Abstract
For the in-situ nondestructive fat quantification of fresh tuna meat, an original lightweight (5.7 kg) hand-held sensor that consists of a planar radio-frequency coil and a single-sided magnetic circuit was developed as a subunit of a time-domain proton magnetic resonance (MR) scanner system. The investigation depth of the sensor unit is 12 mm, which is sufficient to probe the meat section beneath thick skin with scales and the underlying subcutaneous fat layer of large fish such as tuna. The scanner was successfully applied in a laboratory to a fillet of a bluefin tuna (Thunnus thynnus) to measure meat sections 12 mm beneath the skin. The required measurement time was 100 s for each section. The results of MR scan at 11 locations on the fillet were compared with those of conventional destructive food analysis. Reasonable agreement with an error (root-mean-square residual) of as small as 1.8 wt% was obtained for fat quantification. The time-domain MR relaxometry for the same tuna fillet also allowed lean meat quantification with a small root-mean-square residual of 6.7 wt%.
Similar content being viewed by others
Introduction
The fat content of tuna meat affects consumers’ sense of taste and ultimately determines price [1, 2]. Thus, it is desirable to nondestructively measure the fat content of fresh tuna using lightweight hand-held sensors at markets and sea-food processing plants [3,4,5,6,7,8] to accurately assess the value and price of whole fish. Portable surface scanners that employ low-field single-sided (i.e. unilateral) magnetic resonance (MR) relaxometry [9,10,11,12,13,14] are one of the most promising techniques for the in-situ nondestructive quantification of the fat content of fish. The advantages of MR scanning over other non-destructive methods, such as near-infrared spectroscopy, microwave attenuation, and electrical impedance [4, 15,16,17,18,19], are that (i) the sensed region is compact and its location is accurately known and (ii) the undesirable effects of bones and skin with scales can be eliminated if the investigation depth (distance from the sensor to the center of the sensed region) is carefully designed [20]. Veliyulin et al. [21] and Aursand et al. [22] successfully applied a single-sided MR surface scanner with an investigation depth of approximately 6 mm to the fat mapping of Atlantic salmon (Salmo salar), which has relatively thin skin. However, the investigation depth of this MR scanner with a hand-held sensor unit is too shallow to probe the meat sections beneath the thick skin and subcutaneous fat layers of bluefin tuna. Nakashima [20] measured bluefin tuna (Thunnus thynnus) using an MR surface scanner with an investigation depth of 30 mm, which is sufficient to probe meat sections beneath thick skin and subcutaneous fat layers. However, the sensor unit consisted of a planar radio-frequency (RF) coil and a large permanent magnetic circuit with an aluminum water-circulating disk, making it heavy (approximately 48 kg) and thus unsuitable for hand-held use.
In the present study, an original lightweight sensor unit (5.7 kg) is designed and constructed. Its investigation depth is 12 mm, which is sufficient to probe meat sections beneath thick skin with scales and subcutaneous fat layers. The detailed properties of the sensor unit, such as the spatial distribution of the magnetic field and MR sensitivity, and calibration results are examined. An MR surface scanner system with the proposed hand-held senor unit was applied to a fillet of a fresh bluefin tuna to perform one-dimensional fat mapping along a baseline. Reasonable agreement in terms of fat content was found between the nondestructive MR measurements and destructive conventional food analysis results. Time-domain MR scanning allows the simultaneous quantification of the lean meat content as well as fat content by utilizing the difference in the proton spin–spin relaxation times (T2). Successful measurements of lean meat at the locations used for fat quantification are also reported.
Methods and materials
Single-sided sensor unit
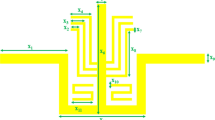
The original hand-held MR sensor unit developed in the present study is shown in Fig. 1a. It consists of a double-D-shaped planer RF coil with an aluminum tuning/matching (T/M) box (1.2 kg) and a single-sided magnetic circuit with handles (4.5 kg). The total weight is 5.7 kg. A double-D design [23] was employed for the coil because the direction of the RF magnetic field oscillating at the Larmor frequency should be normal to that of the static magnetic field in the sensed region. As a result, the direction of the RF magnetic field is in the y-direction while that of the static field is in the z-direction. Numerical simulations of the RF field [9, 12] were performed to determine the detailed geometry and dimensions of the planer RF coil. The design of the RF circuit is schematically shown in the Electronic Supplementary Material, Fig. ESM_1. The Larmor frequency tuning and 50 Ω matching operation are performed by nonmagnetic trimmer capacitors. The coil is divided into 4 units by four tuning capacitors to stabilize the T/M operation. During MR measurements, a glass-fiber-reinforced plastic (GFRP) plate (thickness: 1 mm) was placed on the RF coil shown in Fig. 1a to mechanically protect the coil and electromagnetically decouple the coil and samples (Fig. 1b).
Proposed hand-held sensor unit of the MR surface scanner system. The unit consists of permanent magnets with handles and an RF coil with a T/M box. a Photograph of the sensor unit. The RF coil is connected to the console via the BNC cable. The exact location of the origin of the x–y–z coordinate system is shown in Fig. ESM_3a. b A GFRP plate (thickness: 1 mm) was placed on the RF coil. The sensor unit weighs 5.7 kg
The single-sided (unilateral) magnetic circuit employs Nd-Fe-B permanent magnets, NEOMAX® (Hitachi Metals, Ltd., Tokyo, Japan). With the single-sided magnet geometry [9, 24], one side of the planar RF coil is exposed to free space (i.e. z > 0 in Fig. 1a), enabling the nondestructive surface scanning of large objects such as whole tuna. The magnetic circuit is axisymmetric and consists of an outer part (ring magnet) and an inner part (cylindrical magnet). The two magnets, which have the same polarity, are magnetized in the z-direction shown in Fig. 1a to produce strong and homogeneous magnetic fields far from their end surfaces. Computer simulations of the static magnetic fields [9, 11] were performed to determine the dimensions and relative position of the two magnets. The experimentally measured spatial distribution of the magnetic fields is shown in Fig. ESM_2. The dimensions and location of the sensed region of the sensor unit were measured using silicon rubber sheets [23, 25, 26] before the tuna measurements. The procedure and results are described in Fig. ESM_3. The investigation depth (distance from the RF coil to the center of the sensed region) was 12 mm and the dimensions of the sensed region were 30, 24, and 7 mm along the x-, y-, and z-axes, respectively. The RF coil and the magnetic circuit are carefully designed such that the layer within ≈8 mm from the coil surface does not significantly contribute to the MR signals (Fig. ESM_3b). Thus, MR signals from the subcutaneous fat layer and skin of fish are eliminated, leaving those from a region composed of only tuna meat away from the skin.
Sample description
A 7.1-kg fillet of a fresh bluefin tuna (Thunnus thynnus) farmed in Nagasaki, Japan, with skin and spine (Fig. 2a) was purchased from a local store. An X-ray computed tomography (CT) image of the fillet is shown in Fig. ESM_4. The fillet was left in the laboratory overnight to allow it to equilibrate to ambient room temperature (28 to 30 °C), and then the 11 locations marked in Fig. 2a were scanned by the MR sensor (Fig. 2b). An anatomical examination of the fillet performed after the MR measurements and the X-ray CT image analysis revealed that the total thickness of the skin with scales and the underlying subcutaneous fat layer ranged from approximately 4 mm (back) to 8 mm (abdomen). This confirms that the sensed region did not overlap with the skin and the subcutaneous fat layer, and that reliable quantification of meat sections was performed.
Photograph of a fillet of bluefin tuna measured in the present study. a Top view of the fillet (weight: 7.1 kg). Eleven locations on a dotted baseline (L1 to L11, separated by 3 cm along the curved fish body) scanned by the MR sensor are marked. An X-ray CT image taken along the dotted line is shown in Fig. ESM_4. b Location L6 being scanned with the proposed sensor unit. A wider-angle photograph of this scene is shown in Fig. ESM_5
The number of measured locations in Fig. 2a (i.e. 11) may be insufficient for evaluating the performance of the developed MR sensor. Thus, additional samples, namely (i) and (ii) below, whose meat quality ranged broadly from lean meat to fatty meat, were prepared and measured by the same MR scanner. These additional samples were all boneless meat block samples, typically 100 × 100 × 50 mm3, large enough to completely cover the sensed region (30 × 24 × 7 mm3). Each sample was vacuum-sealed in a thin plastic film package to prevent undesired water evaporation and fat alteration during the MR experiments. (i) After the MR scanning of the 11 locations shown in Fig. 2a, four samples were collected from sections located deeper than the sensed region shown in Fig. ESM_4 (typically 10 cm beneath the skin of the fillet). A total of four locations were measured for the vacuum-sealed meat blocks without skin. The T2 value of the leanest meat sample (i.e. that with the least fat content) among the four was assumed to be the T2 value of pure lean meat as an end member. (ii) Fresh tuna meat blocks from a different bluefin tuna fish farmed in Kagoshima, Japan, were purchased from a local store. Twelve sections of the vacuum-sealed blocks without skin were measured by the MR scanner. A total of 27 samples (11 + 4 + 12) were thus measured to estimate the fat and lean content in the present study.
Furthermore, an almost pure subcutaneous fat sample without muscle was collected from near the abdomen of the fillet shown in Fig. 2a. Because the fat sample was small in volume compared with the sensed region, the MR signal amplitude was not plotted in the calibration diagram. Instead, the MR data were used to determine the T2 value of pure fat as an end member, as described in the next section.
MR scanning of tuna samples
The transient transverse relaxation of protons (1H nuclei) in the tuna samples was measured using the MR console system (MRTechnology Inc., Tsukuba, Japan) shown in Fig. ESM_5. The target nuclei were the protons of water molecules in muscle and those of fat molecules. The proton Larmor frequency corresponding to the magnetic flux density of the sensed region (Fig. ESM_2) was 4.3 MHz. The center (i.e. x = y = 0) of the RF coil shown in Fig. 1 was placed on one of the 11 locations marked in Fig. 2a and adjusted for T/M by the capacitors shown in Fig. ESM_1 using the Smith chart displayed on a network analyzer, and then the transient proton relaxation was measured. The resultant quality factor Q of the RF coil after the T/M adjustment ranged from 18 to 19 depending on the meat section. The RF shield cloth (Fig. 2b) was connected to the outer shield of the BNC cable (Fig. ESM_1) to reduce environmental RF noise.
A phase-alternated pair stacking (PAPS) Carr–Purcell–Meiboom–Gill (CPMG) pulse sequence [27] was employed to acquire the proton transverse relaxation data. The parameters of the CPMG sequence were as follows. The duration of the 90° and 180° pulses was 0.2 ms, the echo spacing was 0.5 ms, the sequence repetition time was 2 s (i.e., the T1 full relaxation condition, where T1 refers to the spin–lattice relaxation time), and the number of echoes was 1100. Two values (50 and 100) were employed for the number of stacked signals, Nstack, to evaluate the effect of Nstack on measurement accuracy. Thus, the required MR measurement times for each location were 2 s × 50 times = 100 s and 2 s × 100 times = 200 s, respectively.
The principle of the CPMG data processing for the fat and lean meat quantification is described here based on T2 relaxometry. The raw time-series data obtained using the CPMG pulse sequence consisted of 1100 echo data points. The first four echoes were discarded because the initial echoes are inevitably transient and distorted due to the grossly inhomogeneous magnetic and RF fields [28]. The time-series data were then divided by Q0.5; as stated previously, Q varied in the range of 18 ≤ Q ≤ 19 among the measured samples. This quality factor correction was needed because the magnitude of the CPMG signal increases with Q0.5 [29, 30] and thus must be scaled by Q0.5 to quantitatively compare CPMG datasets obtained in cases with different Q values.
As a final step, it was assumed that boneless fatty meat is a mechanical mixture of lean meat (water + protein) and fat. This assumption has been validated by previous studies on beef and tuna meat [20, 25]. Although it is difficult to obtain proton relaxation signals from protein macromolecules using low-field time-domain MR, they can be readily obtained from small water and fat molecules with relatively high mobility. Thus, the time-series data after the Q-value correction mentioned above, f(t), were a mixture of proton relaxation signals from the two end members, fat and water, written as
where t is time, Afat and Alean are the MR signal amplitudes for fat molecules and water molecules in lean meat, respectively, and T2fat and T2lean are the T2 values for fat and lean meat, respectively. The values of Afat and Alean were determined from the bi-exponential model, Eq. 1, using the method of least squares, and the constants T2fat and T2lean were determined using the following method.
CPMG data for the two end members mentioned in the previous section (lean meat with negligible fat content sampled from a deep section of the fillet, and almost pure subcutaneous fat without muscle collected from near the abdomen of the fillet) are shown in Fig. 3 for Nstack = 100 and in Fig. ESM_6 for Nstack = 50. A mono-exponential model assuming a single T2 value was fitted to the CPMG data using the method of least squares. As a result, T2fat and T2lean were determined to be 143 and 49 ms for Nstack = 100, and 148 and 50 ms for Nstack = 50, respectively. It should be noted that T2fat and T2lean are significantly different (≈threefold difference). Similar results have been reported for salmon [21] and beef [25]. This large difference ensures that the fat and lean meat contained in the samples can be differentiated with reasonable accuracy.
Example of the CPMG time-series data, f(t), for three portions (subcutaneous fat, lean meat, and fatty meat) of a single fish. Nstack = 100. According to the conventional food analysis, the fat content and water content were 72.1 and 16.6 wt% for the subcutaneous fat sample, and 2.3 and 71.3 wt% for the lean meat sample, respectively. The bi-exponential model, Eq. (1), was fitted to the fatty meat data, which are for location L7 in Fig. 2a, by the least squares method using a solid line. A mono-exponential model assuming a single T2 value was fitted to the subcutaneous fat and lean meat samples by solid lines; T2 values of 143 and 49 ms, respectively, were obtained
After the MR measurements, a small piece of meat with dimensions of 40, 30, and 15 mm along the x-, y-, and z-axes, respectively, was cut from each section using a knife, with care taken to completely include the sensed region (30 × 24 × 7 mm3). The fat and water content of these tuna meat pieces was then analyzed using conventional food chemical analysis techniques to determine the true values. The fat and water were quantified using the Soxhlet extraction method and the air oven method, respectively. The analytical techniques followed the Standard Tables of Food Composition in Japan 2015 (7th Ed.) determined by the Ministry of Education, Culture, Sports, Science and Technology, Japan. Other proximate compositions (ash and protein) were not quantified by the conventional food chemical analysis in the present study. This is because the ash content is negligible (approximately 1 wt%) for boneless fish meat [31, 32] and difficult to measure by the low-field proton MR scanning, and the protein content correlates well with the water content as follows.
For lean meat, which consists of water and protein, the weight ratio of protein to water is constant [20, 25, 33]. Thus, it is possible to convert the water content value obtained using the air oven method into lean meat content. Employing 72.6 wt% [33] as the water fraction, the lean meat content was calculated in the present study by multiplying a constant of 1/0.726 ≈ 1.38 by the water content value. The results for the tuna meat analysis by the conventional food chemical analysis are shown in Table 1 and Tables ESM_1 and ESM_2. The sum of the fat and lean meat content is almost 100 wt%, suggesting that the conversion method of water content into lean meat content is acceptable.
Results
An example of the CPMG data for a fatty section of meat (location L7 in Fig. 2a) is shown in Fig. 3 for Nstack = 100 and in Fig. ESM_6 for Nstack = 50. Although the noise level is higher for Nstack = 50 compared with that for Nstack = 100, the degree of fitting with Eq. (1) is reasonable for both cases. The apparent decay rate for L7 falls between those for the subcutaneous fat and lean meat samples, which is a consequence of the bi-exponential model, Eq. (1), where fatty meat is a mechanical mixture of pure fat and lean meat.
Unknown quantities Afat and Alean for 27 sections of the tuna samples were determined by fitting the bi-exponential model, Eq. (1), with known T2fat and T2lean values to the CPMG relaxation data. These determined quantities are plotted against the conventional food analytical results in Fig. 4 for Nstack = 100 and in Fig. ESM_7 for Nstack = 50. The 27 data points were fitted to theoretical calibration curves, Eq. (2) for fat and Eq. (3) for lean meat, written as
and
where Bfat and Blean are constants, wfat and wlean are the weight fractions of fat and lean meat (in wt%), respectively, and ρfat and ρlean are the bulk densities of fat and lean meat, respectively. Based on the results of Kraybill et al. [33], ρfat and ρlean were taken to be 0.92 and 1.06 g/cm3, respectively, in the present study. Figure 4 and Fig. ESM_7 show that Eqs. (2) and (3) are not exactly linear but slightly curved with respect to the weight fraction of fat and lean meat. This is a consequence of the bulk density of fat being different from that of lean meat. Thus, it should be noted that the linear calibration operation employed in previous studies [20, 25] are not exactly correct.
Results of the single-sided MR measurements of fresh bluefin tuna for Nstack = 100. For the tuna farmed in Nagasaki, 11 shallow sections beneath the skin (L1 to L11 in Fig. 2a) and 4 packed meat blocks without skin sampled from deeper sections were measured. Twelve packed meat blocks without skin sampled from a different tuna (farmed in Kagoshima) were also measured. a Afat of Eq. (1) plotted against the fat content obtained by the conventional food analysis, and fitted by Eq. (2). The horizontal axis is wfat in Eq. (2). b Alean of Eq. (1) plotted against the lean content obtained by the conventional food analysis, and fitted by Eq. (3). The horizontal axis is wlean in Eq. (3)
The constants Bfat and Blean in Eqs. (2) and (3) were determined using the data points in Fig. 4 and Fig. ESM_7 by the method of least squares. Then, the vertical axes of Fig. 4 and Fig. ESM_7 were rewritten using the determined constants to obtain Fig. 5 and Fig. ESM_8, respectively. The root-mean-square residuals (RMSRs) for the 11 data points (L1 to L11) shown in Fig. 2a were 2.0 wt% for fat in Fig. 5a and 5.8 wt% for lean meat in Fig. 5b (Nstack = 100); they were 1.8 wt% for fat in Fig. ESM_8a and 6.7 wt% for lean meat in Fig. ESM_8b (Nstack = 50). The sum of the fat and lean content for each sample was calculated using Fig. 5 and Fig. ESM_8. The results are plotted in Fig. 6. The numerical data set for the eleven points (L1 to L11) shown in Fig. 2a is summarized in Table 1; that for the other data points shown in Figs. 5, 6 and Fig. ESM_8 is summarized in Tables ESM_1 and ESM_2.
Cross-plot of the a fat and b lean content for the 27 samples in Fig. 4 measured by MR scanning (Nstack = 100) and by conventional food analytical methods. Three error contours, corresponding to 0 and ± 5 wt%, are indicated by solid and dotted lines, respectively. The RMSRs of the 27 data points for the fat and lean contents were 2.0 and 5.6 wt%, respectively
Cross-plot of the total meat amount (fat + lean meat) measured by single-sided MR scanning and by conventional food analytical methods. Data were collected from Fig. 5 and Fig. ESM_8. The true value (100 wt%) is indicated by dotted lines. The arithmetic average and standard deviation of the 27 data points were 98.8 and 1.5 wt%, respectively, for the food analysis, and 99.8 and 6.3 wt% (Nstack = 50) and 99.8 and 5.5 wt% (Nstack = 100), respectively, for MR scanning
Discussion
The experimental results described in the previous section show that accurate fat quantification was performed for the fresh tuna samples using the proposed sensor unit. The RMSR for Nstack = 100 was 2.0 wt% for both the 11 locations of Fig. 2a and the 27 samples in Fig. 5a; that for Nstack = 50 was 1.8 wt% for the 11 locations of Fig. 2a and 1.9 wt% for the 27 samples in Fig. ESM_8a. These error levels are comparable to that obtained with near-infrared spectroscopy for yellowfin tuna without skin [34]. This reasonable agreement with respect to fat content between MR scanning and conventional food analysis demonstrates that MR scanning is a promising technique for the fat quantification of fresh tuna. The scanner system with the hand-held sensor unit (Fig. ESM_5) is portable and works with an AC 100-V power supply. Thus, this system is promising for the in-situ, accurate, rapid, and nondestructive fat quantification of large fish such as tuna with thick skin and subcutaneous fat layers at markets and sea-food processing plants [3,4,5,6,7,8].
Figures 5 and ESM_8 show that the scattering of the data points is almost independent of the fish farmed in Nagasaki or in Kagoshima. The independence from the individual with respect to the fat and lean meat quantification has been reported by the previous studies on the MR scanning of tuna [20] and beef [25] samples. This suggests that the employed bi-exponential model, Eq. (1), where fatty meat is a mechanical mixture of pure fat and lean meat, is reasonable.
The advantage of MR scanning over conventional nondestructive fat quantification methods should be noted. The measurement accuracy of conventional methods, such as near-infrared spectroscopy, microwave attenuation, and electrical impedance [4, 15,16,17,18], can be degraded by skin with scales and by bones, which can scatter, absorb, and distort near-infrared rays, microwaves, and electric currents. In contrast, even though 11 data points were for samples with skin and 16 data points were for those without skin in Fig. 4 and Fig. ESM_7, all 27 data points obey a single calibration curve, Eqs. (2) or (3). This negligible effect of skin on the MR data has been reported by a previous study [20]. This is a consequence of the dimensions of the sensed region being compact (30 × 24 × 7 mm3) and the lack of skin and bones in the sensed region, as shown in Fig. ESM_4. Because the MR data are not affected by skin and bones, the proposed MR scanning method has an advantage over other methods for characterizing whole fish. As a next possible research step, it is desirable to perform scanning of a whole tuna with skin by MR scanning and these conventional nondestructive methods to confirm the advantage of the MR scanning method mentioned above. Supplementary analysis by destructive methods for meat pieces employing infrared or visible-light spectroscopy and differential scanning calorimetry [35,36,37] would be helpful to the interpretation of the raw data obtained by the nondestructive methods.
Time-domain MR relaxometry based on Eq. (1) allowed us to successfully quantify lean content as well as fat content. The RMSR for Nstack = 100 was 5.8 wt% for the 11 locations shown in Fig. 2a and 5.6 wt% for the 27 samples in Fig. 5b; that for Nstack = 50 was 6.7 wt% for the 11 locations shown in Fig. 2a and 6.5 wt% for the 27 samples in Fig. ESM_8b. These values are slightly larger than the RMSRs for fat quantification. A possible reason for these larger RMSR values is that because protein in lean meat cannot be directly detected by the low-field MR relaxometry, the water content obtained by MR measurement was multiplied by 1.38 to estimate the lean meat quantity. This straightforward magnification of the water data could have magnified the error in the lean meat estimation.
The sum of fat and lean content should be 100 wt% for self-consistency of the MR method because there are no bones in the sensed region, as shown in Fig. ESM_4. The sum of fat and lean content obtained by MR scanning is plotted against that obtained by conventional food analysis in Fig. 6. The slightly larger standard deviation for the nondestructive MR scanning compared with that for the destructive food analytical method implies that the intrinsic measurement error is slightly larger for the MR scanning method. However, the standard deviation is as small as 6.3 wt% (Nstack = 50) and 5.5 wt% (Nstack = 100) in Fig. 6, demonstrating self-consistency and thus the reliability of MR scanning followed by data analysis using Eqs. (1)–(3).
Two values for Nstack (50 and 100) were examined in the present study to evaluate the effects of signal stacking on measurement accuracy. As shown in Figs. 5 and 6 and Fig. ESM_8, no significant difference was observed among the two cases. The required measurement times for each location were 2 s × 50 times = 100 s and 2 s × 100 times = 200 s, respectively. Thus, Nstack of 50 is recommended for reducing measurement time. If the RF shielding against environmental noise is improved, for example, by employing a shield tent [25], better CPMG data with a higher signal-to-noise ratio could be acquired, allowing the use of a lower Nstack value and thus lowering measurement time. Because the MR scanning method is noninvasive, lowered measurement time would enable to scan live fish at fish farms and to release them with negligible damage after scanning, which would be useful for monitoring the growth path in terms of the nutritional management.
The successful application of the single-sided MR scanner to tuna demonstrated in the present study suggests that the scanner would be promising for the fat quantification of different sea-foods (e.g. prawns [38] and crabs [39]). When performing MR scan of such fishery products at real markets and sea-food processing plants, however, special care to the sample temperature should be taken. The settings for the T2fat and T2lean values in Eq. (1) employed in the present study (i.e. 143 and 49 ms for Nstack = 100, and 148 and 50 ms for Nstack = 50) are not universal but valid only for the sample temperature of 28–30 °C. The values significantly depend on the sample temperature [40, 41]. The preliminary study on tuna meat (Fig. ESM_9) suggests that smaller T2fat and T2lean values should be employed for Eq. (1) when quantifying fat and lean meat of chilled sea-foods.
There are strong demands for the nondestructive measurement and characterization of packed products at fishery and non-fishery food factories [5,6,7, 42,43,44,45,46]. Low-field unilateral (i.e. single-sided) and bilateral MR scanning has been tried for such purposes [21, 22, 47,48,49,50,51]. Examples include animal meat [3, 4, 25, 40] and fruit [52,53,54,55,56]. Thus, the single-sided MR surface scanner developed (Fig. 1) with a reasonable investigation depth could be applicable to such foods, particularly those that require deeper investigation depths such as beef carcasses with thick subcutaneous fat layers and large fruits with thick skin.
Conclusions
An MR surface scanner based on proton relaxometry enables the nondestructive quantification of the fat content of meat by utilizing the difference in the spin–spin relaxation times of water molecules in lean meat and fat molecules. An original hand-held sensor unit was developed for the MR surface scanner to quantify the fat content of fresh tuna. The investigation depth is as deep as 12 mm and the dimensions of the sensed region are 30 × 24 × 7 mm3. The MR scanner was successfully applied in a laboratory to a fillet of a bluefin tuna to measure 11 meat sections 12 mm beneath the skin; the measurement time was 100 s for each section. The RMSR value, adopted as the measurement error, for the MR scanning was as small as 1.8 wt% when compared with the true values obtained by conventional destructive food analysis. The MR scanner also allows lean meat quantification with an RMSR value of 6.7 wt%. This portable MR scanner is a promising tool for the in-situ, rapid, accurate, and nondestructive quantification of fat and lean meat of tuna with thick skin and subcutaneous fat layers at markets and sea-food processing plants.
References
K.E. McConnell, I.E. Strand, Am. J. Agric. Econ. (2000). https://doi.org/10.1111/0002-9092.00011
T. Shimose, M. Kanaiwa, T. Tanabe, Fish Res. (2018). https://doi.org/10.1016/j.fishres.2018.03.001
M.F. Marcone, S. Wang, W. Albabish, S. Nie, D. Somnarain, A. Hill, Food Res. (2013). https://doi.org/10.1016/j.foodres.2012.12.046
F. Tao, M. Ngadi, Crit. Rev. Food Sci. Nutr. (2018). https://doi.org/10.1080/10408398.2016.1261332
Z. Xiaobo, H. Xiaowei, M. Povey, Analyst (2016). https://doi.org/10.1039/c5an02152a
S. Abasi, S. Minaei, B. Jamshidi, D. Fathi, Trends Food Sci. Technol. (2018). https://doi.org/10.1016/j.tifs.2018.05.009
H.S. El-Mesery, H. Mao, A.E.F. Abomohra, Sensors (2019). https://doi.org/10.3390/s19040846
K. Fan, M. Zhang, Crit. Rev. Food Sci. Nutr. (2019). https://doi.org/10.1080/10408398.2018.1441124
B. Blümich, J. Perlo, F. Casanova, Prog. Nucl. Magn. Reson. Spectrosc. 52, 197–269 (2008)
M. Greer, C. Chen, S. Mandal, J. Magn. Reson. (2019). https://doi.org/10.1016/j.jmr.2019.106591
A. Bashyam, M. Li, M.J. Cima, J. Magn. Reson. (2018). https://doi.org/10.1016/j.jmr.2018.05.004
N.P. Gaunkar, S. Utsuzawa, Y.Q. Song, M. Mina, D. Jiles, IEEE Trans. Magn. (2018). https://doi.org/10.1109/TMAG.2018.2845664
O. Sucre, C. Rondeau-Mouro, J. Magn. Reson. (2019). https://doi.org/10.1016/j.jmr.2019.106619
B. Blümich, in Advanced Characterization Techniques, Diagnostic Tools and Evaluation Methods in Heritage Science, ed. by D. Bastidas, E. Cano (Springer, Cham, 2018)
M. Kent, Food Control (1990). https://doi.org/10.1016/0956-7135(90)90121-R
J. Shimamoto, S. Hiratsuka, K. Hasegawa, M. Sato, S. Kawano, Fish Sci. (2003). https://doi.org/10.1046/j.1444-2906.2003.00698.x
K. Kubo, Y. Matsumoto, K. Kuwahara, S. Okabe, S. Taniyama, K. Tachibana, M. Murata, Nippon Suisan Gakkaishi 82, 743–752 (2016). (in Japanese with English abstract and figure captions)
D. Nielsen, G. Hyldig, J. Nielsen, H.H. Nielsen, LWT (2005). https://doi.org/10.1016/j.lwt.2004.07.010
J. Willis, A.J. Hobday, Fish Res. (2008). https://doi.org/10.1016/j.fishres.2008.02.010
Y. Nakashima, J. Aquat. Food Prod. Technol. (2019). https://doi.org/10.1080/10498850.2019.1569742
E. Veliyulin, C. van der Zwaag, W. Burk, U.W. Erikson, J. Sci. Food Agric. (2005). https://doi.org/10.1002/jsfa.2117
I.G. Aursand, E. Veliyulin, U. Erikson, in Modern Magnetic Resonance, ed. by G.A. Webb (Springer, Dordrecht, 2008)
S. Utsuzawa, E. Fukushima, J. Magn. Reson. (2017). https://doi.org/10.1016/j.jmr.2017.07.006
B. Manz, A. Coy, R. Dykstra, C.D. Eccles, M.W. Hunter, B.J. Parkinson, P.T. Callaghan, J. Magn. Reson. (2006). https://doi.org/10.1016/j.jmr.2006.07.017
Y. Nakashima, Appl. Magn. Reson. (2015). https://doi.org/10.1007/s00723-015-0657-4
Y. Nakashima, T. Sawatsubashi, S. Fujii, Int. J. Coal Prep. Util. (2020). https://doi.org/10.1080/19392699.2020.1722656
G.R. Coates, L. Xiao, M.G. Prammer, NMR Logging Principles and Applications (Gulf Professional Publishing, Houston, 2001)
M.D. Hürlimann, D.D. Griffin, J. Magn. Reson. (2000). https://doi.org/10.1006/jmre.1999.1967
F.D. Doty, R.R. Inners, P.D. Ellis, J. Magn. Reson. (1981). https://doi.org/10.1016/0022-2364(81)90051-2
D.I. Hoult, B. Bhakar, Concepts Magn. Reson. (1997). https://doi.org/10.1002/(SICI)1099-0534(1997)9:5%3C277:AID-CMR1%3E3.0.CO;2-W
M.I. Yeannes, M.E. Almandos, J. Food Comp. Anal. (2003). https://doi.org/10.1016/S0889-1575(02)00168-0
Y.N. Nakamura, M. Ando, M. Seoka, K.I. Kawasaki, Y. Tsukamasa, Fish Sci. (2005). https://doi.org/10.1111/j.1444-2906.2005.01004.x
H.F. Kraybill, H.L. Bitter, O.G. Hankins, J. Appl. Physiol. (1952). https://doi.org/10.1152/jappl.1952.4.7.575
K. Khodabux, M.S.S. L’Omelette, S. Jhaumeer-Laulloo, P. Ramasami, P. Rondeau, Food Chem. (2007). https://doi.org/10.1016/j.foodchem.2006.05.057
R.A. Mancini, M. Hunt, Meat Sci. (2005). https://doi.org/10.1016/j.meatsci.2005.03.003
M. Lucarini, A. Durazzo, J.S. Del Pulgar, P. Gabrielli, G. Lombardi-Boccia, Food Chem. (2018). https://doi.org/10.1016/j.foodchem.2017.11.042
T. Mihociu, M. Negoita, A. Culetu, Int. J. Innov. Approach Agric. Res. (2018). https://doi.org/10.29329/ijiaar.2018.174.1
M. Li, B. Li, W. Zhang, Food Chem. (2018). https://doi.org/10.1016/j.foodchem.2017.08.086
J.P. Wold, M. Kermit, A. Woll, Appl. Spectrosc. (2010). https://doi.org/10.1366/000370210791666273
E. Micklander, B. Peshlov, P.P. Purslow, S.B. Engelsen, Trends Food Sci Technol. (2002). https://doi.org/10.1016/S0924-2244(02)00163-2
Y. Nakashima, Japanese patent (application number 2020-015284) (2020).
S. Pirsa, S. Chavoshizadeh, Polym. Adv. Technol. (2018). https://doi.org/10.1002/pat.4250
S. Pirsa, M. Zandi, H. Almasi, J. Food Process Eng. (2016). https://doi.org/10.1111/jfpe.12219
S. Chavoshizadeh, S. Pirsa, F. Mohtarami, Eur. J. Lipid Sci. Tech. (2020). https://doi.org/10.1002/ejlt.201900385
S. Pirsa, I. Karimi Sani, S. Khodayvandi, Polym. Adv. Technol. (2018). https://doi.org/10.1002/pat.4397
S. Pirsa, T. Shamusi, Mater. Sci. Eng. C (2019). https://doi.org/10.1016/j.msec.2019.02.021
P.Ł. Kowalczewski, K. Walkowiak, Ł. Masewicz, H.M. Baranowska, Open Agric. (2019). https://doi.org/10.1515/opag-2019-0038
M. Gudjónsdóttir, R.J. Napitupulu, H.T. Petty Kristinsson, Magn. Reson. Chem. (2019). https://doi.org/10.1002/mrc.4855
E. Kirtil, S. Cikrikci, M.J. McCarthy, M.H. Oztop, Curr. Opin. Food Sci. (2017). https://doi.org/10.1016/j.cofs.2017.07.005
U. Erikson, I.B. Standal, I.G. Aursand, E. Veliyulin, M. Aursand, Magn. Reson. Chem. (2012). https://doi.org/10.1002/mrc.3825
M. Grootveld, B. Percival, M. Gibson, Y. Osman, M. Edgar, M. Molinari, M.L. Mather, F. Casanova, P.B. Wilson, Anal. Chim. Acta. (2019). https://doi.org/10.1016/j.aca.2019.02.026
D. Capitani, A.P. Sobolev, V. Di Tullio, L. Mannina, N. Proietti, Chem. Biol. Technol. Agric. (2017). https://doi.org/10.1186/s40538-017-0100-1
N. Proietti, G. Adiletta, P. Russo, R. Buonocore, L. Mannina, A. Crescitelli, D. Capitani, J. Food Eng. (2018). https://doi.org/10.1016/j.jfoodeng.2018.02.028
M.N. Martin, B.J. Balcom, M.J. McCarthy, A.P. Augustine, J. Food Sci. (2019). https://doi.org/10.1111/1750-3841.14778
C.S.W. Miaw, P.M. Santos, A.R.C.S. Silva, A. Gozzi, N.C.C. Guimarães, M.P. Callao, I. Ruisánchez, M.M. Sena, S.V.C. de Souza, Food Anal. Methods (2019). https://doi.org/10.1007/s12161-019-01522-7
M. Pinter, T. Harter, M. McCarthy, M. Augustine, Sensors (2014). https://doi.org/10.3390/s140304167
Acknowledgements
Comments by anonymous reviewers were helpful.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author is an inventor of a Japanese patent of Ref. [41].
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nakashima, Y. Development of a hand-held magnetic resonance sensor for the nondestructive quantification of fat and lean meat of fresh tuna. Food Measure 14, 2947–2955 (2020). https://doi.org/10.1007/s11694-020-00539-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11694-020-00539-5