Abstract
Waste water treatment plant (WWTP) is considered as an important source of surface water contamination by enteric pathogens. In this study, we describe the occurrence of enteric viruses (group A rotaviruses, noroviruses, astroviruses, sapoviruses, hepatitis A virus, and hepatitis E virus) and Clostridium difficile in the effluent of a wastewater treatment plant during a 1-year period. Enteric viruses were simultaneously and efficiently concentrated in a single step using methacrylate monolithic chromatographic support. Rotaviruses, noroviruses (genogroup I and II), and sapoviruses were detected in all 12 concentrated samples, whereas astroviruses were not detected in August and September and hepatitis A and E viruses were not detected at all. Clostridium difficile was detected in all samples and altogether 121 strains were isolated and grouped into 32 different ribotypes of which 014/020 and 010 were most prevalent. Pathogens detected in WWTP effluent partially reflect the epidemiological situation of enteric viruses and C. difficile in human population and open the discussion on implementation of possible techniques for virus and bacteria removal from WWTP effluent prior to release into the surface water system.
Similar content being viewed by others
Introduction
Enteric pathogens are associated with high public health burden throughout the world and most common representatives include rotaviruses, noroviruses, adenoviruses, Salmonella sp., pathogenic E. coli, Campylobacter spp., and Clostridium difficile (Kolling et al. 2012).
In general, the most important way of transmission for enteric pathogens is the fecal-oral route. However, after the development of new, sensitive, and reliable techniques in environmental microbiology, the importance of indirect transmission of bacterial, parasitic, and viral pathogens became recognized (Bosch et al. 2008; Bouzid et al. 2008). Contaminated water used for irrigation or food preparation can be a source of food contamination (Koopmans and Duizer 2004). Several studies support this theory showing the high prevalence of enteric viruses in surface and/or groundwater systems (Steyer et al. 2011b; Lodder and de Roda Husman 2005; Williamson et al. 2011). Enteric viruses enter the surface- or groundwater systems from septic tanks, defective sewage collecting system, or manure wash-off disposed to agriculture field for irrigation and enriching of the soil (Laine et al. 2011; Paul et al. 1997; Beller et al. 1997). For the surface water system, treated effluent from wastewater treatment plant (WWTP) is likely to be one of the most important sources of enteric pathogen contamination of the aquatic environment. Several studies have shown, that the concentration of enteric viruses throughout the treatment plant process decreases to 30–90 % (up to 4.2 log units), but effluents could still contain up to 107 genome copies of enteric viruses/L (Simmons and Xagoraraki 2011; Okoh et al. 2010; La Rosa et al. 2010; Carducci et al. 2008; Hata et al. 2013). Moreover, even low concentrations of such viruses can be infective (Koopmans and Duizer 2004) and therefore still constitute a health risk. Such low concentrations may have been overlooked in studies or screenings where efficient concentration steps were not included in the analysis.
The selected bacterial intestinal pathogen in this study was C. difficile, because of its environmental stability due to spore production and because of its current importance in public health (Magill et al. 2014). Not much is known on the prevalence of C. difficile in water or environment in general because hospitals were previously seen as most important reservoir. But long-term epidemiology data showing constant introduction of new genotypes into the hospitals and increasing incidence of community C. difficile infections are indicating that other possible reservoirs and transmission routes are important as well. C. difficile contamination of food has been well studied (Rupnik and Songer 2010; Weese 2010) and some early reports described C. difficile in drinking water and swiming pools (al Saif and Brazier 1996). But to date only a single study in Switzerland showed a high prevalence and diversity of C. difficile in treated wastewater samples (Romano et al. 2012). Distribution of C. difficile in rivers has suggested the release of WWTP effluent into surface waters as a potential source (Zidaric et al. 2010). It was also shown, that certain C. difficile ribotypes were detected in environmental samples (WWTP effluents, surface waters) as well as in patients within the same area (Romano et al. 2012; Zidaric et al. 2010). Clostridium difficile can be namely further divided into more than 300 ribotypes based on the PCR amplification of intergenic spacer regions of multicopy ribosomal operon. Additional important information for strain is the combination of three known toxins, toxin A, toxin B, and binary toxin CDT. According to changes in the region coding for toxins A and B, called PaLoc, strains could be divided into 34 toxinotypes (Rupnik 2010).
The aim of the study was to follow the release to the environment of enteric viruses and a single enteric bacterial pathogen (C. difficile) in the effluent of a large WWTP during a 1-year period. A monolithic chromatography-based method was for the first time used for simultaneous concentration of the selected enteric viruses prior to detection. In addition, physico-chemical parameters were monitored in the effluent to link possible variations with enteric viruses or C. difficile detection rate.
Materials and Methods
Description of WWTP
The selected WWTP is a conventional two-stage activated sludge plant located in central Slovenia. The capacity of the plant, designed for organic matter removal from wastewater, is 200,000 pollution equivalents (PE), and an average daily inflow of approximately 20,000 m3. The facility collects waste waters from five communities (including households), farms, and industry. The plant influent consists of 35.9 % municipal and 11.1 % of industrial wastewater. The BOD5 loading is about 6,344 kg/day (85 centil). The existing plant satisfactorily eliminates carbon components: the reduction of COD is 95.8 %, BOD5 98.6 %. As the plant is not designed for specific nutrients elimination, it is able to reduce TP 57.2 % and TN 46.1 %.
Sampling and Storage of Samples
Samples were collected from the effluent on a monthly basis from January to December 2012 with flow-proportional automatic sampler, which collects 10 l of effluent in 24 h or even more, depending on the flow. For further analysis 5 l of effluent was collected in a plastic container and stored at +4 °C during transport to laboratories and until further processing. Immediately after sampling, physico-chemical parameters were analyzed (Suppl. 1).
Concentration of Water Sample for Viral Detection
Before concentration the samples were filtrated through filter paper to remove larger precipitates. The concentration of 5 l of each sample was done using convective interaction media (CIM) quaternary amine (QA) 8 ml tube monolithic column (BIA separations, Slovenia) on an AKTA 100 purifier (GE Healthcare, Uppsala, Sweden) as described previously (Gutierrez-Aguirre et al. 2009, 2011). Briefly samples were loaded at 40–80 ml/min flow rate into the CIM QA column. Bound viruses were eluted at 4 ml/min using 50 mM HEPES and 1 M NaCl, pH 7. The elution was controlled by inline monitoring the absorption at 280 nm. The final volume after concentration was 20 ml.
Detection of Viruses
A 140 µl sample volume was used for nucleic acid extraction from 5 l of nonconcentrated samples (raw) and 20 ml of concentrated samples using the QIAamp viral RNA mini kit (QIAgen, CA, USA). As a control of the RNA extraction and to assess the presence of potentially co-extracted inhibitors 2 ng of luciferase control RNA (Promega, WI, USA) was added to each sample before RNA extraction. For each target tested with molecular methods, a specific positive control was included (for GARV, NoV-I and II, HAstV, and HSaV previously characterized strains from clinical samples were used, for HEV a porcine strain characterized as genotype 3 with sequence analysis was selected, and for HAV a cell culture propagated strain HM 175/18f was used).
For enteric virus panel we selected group A rotaviruses (GARV), noroviruses genogroup I and II (NoV-I, NoV-II), human astroviruses (HAstV), human sapoviruses (HSaV), and hepatitis A and E viruses (HAV, HEV). Real-time quantitative PCR (qPCR) assays specific for each virus were selected from previous publications (Gutierrez-Aguirre et al. 2008; Svraka et al. 2009; Kageyama et al. 2003; Costafreda et al. 2006; Jothikumar et al. 2006) and used for the detection of the mentioned pathogens (Suppl. 2). Luciferase qPCR assay was described previously by Toplak et al. (Toplak et al. 2004). One-step reverse transcription qPCR (RT-qPCR) was performed, using AgPath One-step RT-PCR kit (Life Technologies, Applied Biosystems Division, Foster City, CA) on the StepOne Real-time PCR System (Life Technologies) and ABI 7900HT system (Life Technologies). Two µl of extracted RNA was used for the detection of viral targets in a 10 µl final reaction volume. Quantification cycles (C q), which are inversely proportional to the initial target concentration, were derived for each pathogen using the SDS 2.3 and SpetOne v2.2.2 software (Life Technologies).
Detection and Characterization of C. difficile
For C. difficile isolation 50 ml of nonconcentrated water sample was subjected to heat shock at 70 °C for 20 min and then filtered through a 0.2-µm pore size cellulose nitrate membrane filter (Whatman, GE Healthcare Ltd., UK). Filters were placed on commercial selective medium chromID™ C. difficile agar (bioMérieux, France) and incubated anaerobically at 37 °C for up to 3 days. After incubation, 20 black colonies were picked from each filter and subcultured on fresh medium. Clostridium difficile was confirmed by detection of molecular marker cdd3 as described in Zidaric et al. (Zidaric et al. 2010) and characterized by PCR ribotyping (Bidet et al. 1999) and toxinotyping (Rupnik 2010).
Correlations
The linear dependancy between the independent continuous variables has been statistically measured by Pearson’s correlation coefficient which was calculated using Matlab software package. The value of Pearson’s correlation coefficient varies between −1 and 1, where 1 and −1 mean a perfect positive or negative correlation, respectively, while 0 means that there is no linear relationship between the two compared variables. As variables we used selected physical and chemical measurements at WWTP effluent and presence or absence of different viral pathogens.
Results
Detection of Enteric Viruses and C. difficile
In this study, twelve monthly samples were analyzed for the presence of seven viral enteric pathogens and C. difficile.
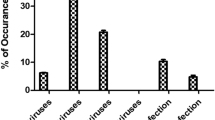
Four of seven tested viruses (GARV, NoV-I, NoV-II, HSaV) were detected in all 12 samples. HAstV was present in ten samples but remained undetectable even after the concentration step in samples from August and September. HEV and HAV were not detected during the whole length of the study, nor before neither after the concentration step (Table 1).
While viral monitoring was focused mostly on the effect of sample concentration, on the detection and on different correlations (as described below), C. difficile part was focused on isolation and characterization of strains. Clostridium difficile was isolated from each of the 12 samples analyzed during a 1-year sampling period. Altogether 121 strains were isolated and were distributed into 32 different ribotypes (Table 2). Ribotype variability differed between months with the highest number of ribotypes being detected in February to April and in July and August (Table 2). The most prevalent ribotypes were 014/020 (43 strains were found in all 12 samples), 010 (17 strains were found in 8 out of 12 samples), 046 (7 strains were found in 5 out of 12 samples). Three further ribotypes were detected in three samples (002, 005, 009), three ribotypes were detected in two samples, and 23 ribotypes were detected only in a single sample.
Detected strains belong to toxinotypes 0, I, IX, V, and XXIV (Table 2). A large proportion of strains was also nontoxigenic (45 strains; 37.2 %) and only four strains (3.3 %) had binary toxin genes.
The Effect of CIM Monolith-Based Concentration Method on Detection of Viruses
The CIM QA monolith-based concentration step used in this study was optimized previously for the concentration of rotaviruses from environmental waters (Gutierrez-Aguirre et al. 2009, 2011) and HAV and feline caliciviruses (FCV) from bottled water (Kovac et al. 2009). In those studies, the maximum viral recoveries achieved were close to 99, 40, and 20 % for rotavirus in buffer, HAV in bottled water, and FCV in bottled water, respectively. In this study the CIM-based method was applied for the first time to a complex sample such as wastewater effluent. The rotavirus recoveries, estimated from qPCR results in 6 out of the 12 samples ranged from 31 to 98 %, indicating that the method worked satisfactorily also with such complex sample. Moreover, the method proved to be applicable for simultaneous concentration of five different enteric viruses (GARV, NoV-I and II, HSaV, and HAstV) in a single step, as seen from the decrease in the C q values (inversely proportional to the target concentration) after concentration (Table 1). In 16 out of 60 performed analysis, different viruses would not be detected in the absence of the concentration step (Table 1). The concentration of WWTP effluent resulted in an increase of each virus concentration of approximately two orders of magnitude (corresponding to a decrease of 4.5 to 7 units in the C q values (ΔC q) between concentrated and nonconcentrated sample) (Table 1). The achieved concentration varied depending on the month and virus. Looking at the targets separately, the ΔC q value differed from month to month indicating a varying concentration efficiency throughout the sampling period (Table 1). The luciferase C q obtained for each sample analyzed within a particular month did not differ among themselves, indicating an optimal performance of both RNA extractions and RT-qPCR reaction (Suppl. 3).
Correlations in Concurrent Presence of Different Viral Pathogens or in Single Viral Pathogen and WWTP Parameters
Correlations were calculated to test if certain pathogens tend to be present or absent in the effluent at the same time or whether their presence or absence correlated with some physico-chemical parameters. During the study period, the detection of HAstV and HSaV correlated significantly in nonconcentrated as well as in concentrated samples (Table 3). When looking at the concentrated samples, NoV-I also showed correlation with the detection of HAstV and HSaV (Table 3).
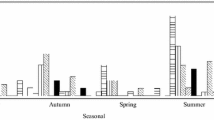
In addition, HAstV and SaV correlate also with some of the monitored physical or chemical parameters (temperature, total N, and nitrate (NO3−)) (Table 4). During the summer months, higher effluent temperature was noted and during this period, HAstV C q values were higher, meaning that viruses were present at lower concentration or were not detected at all (August, September). Similar trend, although with a lower correlation was observed also for HSaV, whereas NoV-II concentration seemed to increase during the period of higher temperature (Table 1; Fig. 1).
There was no significant correlation between different viruses and the detection of specific C. difficile ribotypes, nor between simultaneous presence of different C. difficile ribotypes.
Discussion
WWTP effluents are an important potential source of pathogens that follow the fecal-oral route of transmission. They are most likely introduced into the WWTP through municipal waste and are released to the surface water with the WWTP effluent due to inefficient removal of pathogenic viruses and bacteria. As at least 10 % of the world’s population is thought to consume foods produced by irrigation with waste water (Smit and Nasr 1992), the release of such pathogens constitute a potential risk for environment and public health. In our study C. difficile and four out of seven viral enteric pathogens (GARV, NoV-I, NoV-II, HSaV) were detected in every monthly sample during a 1-year period. Among the remaining viral pathogens, one (HAstV) was detected very often and two (HAV, HEV) never.
Concentration step contributed considerably to the detection rate of the selected viral targets. The used CIM chromatographic method was already tested previously and showed good results in concentration of single virus–rotaviruses, feline caliciviruses, or HAV in surface water, drinking water, or bottled water (Gutierrez-Aguirre et al. 2009; Kovac et al. 2009). In this study, the recoveries for rotaviruses (ranging from 31 to 98 % after estimation in 6 out of the 12 samples) and the decrease in the C qs for all viruses (except for HAV and HEV that were never detected) suggest that the method worked satisfactorily also with such complex sample as is the effluent of a WWTP. Moreover, the method proved to be effective for simultaneous concentration of most important enteric viruses in one step. According to the ΔC t values, the concentration step in our study was acceptable for all the detected viruses (Table 2). However, variation in ΔC q from sample to sample was observed. There are different possible explanations for such variations. As our concentration technique is a chromatographic method, there are probably many factors inherent to the effluent sampled each month that can influence the virus binding capacity of the column. Moreover, the qPCR detection method is known to show stochastic variations between measurements at low concentrations (Ellison et al. 2006), which could also be behind the monthly variations observed in the ΔC q for each virus. Interestingly, the highest variations in the ΔC q were observed in the samples of NoV-I and II, which are the ones showing the highest C q (lowest target concentrations) and are therefore more prone to variations originating from the qPCR stochastic effects (Table 1). HAV and HEV were never detected nor before neither after the concentration. For HAV, there is proof in the literature that the CIM-based method works, but in that study (Kovac et al. 2009) they used bottled water instead of wastewater effluent. For HEV, there is no proof, neither in this study nor in previous works. Therefore, we cannot conclude if the HEV and HAV were absent, or if they were present at such low concentrations that, even after concentration step, remained below the LOD of qPCR.
Dynamics of GARV in the WWTP effluent did not reflect the general situation in clinical infections in Slovenia. In our study, GARV was present in the WWTP effluent without a significant increase of relative concentration indicating some seasonality trend, which was an unexpected result. In Slovenia, GARV epidemiological peak is usually from February to April (Steyer et al. 2009). Thus, it was expected that higher concentration of GARV would be detected in these months. However, for a real seasonality trend of rotaviruses in the WWTP effluent, more samples should be collected in each season. In Slovenia, NoV infections appear mainly throughout the year and only a weak epidemiological peak is noted during the autumn–winter months (October–February), with NoV-II being the most prevalent. It is to assume that many outpatient cases of gastroenteritis, not reported by hospitals or local physicians and therefore not reflected in clinical epidemiologic data, are also contributing to the enteric virus present in WWTP’s.
Negative results of the effluent samples for HAV and HEV were not surprising. The official data of HAV incidence in Slovenia are very low (0.5 cases/100,000 in 2012, corresponding to 11 cases; National Institute of Public Health, http://www.nijz.si). In humans, no official data on HEV infections are available, but according to the laboratory report, confirmed cases of acute HEV are rare (Steyer et al. 2011a). HEV prevalence in pigs in Slovenia is 5.3–28.6 % (Steyer et al. 2011a) but pig farm wastewaters are collected and treated separately. Absence of HEV in WWTP effluent in this study is in concordance with the result of a similar study in Switzerland, where HEV was detected in low concentration in inlet, but was not detected in effluent of a WWTP (Masclaux et al. 2013).
For HAstV and HSaV no epidemiological data are available as HAstV and HSaV gastroenteritis is a nonreportable disease in Slovenia and there are only few laboratories which perform testing for these two pathogens. In our samples, both were usually detected together with similar dynamic of the C q values. This correlation is hard to explain with our local clinical data as HSaV is not included in the routine diagnostic scheme for enteric viruses in sporadic cases. In the recent literature, the described epidemiological profiles of HSaV and HAstV are mainly limited to hospitalized children with gastroenteritis, showing a low prevalence with up to 5.4 % (Chan-it et al. 2010; Medici et al. 2012; Chhabra et al. 2013; Gonzalez et al. 2011). HAstV and HSaV are mainly detected throughout the year, with slight increase of the detection rate in colder months, which is the trend we have observed in the effluent samples of this study. However, to obtain a real correlation between the detection rate of these viruses in human population and WWTP effluent, a cohort study should be performed and tested for viruses together with WWTP samples. Clinical data from hospitalized patients are most probably not a representative sample for such a study.
The results for C. difficile ribotypes present in WWTP effluent correspond well with the epidemiological situation known for Slovenia. The most prevalent ribotype in the strain collection from various sources covering years 2008–2010 is the only one that was present throughout the year in the WWTP effluent (Janezic et al. 2012). Other most prevalent types found in WWTP effluent are also readily isolated from humans, animals, or surface waters (Janezic et al. 2012). In the study reporting C. difficile ribotypes from patients from the same broad region as WWTP, five most prevalent ribotypes were 014/020, 150, 023, 011/049,010 (41). Two of them (014/020 and 010) were most prevalent in WWTP effluent in this study through out the year, and two others (150 and 011/049) were found sporadically.
Although the limitation of the study is low number of samples (one sample per month) collected at single WWTP treatment plant, the importance of WWTP effluent on enteric viruses and C. difficile release to the environment was clearly presented. It should be also emphasized that due to a low sampling frequency, it cannot be speculated on real seasonality trend of the detected enteric pathogens.
In summary, detection of the selected intestinal pathogens provides limited epidemiologic data for the geographic region covered by WWTP. The obtained results also confirmed that WWTP effluent can be a source of surface water contamination with intestinal pathogens. An efficient method for removal of these microorganisms would contribute to the decreasing trend of waterborne infections and infections linked to the contaminated surface water sources.
The average decrease in C q (4.5–7.1) observed after concentration step for GARV, NoV-I and II, HAstV, and HSaV, indicates that monolith chromatography can be used to concentrate these five enteric viruses simultaneously from wastewater effluent samples.
References
Al Saif, N., & Brazier, J. S. (1996). The distribution of Clostridium difficile in the environment of South Wales. Journal of Medical Microbiology, 45(2), 133–137.
Beller, M., Ellis, A., Lee, S. H., Drebot, M. A., Jenkerson, S. A., Funk, E., et al. (1997). Outbreak of viral gastroenteritis due to a contaminated well—International consequences. Jama-Journal of the American Medical Association, 278(7), 563–568. doi:10.1001/jama.278.7.563.
Bidet, P., Barbut, F., Lalande, V., Burghoffer, B., & Petit, J. C. (1999). Development of a new PCR-ribotyping method for Clostridium difficile based on ribosomal RNA gene sequencing. FEMS Microbiology Letters, 175(2), 261–266.
Bosch, A., Guix, S., Sano, D., & Pinto, R. M. (2008). New tools for the study and direct surveillance of viral pathogens in water. Current Opinion in Biotechnology, 19(3), 295–301. doi:10.1016/j.copbio.2008.04.006.
Bouzid, M., Steverding, D., & Tyler, K. M. (2008). Detection and surveillance of waterborne protozoan parasites. Current Opinion in Biotechnology, 19(3), 302–306. doi:10.1016/j.copbio.2008.05.002.
Carducci, A., Morici, P., Pizzi, F., Battistini, R., Rovini, E., & Verani, M. (2008). Study of the viral removal efficiency in a urban wastewater treatment plant. Water Science and Technology: A Journal of the International Association on Water Pollution Research, 58(4), 893–897. doi:10.2166/wst.2008.437.
Chan-it, W., Thongprachum, A., Okitsu, S., Mizuguchi, M., & Ushijima, H. (2010). Epidemiology and molecular characterization of sapovirus and astrovirus in Japan, 2008-2009. Japanese Journal of Infectious Diseases, 63(4), 302–303.
Chhabra, P., Payne, D. C., Szilagyi, P. G., Edwards, K. M., Staat, M. A., Shirley, S. H., et al. (2013). Etiology of viral gastroenteritis in children <5 years of age in the United States, 2008-2009. The Journal of Infectious Diseases, 208(5), 790–800. doi:10.1093/infdis/jit254.
Costafreda, M. I., Bosch, A., & Pinto, R. M. (2006). Development, evaluation, and standardization of a real-time TaqMan reverse transcription-PCR assay for quantification of hepatitis A virus in clinical and shellfish samples. Applied and Environmental Microbiology, 72(6), 3846–3855. doi:10.1128/AEM.02660-05.
Ellison, S. L., English, C. A., Burns, M. J., & Keer, J. T. (2006). Routes to improving the reliability of low level DNA analysis using real-time PCR. BMC Biotechnology, 6, 33. doi:10.1186/1472-6750-6-33.
Gonzalez, G. G., Liprandi, F., & Ludert, J. E. (2011). Molecular epidemiology of enteric viruses in children with sporadic gastroenteritis in Valencia, Venezuela. Journal of Medical Virology, 83(11), 1972–1982. doi:10.1002/jmv.22185.
Gutierrez-Aguirre, I., Banjac, M., Steyer, A., Poljsak-Prijatelj, M., Peterka, M., Strancar, A., et al. (2009). Concentrating rotaviruses from water samples using monolithic chromatographic supports. Journal of Chromatography A, 1216(13), 2700–2704. doi:10.1016/j.chroma.2008.10.106.
Gutierrez-Aguirre, I., Steyer, A., Banjac, M., Kramberger, P., Poljsak-Prijatelj, M., & Ravnikar, M. (2011). On-site reverse transcription-quantitative polymerase chain reaction detection of rotaviruses concentrated from environmental water samples using methacrylate monolithic supports. Journal of Chromatography A, 1218(17), 2368–2373. doi:10.1016/j.chroma.2010.10.048.
Gutierrez-Aguirre, I., Steyer, A., Boben, J., Gruden, K., Poljsak-Prijatelj, M., & Ravnikar, M. (2008). Sensitive detection of multiple rotavirus genotypes with a single reverse transcription-real-time quantitative PCR assay. Journal of Clinical Microbiology, 46(8), 2547–2554. doi:10.1128/Jcm.02428-07.
Hata, A., Kitajima, M., & Katayama, H. (2013). Occurrence and reduction of human viruses, F-specific RNA coliphage genogroups and microbial indicators at a full-scale wastewater treatment plant in Japan. Journal of Applied Microbiology, 114(2), 545–554. doi:10.1111/jam.12051.
Janezic, S., Ocepek, M., Zidaric, V., & Rupnik, M. (2012). Clostridium difficile genotypes other than ribotype 078 that are prevalent among human, animal and environmental isolates. BMC Microbiology, 12, 48. doi:10.1186/1471-2180-12-48.
Jothikumar, N., Cromeans, T. L., Robertson, B. H., Meng, X. J., & Hill, V. R. (2006). A broadly reactive one-step real-time RT-PCR assay for rapid and sensitive detection of hepatitis E virus. Journal of Virological Methods, 131(1), 65–71. doi:10.1016/j.jviromet.2005.07.004.
Kageyama, T., Kojima, S., Shinohara, M., Uchida, K., Fukushi, S., Hoshino, F. B., et al. (2003). Broadly reactive and highly sensitive assay for Norwalk-like viruses based on real-time quantitative reverse transcription-PCR. Journal of Clinical Microbiology, 41(4), 1548–1557.
Kolling, G., Wu, M., & Guerrant, R. L. (2012). Enteric pathogens through life stages. Frontiers in Cellular and Infection Microbiology, 2, 114. doi:10.3389/fcimb.2012.00114.
Koopmans, M., & Duizer, E. (2004). Foodborne viruses: An emerging problem. International Journal of Food Microbiology, 90(1), 23–41. doi:10.1016/S0168-1605(03)00169-7.
Kovac, K., Gutierrez-Aguirre, I., Banjac, M., Peterka, M., Poljsak-Prijatelj, M., Ravnikar, M., et al. (2009). A novel method for concentrating hepatitis A virus and caliciviruses from bottled water. Journal of Virological Methods, 162(1–2), 272–275. doi:10.1016/j.jviromet.2009.07.013.
La Rosa, G., Pourshaban, M., Iaconelli, M., & Muscillo, M. (2010). Quantitative real-time PCR of enteric viruses in influent and effluent samples from wastewater treatment plants in Italy. Annali dell’Istituto superiore di sanita, 46(3), 266–273. doi:10.4415/ANN_10_03_07.
Laine, J., Huovinen, E., Virtanen, M. J., Snellman, M., Lumio, J., Ruutu, P., et al. (2011). An extensive gastroenteritis outbreak after drinking-water contamination by sewage effluent, Finland. Epidemiology and Infection, 139(7), 1105–1113. doi:10.1017/S0950268810002141.
Lodder, W. J., & de Roda Husman, A. M. (2005). Presence of noroviruses and other enteric viruses in sewage and surface waters in The Netherlands. Applied and Environmental Microbiology, 71(3), 1453–1461. doi:10.1128/AEM.71.3.1453-1461.2005.
Magill, S. S., Edwards, J. R., Bamberg, W., Beldavs, Z. G., Dumyati, G., Kainer, M. A., et al. (2014). Multistate point-prevalence survey of health care-associated infections. The New England Journal of Medicine, 370(13), 1198–1208. doi:10.1056/NEJMoa1306801.
Masclaux, F. G., Hotz, P., Friedli, D., Savova-Bianchi, D., & Oppliger, A. (2013). High occurrence of hepatitis E virus in samples from wastewater treatment plants in Switzerland and comparison with other enteric viruses. Water Research, 47(14), 5101–5109. doi:10.1016/j.watres.2013.05.050.
Medici, M. C., Tummolo, F., Albonetti, V., Abelli, L. A., Chezzi, C., & Calderaro, A. (2012). Molecular detection and epidemiology of astrovirus, bocavirus, and sapovirus in Italian children admitted to hospital with acute gastroenteritis, 2008-2009. Journal of Medical Virology, 84(4), 643–650. doi:10.1002/jmv.23231.
Okoh, A. I., Sibanda, T., & Gusha, S. S. (2010). Inadequately treated wastewater as a source of human enteric viruses in the environment. International Journal of Environmental Research and Public Health, 7(6), 2620–2637. doi:10.3390/ijerph7062620.
Paul, J. H., Rose, J. B., Jiang, S. C., Zhou, X. T., Cochran, P., Kellogg, C., et al. (1997). Evidence for groundwater and surface marine water contamination by waste disposal wells in the Florida Keys. Water Research, 31(6), 1448–1454. doi:10.1016/S0043-1354(96)00374-0.
Romano, V., Pasquale, V., Krovacek, K., Mauri, F., Demarta, A., & Dumontet, S. (2012). Toxigenic Clostridium difficile PCR ribotypes from wastewater treatment plants in southern Switzerland. Applied and Environmental Microbiology, 78(18), 6643–6646. doi:10.1128/AEM.01379-12.
Rupnik, M. (2010). Clostridium difficile toxinotyping. Methods in Molecular Biology, 646, 67–76. doi:10.1007/978-1-60327-365-7_5.
Rupnik, M., & Songer, J. G. (2010). Clostridium difficile: Its potential as a source of foodborne disease. Advances in Food and Nutrition Research, 60, 53–66. doi:10.1016/S1043-4526(10)60003-4.
Simmons, F. J., & Xagoraraki, I. (2011). Release of infectious human enteric viruses by full-scale wastewater utilities. Water Research, 45(12), 3590–3598. doi:10.1016/j.watres.2011.04.001.
Smit, J., & Nasr, J. (1992). Urban agriculture for sustainable cities: Using wastes and idle land and water bodies as resources. Environment and Urbanization, 4(2), 141–152. doi:10.1177/095624789200400214.
Steyer, A., Bajzelj, M., Znuderl, K., Berce, I., Drinovec, B., Harlander, T., et al. (2009). Molecular epidemiology of rotaviruses during rotavirus vaccine introduction in Slovenia. Zdravniski Vestnik-Slovenian Medical Journal, 78(8), 381–386.
Steyer, A., Naglic, T., Mocilnik, T., Poljsak-Prijatelj, M., & Poljak, M. (2011a). Hepatitis E virus in domestic pigs and surface waters in Slovenia: Prevalence and molecular characterization of a novel genotype 3 lineage. Infection Genetics and Evolution, 11(7), 1732–1737. doi:10.1016/j.meegid.2011.07.007.
Steyer, A., Torkar, K. G., Gutierrez-Aguirre, I., & Poljsak-Prijatelj, M. (2011b). High prevalence of enteric viruses in untreated individual drinking water sources and surface water in Slovenia. International Journal of Hygiene and Environmental Health, 214(5), 392–398. doi:10.1016/j.ijheh.2011.05.006.
Svraka, S., van der Veer, B., Duizer, E., Dekkers, J., Koopmans, M., & Vennema, H. (2009). Novel approach for detection of enteric viruses to enable syndrome surveillance of acute viral gastroenteritis. Journal of Clinical Microbiology, 47(6), 1674–1679. doi:10.1128/JCM.00307-09.
Toplak, N., Okršlar, V., Stani-Racman, D., Gruden, K., & Žel, J. (2004). A high-throughput method for quantifying transgene expression in transformed plants with real-time PCR analysis. Plant Molecular Biology Reporter, 22, 237–250.
Weese, J. S. (2010). Clostridium difficile in food–innocent bystander or serious threat? Clinical microbiology and infection : the official publication of the European Society of Clinical Microbiology and Infectious Diseases, 16(1), 3–10. doi:10.1111/j.1469-0691.2009.03108.x.
Williamson, W. M., Ball, A., Wolf, S., Hewitt, J., Lin, S., Scholes, P., et al. (2011). Enteric viruses in New Zealand drinking-water sources. Water Science and Technology: A Journal of the International Association on Water Pollution Research, 63(8), 1744–1751.
Zidaric, V., Beigot, S., Lapajne, S., & Rupnik, M. (2010). The occurrence and high diversity of Clostridium difficile genotypes in rivers. Anaerobe, 16(4), 371–375. doi:10.1016/j.anaerobe.2010.06.001.
Acknowledgments
This work was supported by Slovenian National research Agency (ARRS) Grant L2-4314.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Steyer, A., Gutiérrez-Aguirre, I., Rački, N. et al. The Detection Rate of Enteric Viruses and Clostridium difficile in a Waste Water Treatment Plant Effluent. Food Environ Virol 7, 164–172 (2015). https://doi.org/10.1007/s12560-015-9183-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12560-015-9183-7