Abstract
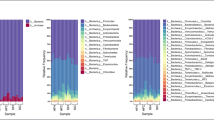
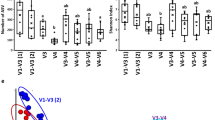
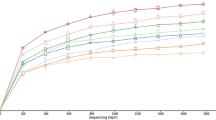
To explore the fungal diversity in ruminant feces for bioenergy, libraries based on internal transcribed spacer (ITS), 18S rRNA, and 28S rRNA regions were constructed, respectively. Although the libraries were constructed from the same DNA extracts, the fungal taxa analyses based on these libraries are different. The ITS and 28S libraries comprised higher proportions of fungal clones than 18S libraries, and the ITS libraries converged into the lower diversities. The ITS libraries could be used to analyze the fungal community. The 18S libraries were suitable for the fungi and protozoa community. However, the 28S are suitable for analysis of Ascomycota fungi. The major fungal taxa in cattle feces analyzed by ITS, 18S, and 28S libraries are similar to those of sheep feces, respectively. The fungal taxa detected by the ITS library comprised only 20 % fungal taxa detected by the three libraries. The 18S library comprised 30 % fungal taxa; the 28S library comprised about 50 % fungal taxa. The results indicated that primer sets toward different DNA regions lead to the difference in structures of fungal community. So the selection of primer sets may influence the fungal communities, and libraries based on single primer sets may underestimate the fungal diversity. The comparison of ITS, 18S, and 28S libraries could fid more diverse fungi than that based on only one library.



Similar content being viewed by others
References
Abranches J, Valente P, Nóbrega HN, Fernandez FAS, Mendonca-Hagler LC, Hagler AN (1998) Yeast diversity and killer activity dispersed in fecal pellets from marsupials and rodents in a Brazilian tropical habitat mosaic. FEMS Microbiol Ecol 26:27–33
Baker SE, Thykaer J, Adney WS, Brettin TS, Brockman FJ (2008) Fungal genome sequencing and bioenergy. Fungal Biol Rev 22:1–5
Bellemain E, Carlsen T, Brochmann C, Coissac E, Taberlet P, Kauserud H (2010) ITS as an environmental DNA barcode for fungi: an in silico approach reveals potential PCR biases. BMC Microbiol. doi:10.1186/1471-2180-10-189
Buée M, Reich M, Murat C, Morin E, Nilsson RH, Uroz S, Martín F (2009) 454 pyrosequencing analyses of forest soils reveal an unexpectedly high fungal diversity. New Phytol 184:449–456
Griffith GW, Ozkose E, Theodorou MK, Davies DR (2009) Diversity of anaerobic fungal populations in cattle revealed by seletive enrichment culture using different carbon sourcers. Fungal Ecol 2:87–97
Jebaraj CS, Raghukumar C, Behnke A, Stoeck T (2010) Fungal diversity in oxygen-depleted regions of the Arabian Sea revealed by targeted environmental sequencing combined with cultivation. FEMS Microbiol Ecol 71:399–412
Lai X, Cao L, Tan H, Fang S, Huang Y, Zhou S (2007) Fungal communities from methane hydrate-bearing deep-sea marine sediments in South China sea. ISME J 1:756–762
Liggenstoffer A, Youssef NH, Couger MB, Elshahed MS (2010) Phylogenetic diversity and community structure of anaerobic gut fungi (phylum Neocallimastigomycota) in ruminant and non-ruminant herbivores. ISME J 4:1225–1235
Lord NS, Kaplan CW, Shank P, Kitts CL, Elrod SL (2002) Assessment of fungal diversity using terminal restriction fragment (TRF) pattern analysis: comparison of 18S and ITS ribosomal regions. FEMS Microbiol Ecol 42:327–337
Nagpal R, Puniya AK, Sehgal JP, Singh K (2011) In vitro fibrolytic potential of anaerobic rumen fungi from ruminants and non-ruminant herbivores. Mycoscience 52:31–38
Nicholson MJ, McSweeney CS, Mackie RI, Brookman JL, Theodorou MK (2010) Diversity of anaerobic gut fungal populations analysed using ribosomal ITS1 sequences in faeces of wild and domesticated herbivores. Anaerobe 16:66–73
Nilsson RH, Ryberg M, Abarenkov K, Sjökvist E, Kristiansson E (2009) The ITS region as a target for characterization of fungal communities using emerging sequencing technologies. FEMS Microbiol Lett 296:97–101
Nyberg A, Persson I-L (2002) Habital differences of coprophilous fungi on moose dung. Mycol Res 106:1360–1366
Peterson R, Grinyer J, Nevalainen H (2011) Secretome of the coprophilous fungus Doratomyces stemonitis C8, isolated from koala feces. Appl Environ Microbiol 77:3793–3801
Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA, Chen W (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for fungi. Proc Natl Acad Sci U S A 109:6241–6246
Varga GA, Kolver ES (1997) Microbial and animal limitations to fiber digestion and utilization. J Nutr 127:819S-823S
Acknowledgments
This work was supported by grants from the Chinese National Natural Science Foundation (Nos. 31170011 and. 51039007) and the Fundamental Research Funds for the Central Universities.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Tan, H., Cao, L. Fungal diversity in sheep (Ovis aries) and cattle (Bos taurus) feces assessed by comparison of 18S, 28S and ITS ribosomal regions. Ann Microbiol 64, 1423–1427 (2014). https://doi.org/10.1007/s13213-013-0787-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13213-013-0787-6