Abstract
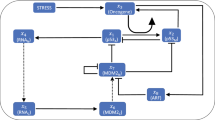
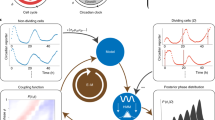
Cells living in a complex environment must constantly detect, process and appropriately respond to changing signals. Therefore, all cellular information processing is dynamic in nature. As a consequence, understanding the process of signal transduction often requires detailed quantitative analysis of dynamic behaviours. Here, we focus on the oscillatory dynamics of the tumour suppressor protein p53 as a model for studying protein dynamics in single cells to better understand its regulation and function.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Cohen, A. A. et al. Dynamic proteomics of individual cancer cells in response to a drug. Science 322, 1511–1516 (2008).
Hoffmann, A., Levchenko, A., Scott, M. L. & Baltimore, D. The IκB–NF-κB signaling module: temporal control and selective gene activation. Science 298, 1241–1245 (2002).
Nelson, D. E. et al. Oscillations in NF-κB signaling control the dynamics of gene expression. Science 306, 704–708 (2004).
Nelson, D. E., See, V., Nelson, G. & White, M. R. Oscillations in transcription factor dynamics: a new way to control gene expression. Biochem. Soc. Trans. 32, 1090–1092 (2004).
Hilioti, Z. et al. Oscillatory phosphorylation of yeast Fus3 MAP kinase controls periodic gene expression and morphogenesis. Curr. Biol. 18, 1700–1706 (2008).
Lane, D. P. Cancer. p53, guardian of the genome. Nature 358, 15–16 (1992).
Vogelstein, B., Lane, D. & Levine, A. J. Surfing the p53 network. Nature 408, 307–310 (2000).
Levine, A. J. p53, the cellular gatekeeper for growth and division. Cell 88, 323–331 (1997).
Horn, H. F. & Vousden, K. H. Coping with stress: multiple ways to activate p53. Oncogene 26, 1306–1316 (2007).
Riley, T., Sontag, E., Chen, P. & Levine, A. Transcriptional control of human p53-regulated genes. Nature Rev. Mol. Cell Biol. 9, 402–412 (2008).
Lev Bar-Or, R. et al. Generation of oscillations by the p53–Mdm2 feedback loop: a theoretical and experimental study. Proc. Natl Acad. Sci. USA 97, 11250–11255 (2000).
Lahav, G. et al. Dynamics of the p53–Mdm2 feedback loop in individual cells. Nature Genet. 36, 147–150 (2004).
Geva-Zatorsky, N. et al. Oscillations and variability in the p53 system. Mol. Syst. Biol. 2, 2006 0033 (2006).
Batchelor, E., Mock, C. S., Bhan, I., Loewer, A. & Lahav, G. Recurrent initiation: a mechanism for triggering p53 pulses in response to DNA damage. Mol. Cell 30, 277–289 (2008).
Hamstra, D. A. et al. Real-time evaluation of p53 oscillatory behavior in vivo using bioluminescent imaging. Cancer Res. 66, 7482–7489 (2006).
Murray, J. D. Mathematical Biology (eds Antman, S. S., Marsden, J. E., Sirovich, L. & Wiggins, S.) 175–256 (Springer, New York, 2002).
Edelstein-Keshet, L. Mathematical Models in Biology 311–380 (Society for Industrial and Applied Mathematics, Philadelphia, 2005).
Strogatz, S. H. Nonlinear Dynamics and Chaos 196–297 (Perseus Books Publishing, LLC, Cambridge, 1994).
Halloy, J., Lauzeral, J. & Goldbeter, A. Modeling oscillations and waves of cAMP in Dictyostelium discoideum cells. Biophys. Chem. 72, 9–19 (1998).
Tyson, J. J. & Murray, J. D. Cyclic AMP waves during aggregation of Dictyostelium amoebae. Development 106, 421–426 (1989).
Leloup, J. C., Gonze, D. & Goldbeter, A. Limit cycle models for circadian rhythms based on transcriptional regulation in Drosophila and Neurospora. J. Biol. Rhythms 14, 433–448 (1999).
Nakajima, M. et al. Reconstitution of circadian oscillation of cyanobacterial KaiC phosphorylation in vitro. Science 308, 414–415 (2005).
Johnson, C. H., Mori, T. & Xu, Y. A cyanobacterial circadian clockwork. Curr. Biol. 18, R816–R825 (2008).
Pomerening, J. R., Sontag, E. D. & Ferrell, J. E. Jr. Building a cell cycle oscillator: hysteresis and bistability in the activation of Cdc2. Nature Cell Biol. 5, 346–351 (2003).
Tyson, J. J., Csikasz-Nagy, A. & Novak, B. The dynamics of cell cycle regulation. Bioessays 24, 1095–1109 (2002).
Novak, B. & Tyson, J. J. Design principles of biochemical oscillators. Nature Rev. Mol. Cell Biol. 9, 981–991 (2008).
Shreeram, S. et al. Wip1 phosphatase modulates ATM-dependent signaling pathways. Mol. Cell 23, 757–764 (2006).
Fujimoto, H. et al. Regulation of the antioncogenic Chk2 kinase by the oncogenic Wip1 phosphatase. Cell Death Differ. 13 1170–1180 (2006).
Lu, X., Nguyen, T. A. & Donehower, L. A. Reversal of the ATM/ATR-mediated DNA damage response by the oncogenic phosphatase PPM1D. Cell Cycle 4, 1060–1064 (2005).
Lu, X., Nannenga, B. & Donehower, L. A. PPM1D dephosphorylates Chk1 and p53 and abrogates cell cycle checkpoints. Genes Dev. 19, 1162–1174 (2005).
Proctor, C. J. & Gray, D. A. Explaining oscillations and variability in the p53–MDM2 system. BMC Syst. Biol. 2, 75 (2008).
Harris, S. L. & Levine, A. J. The p53 pathway: positive and negative feedback loops. Oncogene 24, 2899–2908 (2005).
Hodgkin, A. L. & Huxley, A. F. A quantitative description of membrane current and its application to conduction and excitation in nerve. J. Physiol. 117, 500–544 (1952).
Fitzhugh, R. Impulses and physiological states in theoretical models of nerve membrane. Biophys. J. 1, 445–466 (1961).
Fitzhugh, R. Thresholds and plateaus in the Hodgkin–Huxley nerve equations. J. Gen. Physiol. 43, 867–896 (1960).
Nagumo, J., Arimoto, S. & Yoshizawa, S. An active pulse transmission line simulating nerve axon. Proc. IRE. 50, 2061–2070 (1962).
Suel, G. M., Garcia-Ojalvo, J., Liberman, L. M. & Elowitz, M. B. An excitable gene regulatory circuit induces transient cellular differentiation. Nature 440, 545–550 (2006).
Murray, A. W. & Kirschner, M. W. Dominoes and clocks: the union of two views of the cell cycle. Science 246, 614–621 (1989).
Gonze, D. & Goldbeter, A. A model for a network of phosphorylation–dephosphorylation cycles displaying the dynamics of dominoes and clocks. J. Theor. Biol. 210, 167–186 (2001).
Mirzoeva, O. K. & Petrini, J. H. DNA damage-dependent nuclear dynamics of the Mre11 complex. Mol. Cell. Biol. 21, 281–288 (2001).
Berkovich, E., Monnat, R. J. Jr & Kastan, M. B. Roles of ATM and NBS1 in chromatin structure modulation and DNA double-strand break repair. Nature Cell Biol. 9, 683–690 (2007).
Lee, J. H. & Paull, T. T. ATM activation by DNA double-strand breaks through the Mre11–Rad50–Nbs1 complex. Science 308, 551–554 (2005).
Bekker-Jensen, S., Lukas, C., Melander, F., Bartek, J. & Lukas, J. Dynamic assembly and sustained retention of 53BP1 at the sites of DNA damage are controlled by Mdc1/NFBD1. J. Cell Biol. 170, 201–211 (2005).
Matsuoka, S. et al. ATM and ATR substrate analysis reveals extensive protein networks responsive to DNA damage. Science 316, 1160–1166 (2007).
Cai, L., Dalal, C. K. & Elowitz, M. B. Frequency-modulated nuclear localization bursts coordinate gene regulation. Nature 455, 485–490 (2008).
Wei, C. L. et al. A global map of p53 transcription-factor binding sites in the human genome. Cell 124, 207–219 (2006).
Lahav, G. The strength of indecisiveness: oscillatory behavior for better cell fate determination. Sci. STKE 2004, pe55 (2004).
Zhang, T., Brazhnik, P. & Tyson, J. J. Exploring mechanisms of the DNA-damage response: p53 pulses and their possible relevance to apoptosis. Cell Cycle 6, 85–94 (2007).
Kruse, J. P. & Gu, W. SnapShot: p53 posttranslational modifications. Cell 133, 930–930.e1 (2008).
Brooks, C. L. & Gu, W. Ubiquitination, phosphorylation and acetylation: the molecular basis for p53 regulation. Curr. Opin. Cell Biol. 15, 164–171 (2003).
Bode, A. M. & Dong, Z. Post-translational modification of p53 in tumorigenesis. Nature Rev. Cancer 4, 793–805 (2004).
Banin, S. et al. Enhanced phosphorylation of p53 by ATM in response to DNA damage. Science 281, 1674–1677 (1998).
Canman, C. E. et al. Activation of the ATM kinase by ionizing radiation and phosphorylation of p53. Science 281, 1677–1679 (1998).
Toledo, F. & Wahl, G. M. Regulating the p53 pathway: in vitro hypotheses, in vivo veritas. Nature Rev. Cancer 6, 909–923 (2006).
Li, A. G. et al. An acetylation switch in p53 mediates holo-TFIID recruitment. Mol. Cell 28, 408–421 (2007).
Sims, R. J. 3rd & Reinberg, D. Is there a code embedded in proteins that is based on post-translational modifications? Nature Rev. Mol. Cell Biol. 9, 815–820 (2008).
Murray-Zmijewski, F., Slee, E. A. & Lu, X. A complex barcode underlies the heterogeneous response of p53 to stress. Nature Rev. Mol. Cell Biol. 9, 702–712 (2008).
Bond, G. L. et al. A single nucleotide polymorphism in the MDM2 promoter attenuates the p53 tumor suppressor pathway and accelerates tumor formation in humans. Cell 119, 591–602 (2004).
Hu, W. et al. A single nucleotide polymorphism in the MDM2 gene disrupts the oscillation of p53 and MDM2 levels in cells. Cancer Res. 67, 2757–2765 (2007).
Bond, G. L. et al. MDM2 SNP309 accelerates tumor formation in a gender-specific and hormone-dependent manner. Cancer Res. 66, 5104–5110 (2006).
Petenkaya, A. et al. Lack of association between the MDM2-SNP309 polymorphism and breast cancer risk. Anticancer Res. 26, 4975–4977 (2006).
Campbell, I. G., Eccles, D. M. & Choong, D. Y. No association of the MDM2 SNP309 polymorphism with risk of breast or ovarian cancer. Cancer Lett. 240, 195–197 (2006).
Krekac, D. et al. MDM2SNP309 does not associate with elevated MDM2 protein expression or breast cancer risk. Oncology 74, 84–87 (2008).
Bakkenist, C. J. & Kastan, M. B. DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. Nature 421, 499–506 (2003).
Ahn, J. Y., Schwarz, J. K., Piwnica-Worms, H. & Canman, C. E. Threonine 68 phosphorylation by ataxia telangiectasia mutated is required for efficient activation of Chk2 in response to ionizing radiation. Cancer Res. 60, 5934–5936 (2000).
Matsuoka, S. et al. Ataxia telangiectasia-mutated phosphorylates Chk2 in vivo and in vitro. Proc. Natl Acad. Sci. USA 97, 10389–10394 (2000).
Chehab, N. H., Malikzay, A., Stavridi, E. S. & Halazonetis, T. D. Phosphorylation of Ser-20 mediates stabilization of human p53 in response to DNA damage. Proc. Natl Acad. Sci. USA 96, 13777–13782 (1999).
Khosravi, R. et al. Rapid ATM-dependent phosphorylation of MDM2 precedes p53 accumulation in response to DNA damage. Proc. Natl Acad. Sci. USA 96, 14973–14977 (1999).
Wu, X., Bayle, J. H., Olson, D. & Levine, A. J. The p53-mdm-2 autoregulatory feedback loop. Genes Dev. 7, 1126–1132 (1993).
Acknowledgements
We thank all the members of our laboratory for useful discussions. This work was supported by National Institutes of Health Grant GM083303. E.B. was supported by the American Cancer Society, California Division, Pamela and Edward Taft Postdoctoral Fellowship.
Author information
Authors and Affiliations
Corresponding author
Related links
Rights and permissions
About this article
Cite this article
Batchelor, E., Loewer, A. & Lahav, G. The ups and downs of p53: understanding protein dynamics in single cells. Nat Rev Cancer 9, 371–377 (2009). https://doi.org/10.1038/nrc2604
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrc2604
This article is cited by
-
Long-term p21 and p53 dynamics regulate the frequency of mitosis events and cell cycle arrest following radiation damage
Cell Death & Differentiation (2023)
-
Der Rolle der DNA-Schadensantwort bei granulomatösen Erkrankungen
Zeitschrift für Rheumatologie (2022)
-
Deep learning in cancer pathology: a new generation of clinical biomarkers
British Journal of Cancer (2021)
-
Stability and bifurcation analysis of a delayed genetic oscillator model
Nonlinear Dynamics (2021)
-
p53 dynamics in single cells are temperature-sensitive
Scientific Reports (2020)