Abstract
For more than a century, fungal pathogens and symbionts have been known to orient hyphal growth towards chemical stimuli from the host plant1,2. However, the nature of the plant signals as well as the mechanisms underlying the chemotropic response have remained elusive3. Here we show that directed growth of the soil-inhabiting plant pathogen Fusarium oxysporum towards the roots of the host tomato (Solanum lycopersicum) is triggered by the catalytic activity of secreted class III peroxidases, a family of haem-containing enzymes present in all land plants4. The chemotropic response requires conserved elements of the fungal cell integrity mitogen-activated protein kinase (MAPK) cascade5 and the seven-pass transmembrane protein Ste2, a functional homologue of the Saccharomyces cerevisiae sex pheromone α receptor6. We further show that directed hyphal growth of F. oxysporum towards nutrient sources such as sugars and amino acids is governed by a functionally distinct MAPK cascade. These results reveal a potentially conserved chemotropic mechanism in root-colonizing fungi, and suggest a new function for the fungal pheromone-sensing machinery in locating plant hosts in a complex environment such as the soil.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
de Bary, A. Vergleichende Morphologie und Biologie der Pilze, Mycetozoen, und Bacterien (Wilhelm Engelmann, 1884)
Zentmyer, G. A. Chemotaxis of zoospores for root exudates. Science 133, 1595–1596 (1961)
Brand, A. & Gow, N. A. Mechanisms of hypha orientation of fungi. Curr. Opin. Microbiol. 12, 350–357 (2009)
Passardi, F., Penel, C. & Dunand, C. Performing the paradoxical: how plant peroxidases modify the cell wall. Trends Plant Sci. 9, 534–540 (2004)
Levin, D. E. Cell wall integrity signaling in Saccharomyces cerevisiae . Microbiol. Mol. Biol. Rev. 69, 262–291 (2005)
Arkowitz, R. A. Chemical gradients and chemotropism in yeast. Cold Spring Harb. Perspect. Biol. 1, a001958 (2009)
Berendsen, R. L., Pieterse, C. M. & Bakker, P. A. The rhizosphere microbiome and plant health. Trends Plant Sci. 17, 478–486 (2012)
Dean, R. et al. The Top 10 fungal pathogens in molecular plant pathology. Mol. Plant Pathol. 13, 414–430 (2012)
Pérez-Nadales, E. & Di Pietro, A. The membrane mucin Msb2 regulates invasive growth and plant infection in Fusarium oxysporum . Plant Cell 23, 1171–1185 (2011)
Kim, H. & Borkovich, K. A. A pheromone receptor gene, pre-1, is essential for mating type-specific directional growth and fusion of trichogynes and female fertility in Neurospora crassa . Mol. Microbiol. 52, 1781–1798 (2004)
Segall, J. E. Polarization of yeast cells in spatial gradients of alpha mating factor. Proc. Natl Acad. Sci. USA 90, 8332–8336 (1993)
Berglund, G. I. et al. The catalytic pathway of horseradish peroxidase at high resolution. Nature 417, 463–468 (2002)
Badri, D. V. et al. Root secreted metabolites and proteins are involved in the early events of plant-plant recognition prior to competition. PLoS ONE 7, e46640 (2012)
Widmann, C., Gibson, S., Jarpe, M. B. & Johnson, G. L. Mitogen-activated protein kinase: conservation of a three-kinase module from yeast to human. Physiol. Rev. 79, 143–180 (1999)
Turrà, D., Segorbe, D. & Di Pietro, A. Protein kinases in plant-pathogenic fungi: conserved regulators of infection. Annu. Rev. Phytopathol. 52, 267–288 (2014)
Di Pietro, A., García-MacEira, F. I., Méglecz, E. & Roncero, M. I. A MAP kinase of the vascular wilt fungus Fusarium oxysporum is essential for root penetration and pathogenesis. Mol. Microbiol. 39, 1140–1152 (2001)
Liu, H., Styles, C. A. & Fink, G. R. Elements of the yeast pheromone response pathway required for filamentous growth of diploids. Science 262, 1741–1744 (1993)
Buehrer, B. M. & Errede, B. Coordination of the mating and cell integrity mitogen-activated protein kinase pathways in Saccharomyces cerevisiae . Mol. Cell. Biol. 17, 6517–6525 (1997)
Martínez-Rocha, A. L. et al. Rho1 has distinct functions in morphogenesis, cell wall biosynthesis and virulence of Fusarium oxysporum . Cell. Microbiol. 10, 1339–1351 (2008)
Bar, E. E., Ellicott, A. T. & Stone, D. E. Gβγ recruits Rho1 to the site of polarized growth during mating in budding yeast. J. Biol. Chem. 278, 21798–21804 (2003)
Xue, C., Hsueh, Y. P. & Heitman, J. Magnificent seven: roles of G protein-coupled receptors in extracellular sensing in fungi. FEMS Microbiol. Rev. 32, 1010–1032 (2008)
Kretzschmar, T. et al. A petunia ABC protein controls strigolactone-dependent symbiotic signalling and branching. Nature 483, 341–344 (2012)
López-Berges, M. S., Rispail, N., Prados-Rosales, R. C. & Di Pietro, A. A nitrogen response pathway regulates virulence functions in Fusarium oxysporum via the protein kinase TOR and the bZIP protein MeaB. Plant Cell 22, 2459–2475 (2010)
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nature Methods 9, 671–675 (2012)
Smith, A. T., Sanders, S. A., Thorneley, R. N., Burke, J. F. & Bray, R. R. Characterisation of a haem active-site mutant of horseradish peroxidase, Phe41→Val, with altered reactivity towards hydrogen peroxide and reducing substrates. Eur. J. Biochem. 207, 507–519 (1992)
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007)
Tamura, K. et al. MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011)
Hirokawa, T., Boon-Chieng, S. & Mitaku, S. SOSUI: classification and secondary structure prediction system for membrane proteins. Bioinformatics 14, 378–379 (1998)
Rispail, N., & Di Pietro, A. Fusarium oxysporum Ste12 controls invasive growth and virulence downstream of the Fmk1 MAPK cascade. Mol Plant Microbe Interact 22, 830–839 (2009)
Acknowledgements
The authors are grateful to E. Martínez Aguilera for technical assistance. This work was supported by grants BIO2010-15505 and BIO2013-47870-R from the Spanish Ministerio de Innovación y Competitividad (MINECO), and BIO2008-04479 from MINECO/ERA-NET PathoGenoMics to A.D.P. M.E.G. was supported by the Marie Curie ITN ARIADNE (FP7-PEOPLE-ITN-237936) from the European Commission. F.R. had a fellowship from the ERASMUS student exchange program.
Author information
Authors and Affiliations
Contributions
D.T. and A.D.P. initiated the work and designed the experiments. D.T., M.E.G. and F.R. carried out the experiments and analysed the data. D.T. and A.D.P. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Plate assay for quantitative determination of directed hyphal growth and identification of a F. oxysporum orthologue of the S. cerevisiae α-pheromone precursor.
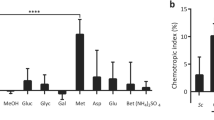
a, Schematic representation of the plate chemotropism assay. Test compound and solvent control are applied to opposite sides of a Petri dish containing a layer of water agar with 2.5 × 106ml−1 F. oxysporum microconidia, at a distance of 0.5 cm from the central scoring line. Chemotropic index was calculated as ((Htest − Hsolv)/Htotal × 100), where Htest is the number of hyphae growing towards the test compound, Hsolv is the number of hyphae growing towards the solvent control, and Htotal is the total number of hyphae counted. b, Visualization of compound diffusion and gradient establishment. The dye Congo red (1% w/v in water) was loaded into the application well on the right side of the scoring line. Diffusion was recorded photographically after the indicated time intervals. c, Dye intensity in experiment b was measured at the indicated distances from the application well after different time intervals, using the Kodak Image Analyzer software. The blue dashed line represents the relative position of the scoring line. Mean values were calculated from measurements of five individual spots per distance. d, Direction of germ tube emergence after 2 h exposure to a gradient of Glu or the solvent (H2O) was quantitatively determined by lectin–FITC staining and expressed as chemotropic index (versus H2O, *P < 0.0001). Data are presented as the mean from two experiments. n = 200 germ tube emergence sites. e, Lengths of germ tubes exposed for 13 h to a gradient of 1% (w/v) cellulose (Cel), 55 mM glucose (Gluc), 295 mM Glu or the solvent (H2O) were measured using the ImageJ software. The mean length of germ tubes growing towards the nutrient chemoattractants is not significantly different from that of germ tubes growing towards the solvent. Data are presented as the mean from two experiments. n = 100 germ tubes. Error bars show s.d. f. The predicted product of the F. oxysporum α-pheromone precursor gene (Fusarium Comparative Database accession FOXG_08636) was aligned with predicted α-pheromone precursors from F. graminearum (FGSG_05061) and F. verticillioides (FVEG_06038). Conserved residues are indicated with an asterisk. Predicted KR and RR cleavage signals for KEX2-like endopeptidases are highlighted in red. Predicted maturation signals characterized by the presence of XA or XP dipeptide repeats are highlighted in yellow. Predicted mature α-pheromone decapeptide repeats are highlighted in grey. Coloured arrowheads indicate differences between the decapeptide repeats at the third amino acid residue. g, Amino acid alignment of predicted mature α-pheromone of F. oxysporum with orthologues from ascomycete fungi. Absolutely and highly conserved residues are shaded in black and in grey, respectively. Residues replaced with alanines in the (Ala1,2) or (Ala6,7) analogues (see Fig. 1c) are indicated with asterisks.
Extended Data Figure 2 Purification of chemoattractant compounds from tomato root exudate reveals secreted peroxidases.
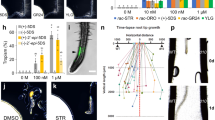
a, Chemotropic growth of germ tubes towards a gradient of tomato root exudate (RE) either untreated (no treat); treated with 1 mg ml−1 proteinase K for 30 min at 37 °C (PK); extracted to obtain an ethyl acetate fraction (EAF) and a water fraction (WF); or the WF subjected to centrifugal ultrafiltration with membranes of 10, 30 or 50 kDa molecular weight cut-off to obtain fractions < 10, 10–30, 30–50 and >50, respectively (*P = 0.006; **P < 0.0001, versus untreated). b, Anion exchange chromatography profile of fraction 30–50 from a. Obtained fractions F1–F5 are indicated. c, Directed growth of F. oxysporum germ tubes towards fractions F1–F5 from b (*P < 0.0001, versus H2O). d, SDS–PAGE of biologically active fraction F1 and inactive fraction F5, followed by staining with Coomassie blue. Protein bands present in the active and absent from the inactive fraction (named B1–B5) are indicated by arrowheads. Relative positions of molecular weight markers are indicated on the right. e, Directed growth of germ tubes towards the proteins eluted from bands B1–B5. f, Peroxidase activity of root exudates obtained from 18 individual tomato plants, indicated as units ml−1 per mg fresh root weight. Data are presented as the mean of three technical replicates. Error bars show s.d. g, Relationship between peroxidase enzymatic activity of root exudates and elicited chemotropic response. Each empty circle represents a root exudate sample from an individual tomato plant (n = 18). Linear regression (solid line) and 95% mean prediction interval (dashed lines) indicate linear correlation of the two variables (P < 0.001). h, Specific inhibitors and oxygen radical scavengers abolish peroxidase enzymatic activity. Activity of 2.5 nM commercial HRP or 100 μl root exudate was measured in the absence (C) or presence of 75 μM of the specific inhibitors thiourea (TU) or SHAM, or 250 μM of the scavenger (+)-sodium l-ascorbate (Asc), and indicated as units ml−1. Data are presented as the mean of three experiments, each with two technical replicates (*P < 0.0002, versus C). i, Peroxidase inhibitors and scavengers do not affect chemotropism towards glucose and α-pheromone. Chemotropic growth of germ tubes towards a gradient of glucose or α-pheromone, in the absence (C) or presence of 60 mM SHAM or 160 mM Asc. No significant differences were observed between treated and untreated samples. a, c, e, g, i, Data represent the mean from two experiments (a, c, g, i) or one representative experiment performed twice (e). n = 500 germ tubes. Error bars show s.d.
Extended Data Figure 3 Identification of chemoattractant proteins from tomato root exudate.
a, Peptide sequences obtained from protein bands B2 and B3 after in-gel tryptic digestion followed by LC-ESI-MS/MS. Masses were calculated by using monoisotopic masses of the occurring amino acid residues and giving peptide masses as [MH] +. b, Amino acid sequence alignment of class III tomato peroxidases TMP1 (P15003), TMP2 (P15004) and CEVI-1 (Q9LWA2), and HRP isoenzyme C (HRP_C1A) (K7ZWW6). Peptides identified in the chemotropically active fraction of tomato root exudate by LC-ESI-MS/MS are underlined in red. Predicted signal peptides are indicated by green boxes. Residues conserved in at least three of the four proteins are shaded in black. Conserved catalytic residues are indicated by orange boxes. Residues Arg 38 and His 42, which were replaced by Ser and Glu, respectively, in the catalytically inactive recombinant TMP2(R38S,H42E) protein (see Fig. 2g, h) are marked with blue asterisks.
Extended Data Figure 4 Conserved elements of the invasive growth MAPK cascade are required for chemotropism towards glucose.
a, b, Identification of ste7Δ (a) and ste11Δ (b) deletion mutants. Genomic DNA of the wild-type strain (WT) and several independent transformants was used as a template for polymerase chain reaction (PCR) with the primer pairs ste7PF + Hyg-G (P) and ste7TR + Hyg-Y (T) and ste11PF + Hyg-G (P) and ste11TR + Hyg-Y (T), respectively. Presence of an amplification product is consistent with homologous replacement of the target gene. c, d, Identification of complemented strains obtained from ste7Δ and ste11Δ mutants. Genomic DNA of independent transformants obtained upon transformation of the indicated mutants with the wild-type ste7 (c) or ste11 gene (d) was used as a template for PCR with primer pairs ste7PFN + ste7GR and ste11PFN + ste11GR, respectively. Presence of an amplification product is consistent with integration of an intact gene copy. e, Elements of the Fmk1 MAPK pathway are required for invasive growth through cellophane membranes. Colonies were grown on PDA plates covered with a cellophane membrane for 2 days at 28 °C (before). The cellophane with the fungal colony was removed and plates were incubated for an additional day (after). The experiment was performed twice, each with three plates. Results shown are from one representative experiment. Scale bar, 2 cm. f, g, Directed growth of germ tubes of the indicated F. oxysporum strains towards a gradient of glucose (Gluc) (f), α-pheromone or tomato root exudate (g) (versus wild type for a given compound, *P < 0.0001). f, g, Data are presented as the mean from two experiments. n = 500 germ tubes. Error bars show s.d.
Extended Data Figure 5 Conserved elements of the CWI MAPK cascade are required for chemotropism towards α-pheromone, root exudate and peroxidase.
a, Directed growth of germ tubes of the wild type or the fmk1Δ mutant towards a gradient of α-pheromone, in the absence or presence of PD98059 (selective p42/44 (ERK-type) MAPK inhibitor) or SB202190 (selective p38/Hog1 MAPK inhibitor) (versus wild type, *P < 0.0001). b, Identification of mpk1Δ and fmk1Δ mpk1Δ deletion mutants by Southern blot analysis. Genomic DNA of the wild-type and 11 independent transformants was treated with EcoRI, separated on a 0.7% agarose gel, transferred to a nylon membrane and hybridized with a DNA probe corresponding to the 3′ flanking region of the mpk1 gene. Transformants #1, #4, #7 (wild-type background) and #1, #2, #4, #7 (fmk1Δ background) show a banding pattern consistent with targeted deletion of the mpk1 gene. c, d, Identification of mkk2Δ (c) and bck1Δ (d) deletion mutants. Genomic DNA of independent transformants was used as template for PCR with the primer pairs mkk2PF + Hyg-G (P) and mkk2TR + Hyg-Y (T), or bck1PF + Hyg-G (P) and bck1TR + Hyg-Y (T), respectively. Presence of an amplification product is consistent with homologous replacement of the target gene. e, f, Identification of complemented strains obtained from mkk2Δ and bck1Δ mutants. Genomic DNA of independent transformants obtained after transformation of the indicated mutants with the wild-type mkk2 (c) or bck1 allele (d) was used as a template for PCR with the primer pairs mkk2PFN + mkk2GR, or bck1PFN + bck1GR, respectively. Presence of an amplification product is consistent with integration of an intact gene copy. g, Elements of the Mpk1 MAPK pathway are required for the cell wall stress response. Colony phenotypes of the indicated strains grown on yeast peptone dextrose medium (YPD) in the absence or presence of the cell-wall-perturbing compounds Calcofluor white (20 μg ml−1) or Congo red (100 μg ml−1). Plates were spot-inoculated with the indicated amount of microconidia, incubated for 4 days at 28 °C and scanned. The experiment was performed twice, each with three plates. Results shown are from one representative experiment. h, i, Directed growth of germ tubes of the indicated F. oxysporum strains towards a gradient of glucose (Gluc) (h), α-pheromone, tomato root exudate or HRP (i) (versus wild type for a given compound, *P < 0.0001). a, h, i, Data are presented as the mean from two experiments. n = 500 germ tubes. Error bars show s.d.
Extended Data Figure 6 Hyphal tip projection angle assay reveals differential roles of Fmk1 and Mpk1 MAPKs in chemotropism towards glutamate and α-pheromone.
a, Schematic representation of the chemotropism plate assay based on measurement of hyphal tip projection angles. b, Average cosine of hyphal tip projection angles of the F. oxysporum wild-type, fmk1Δ or mpk1Δ strains towards a gradient of Glu, α-pheromone or the water control. Data are presented as the mean from three experiments. n = 100 germ tubes. Bars indicate upper and lower 95% significance limits for cosine means according to a t-test. A cosine of 1 means perfect orientation while 0 means random orientation. Chemotropism was considered significant when the lower confidence limit was >0.
Extended Data Figure 7 Loss of Ste2 negatively affects virulence of F. oxysporum on tomato plants.
a, Phylogram of Ste2 orthologues from ascomycete fungi. The analysis was conducted using the MEGA5 program. Distances were inferred using the unweighted pair group method with arithmetic mean (UPGMA). b. Two-dimensional model of the transmembrane topology of F. oxysporum Ste2. The model was generated using the SOSUI software23. Amino acid residues in the primary and secondary transmembrane helix are indicated in dark and light green, respectively. Hydrophobic, positively, and negatively charged residues are marked in black, blue, and red, respectively. c, d, Southern blot analysis to identify ste2Δ (c) and fmk1Δ ste2Δ (d) deletion mutants. Genomic DNA of wild type and the indicated transformants was treated with EcoRI, separated on a 0.7% agarose gel, transferred to a nylon membrane and hybridized with a DNA probe corresponding to the 5′ flanking sequence of the ste2 gene. Transformants #1 and #8 in c and #4, #5 and #9 in d show a banding pattern consistent with targeted deletion of the ste2 gene by homologous integration of a single construct. e, Loss of Ste2 negatively affects virulence of F. oxysporum on tomato seedlings. Surface-sterilized tomato seeds (cultivar Monika) were germinated in glass tubes with 4 ml 0.5% water agar containing 2.5 × 106ml−1 microconidia of the indicated F. oxysporum strains and incubated at 28 °C under a daily cycle of 15 h light and 9 h dark. Plant survival was recorded for 32 days. Plants inoculated with the ste2Δ mutant showed significantly lower mortality than those inoculated with the wild-type and the complemented strain (P = 0.02, log-rank test). n = 40 plants. Results shown are from one representative experiment. Experiments were performed twice with similar results.
Extended Data Figure 8 Mechanism of chemotropic signalling in Fusarium oxysporum.
Chemotropic sensing of nutrients such as glucose or Glu is mediated by the Fmk1 MAPK pathway. Chemotropic sensing of α-pheromone and plant peroxidase-derived signals requires the 7TM-domain receptor Ste2 and the Mpk1 MAPK pathway. Dotted arrows between components denote unknown mechanistic links.
Rights and permissions
About this article
Cite this article
Turrà, D., El Ghalid, M., Rossi, F. et al. Fungal pathogen uses sex pheromone receptor for chemotropic sensing of host plant signals. Nature 527, 521–524 (2015). https://doi.org/10.1038/nature15516
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15516
This article is cited by
-
Fungal Cultivars of Higher Attine Ants Promote Escovopsis Chemotropism
Current Microbiology (2024)
-
Fused expression of Sm1-Chit42 proteins for synergistic mycoparasitic response of Trichoderma afroharzianum on Botrytis cinerea
Microbial Cell Factories (2023)
-
MAPkinases regulate secondary metabolism, sexual development and light dependent cellulase regulation in Trichoderma reesei
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.