Abstract
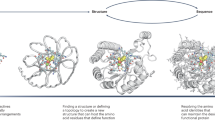
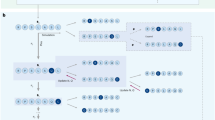
Computational enzyme design holds promise for the production of renewable fuels, drugs and chemicals. De novo enzyme design has generated catalysts for several reactions, but with lower catalytic efficiencies than naturally occurring enzymes1,2,3,4. Here we report the use of game-driven crowdsourcing to enhance the activity of a computationally designed enzyme through the functional remodeling of its structure. Players of the online game Foldit5,6 were challenged to remodel the backbone of a computationally designed bimolecular Diels-Alderase3 to enable additional interactions with substrates. Several iterations of design and characterization generated a 24-residue helix-turn-helix motif, including a 13-residue insertion, that increased enzyme activity >18-fold. X-ray crystallography showed that the large insertion adopts a helix-turn-helix structure positioned as in the Foldit model. These results demonstrate that human creativity can extend beyond the macroscopic challenges encountered in everyday life to molecular-scale design problems.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
References
Rothlisberger, D. et al. Kemp elimination catalysts by computational enzyme design. Nature 453, 190–195 (2008).
Jiang, L. et al. De novo computational design of retro-aldol enzymes. Science 319, 1387–1391 (2008).
Siegel, J.B. et al. Computational design of an enzyme catalyst for a stereoselective bimolecular Diels-Alder reaction. Science 329, 309–313 (2010).
Bar-Even, A. et al. The moderately efficient enzyme: evolutionary and physicochemical trends shaping enzyme parameters. Biochemistry 50, 4402–4410 (2011).
Khatib, F. et al. Crystal structure of a monomeric retroviral protease solved by protein folding game players. Nat. Struct. Mol. Biol. 18, 1175–1177 (2011).
Cooper, S. et al. Predicting protein structures with a multiplayer online game. Nature 466, 756–760 (2010).
Sterner, R. & Hocker, B. Catalytic versatility, stability, and evolution of the (βα)8-barrel enzyme fold. Chem. Rev. 105, 4038–4055 (2005).
Hu, X., Wang, H., Ke, H. & Kuhlman, B. High-resolution design of a protein loop. Proc. Natl. Acad. Sci. USA 104, 17668–17673 (2007).
Murphy, P.M., Bolduc, J.M., Gallaher, J.L., Stoddard, B.L. & Baker, D. Alteration of enzyme specificity by computational loop remodeling and design. Proc. Natl. Acad. Sci. USA 106, 9215–9220 (2009).
Havranek, J.J. & Baker, D. Motif-directed flexible backbone design of functional interactions. Protein Sci. 18, 1293–1305 (2009).
Park, H.S. et al. Design and evolution of new catalytic activity with an existing protein scaffold. Science 311, 535–538 (2006).
Religa, T.L. et al. The helix-turn-helix motif as an ultrafast independently folding domain: the pathway of folding of Engrailed homeodomain. Proc. Natl. Acad. Sci. USA 104, 9272–9277 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
The PyMOL Molecular Graphics System, Version 1.3, Schrödinger, LLC.
Kunkel, T.A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Proc. Natl. Acad. Sci. USA 82, 488–492 (1985).
Siegel, J.B. et al. Computational design of an enzyme catalyst for a stereoselective bimolecular Diels-Alder reaction. Science 329, 309–313 (2010).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods In Enzymology: Macromolecular Crystallography. Part A 276, 307–326 (1997).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Collaborative Computational Project, Number 4. The CCP4 Suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Scharff, E.I., Koepke, J., Fritzsch, G., Luecke, C. & Rueterjans, H. Crystal structure of diisopropylfluorophosphatase from Loligo vulgaris. Structure 9, 493–502 (2001).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Krissinel, E. & Henrick, K. Secondary-structure matching (SSM), a new tool for fast protein structure alignment in three dimensions. Acta Crystallogr. D Biol. Crystallogr. 60, 2256–2268 (2004).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Winn, M.D., Isupov, M.N. & Murshudov, G.N. Use of TLS parameters to model anisotropic displacements in macromolecular refinement. Acta Crystallogr. D Biol. Crystallogr. 57, 122–133 (2001).
Vagin, A.A. et al. Organization of prior chemical knowledge and guidelines for its use. Acta Crystallogr. D Biol. Crystallogr. 60, 2184–2195 (2004).
Laskowski, R.A., Macarthur, M.W., Moss, D.S. & Thornton, J.M. PROCHECK: a program to check the stereochemical quality of protein structures. J. Appl. Cryst. 26, 283–291 (1993).
Acknowledgements
We would like to acknowledge the members of the Foldit team for their help designing and developing the game and all the Foldit players who volunteered to make this work possible. We would also like to thank J. Thompson for useful scripts, as well as B. Siegel and M. Eiben for helpful comments on the manuscript. This work was supported by the Center for Game Science at the University of Washington, US Defense Advanced Research Projects Agency (DARPA) grant N00173-08-1-G025, the DARPA PDP program, the Howard Hughes Medical Institute (D.B.), a National Science Foundation graduate research fellowship to J.B.B. (grant no. DGE-0718124), and a National Science Foundation grant for F.K. (grant no. 0906026).
Author information
Authors and Affiliations
Contributions
C.B.E. analyzed community models, in addition to designing, building and testing the enzyme libraries. J.B.S. designed the experimental and computational methods, and built the initial computational models. F.K. set up the Foldit puzzles and curated the player results for analysis by C.B.E. S.C. led design and development of Foldit. B.L.S., J.B.B. and B.W.S. grew the crystals and collected X-ray diffraction data, processed X-ray data and analyzed the structure. Foldit Players designed new protein backbones and sequences. Z.P. and D.B. contributed to the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Table 1, Supplementary Libraries 1–3, Supplementary Data, Supplementary Methods and Supplementary Figures 1–9 (PDF 4381 kb)
Rights and permissions
About this article
Cite this article
Eiben, C., Siegel, J., Bale, J. et al. Increased Diels-Alderase activity through backbone remodeling guided by Foldit players. Nat Biotechnol 30, 190–192 (2012). https://doi.org/10.1038/nbt.2109
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2109
This article is cited by
-
Building enzymes from scratch
Nature Chemistry (2022)
-
Engineering an efficient and enantioselective enzyme for the Morita–Baylis–Hillman reaction
Nature Chemistry (2022)
-
Exo-selective intermolecular Diels–Alder reaction by PyrI4 and AbnU on non-natural substrates
Communications Chemistry (2021)
-
Computer-aided mind map generation via crowdsourcing and machine learning
Research in Engineering Design (2020)
-
De novo protein design by citizen scientists
Nature (2019)