10.1007/s00253-004-1629-z
Narihiro T, Takebayashi S, Hiraishi A (2004b) Activity and phylogenetic composition of proteolytic bacteria in mesophilic fed-batch garbage composters. Microbes Environ 19:292–300. doi:10.1264/jsme2.19.292
Pedro MS, Haruta S, Hazaka M, Shimada R, Yoshida C, Hiura K, Ishii M, Igarashi Y (2001) Denaturing gradient gel electrophoresis analyses of microbial community from field-scale composter. J Biosci Bioeng 91:159–165. doi:10.1263/jbb.91.159
Pedro MS, Haruta S, Nakamura K, Hazaka M, Ishii M, Igarashi Y (2003) Isolation and characterization of predominant microorganisms during decomposition of waste materials in a field-scale composter. J Biosci Bioeng 95:368–373
Prescott LM, Harley JP, Klein DA (2002) Microbiology, 5th edn. McGraw-Hill, New York
Sakai K, Ezaki Y (2006) Open l-Lactic acid fermentation of food refuse using thermophilic Bacillus coagulans and fluorescence in situ hybridization analysis of microflora. J Biosci Bioeng 101:457–463. doi:10.1263/jbb.101.457
Sneath PHA, Mair NS, Sharpe ME, Holt JG (1986) Bergey’s manual of systematic bacteriology, vol 2. William & Wilkins, Baltimore
Strom PF (1985a) Identification of thermophilic bacteria in solid-waste composting. Appl Environ Microbiol 50:906–913
Strom PF (1985b) Effect of temperature on bacterial species diversity in thermophilic solid-waste composting. Appl Environ Microbiol 50:899–905
Acknowledgments
We thank Mr. Shinichiro Osa for technical assistance, Dr. Teruaki Yoshida for correcting English. This work was supported by the “University-Industry Joint Research” Project for Private Universities and a matching fund subsidy from MEXT (Ministry of Education, Culture, Sports, Science and Technology) of Japan, 2004–2008.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Watanabe, K., Nagao, N., Toda, T. et al. The dominant bacteria shifted from the order “Lactobacillales” to Bacillales and Actinomycetales during a start-up period of large-scale, completely-mixed composting reactor using plastic bottle flakes as bulking agent. World J Microbiol Biotechnol 25, 803–811 (2009). https://doi.org/10.1007/s11274-008-9952-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11274-008-9952-7