Abstract
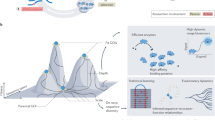
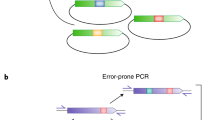
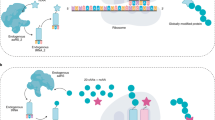
We describe synthetic shuffling, an evolutionary protein engineering technology in which every amino acid from a set of parents is allowed to recombine independently of every other amino acid. With the use of degenerate oligonucleotides, synthetic shuffling provides a direct route from database sequence information to functional libraries. Physical starting genes are unnecessary, and additional design criteria such as optimal codon usage or known beneficial mutations can also be incorporated. We performed synthetic shuffling of 15 subtilisin genes and obtained active and highly chimeric enzymes with desirable combinations of properties that we did not obtain by other directed-evolution methods.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Crameri, A., Raillard, S.-A., Bermudez, E. & Stemmer, W.P.C. DNA shuffling of a family of genes from diverse species accelerates directed evolution. Nature 391, 288–291 (1998).
Chang, C.C. et al. Evolution of a cytokine using DNA family shuffling. Nat. Biotechnol. 17, 793–797 (1999).
Ness, J.E., delCardayre, S.B., Minshull, J. & Stemmer, W.P.C. Adv. Protein Chem. 55, 261–292 (2000).
Raillard, S.-A. et al. Novel enzyme activities and functional plasticity revealed by recombining highly homologous enzymes. Chem. Biol. 8, 891–898 (2001).
Moore, G.L., Maranas, C.D., Lutz, S. & Benkovic, S.J. Predicting crossover generation in DNA shuffling. Proc. Natl. Acad. Sci. USA 98, 3226–3231 (2001).
Sun, F. Modeling DNA shuffling. J. Computational Biol. 6, 77–90 (1999).
Ness, J.E. et al. DNA shuffling of subgenomic sequences of subtilisin. Nat. Biotechnol. 17, 893–896 (1999).
Bott, R. et al. in Subtilisin Enzymes: Practical Protein Engineering Vol. 379 (eds. Bott, R. & Betzel, C.) 277–283 (Plenum, New York, 1996).
Rao, M.B., Tanksale, A.M., Ghatge, M.S. & Deshpande, V.V. Molecular and biotechnological aspects of microbial proteases. Microbiol. Mol. Biol. Rev. 62, 597–635 (1998).
Gupta, R., Beg, Q.K. & Lorenz, P. Bacterial alkaline proteases: molecular approaches and industrial applications. Appl. Microbiol. Biotechnol. 59, 15–32 (2002).
Bryan, P.N. Protein engineering of subtilisin. Biochim. Biophys. Acta 1543, 203–222 (2000).
Siezen, R.J., de Vos, W.M., Leunissen, J.A.M. & Dijkstra, B.W. Homology modelling and protein engineering strategy of subtilases, the family of subtilisin-like serine proteases. Protein Eng. 4, 719–737 (1991).
Graycar, T., Knapp, M., Ganshaw, G., Dauberman, J. & Bott, R. Engineered Bacillus lentus subtilisins having altered flexibility. J. Mol. Biol. 292, 97–109 (1999).
Roberts, R.W. & Szostak, J.W. RNA–peptide fusions for the in vitro selection of peptides and proteins. Proc. Natl. Acad. Sci. USA 94, 12297–12302 (1997).
Hanes, J. & Pluckthun, A. In vitro selection and evolution of functional proteins by using ribosome display. Proc. Natl. Acad. Sci. USA 94, 4937–4942 (1997).
Keefe, A.D. & Szostak, J.W. Functional proteins from a random-sequence library. Nature 410, 715–718 (2001).
Wilson, D.S., Keefe, A.D. & Szostak, J.W. The use of mRNA display to select high-affinity protein-binding peptides. Proc. Natl. Acad. Sci. USA 98, 3750–3755 (2001).
Bryan, P.N. et al. Proteases of enhanced stability: characterization of a thermostable variant of subtilisin. Proteins 1, 326–334 (1986).
Gilliland, G.L., Gallagher, D.T., Alexander, P. & Bryan, P.N. in Subtilisin Enzymes: Practical Protein Engineering (eds. Bott, R. & Betzel, C.) 159–169 (Plenum Press, New York, 1996).
Ostermeier, M., Shim, J.H. & Benkovic, S.J. A combinatorial approach to hybrid enzymes independent of DNA homology. Nat. Biotechnol. 17, 1205–1209 (1999).
Sieber, V., Martinez, C.A. & Arnold, F.H. Libraries of hybrid proteins from distantly related sequences. Nat. Biotechnol. 19, 456–460 (2001).
Kikuchi, M., Ohnishi, K. & Harayama, S. Novel family shuffling methods for the in vitro evolution of enzymes. Gene 236, 159–167 (1999).
Kikuchi, M., Ohnishi, K. & Harayama, S. An effective family shuffling method using single-stranded DNA. Gene 243, 133–137 (2000).
Coco, W.M. et al. DNA shuffling method for generating highly recombined genes and evolved enzymes. Nat. Biotechnol. 19, 354–359 (2001).
Gibbs, M.D., Nevalainen, K.M.H. & Bergquist, P.L. Degenerate oligonucleotide gene shuffling (DOGS): a method for enhancing the frequency of recombination with family shuffling. Gene 271, 13–20 (2001).
Voigt, C.A., Mayo, S.L., Arnold, F.H. & Wang, Z.-G. Computational method to reduce the search space for directed protein evolution. Proc. Natl. Acad. Sci. USA 98, 3778–3783 (2001).
Stemmer, W.P.C., Crameri, A., Ha, K.D., Brennan, T.M. & Heyneker, H.L. Single-step assembly of a gene and entire plasmid from large numbers of oligodeoxyribonucleotides. Gene 164, 49–53 (1995).
Stemmer, W.P. DNA shuffling by random fragmentation and reassembly: in vitro recombination for molecular evolution. Proc. Natl. Acad. Sci. USA 91, 10747–10751 (1994).
Shafikhani, S., Siegel, R.A., Ferrari, E. & Schellenberger, V. Generation of large libraries of random mutants in Bacillus subtilis by PCR-based plasmid multimerization. Biotechniques 23, 304–310 (1997).
Rice, J.A. Mathematical Statistics and Data Analysis Edn 2 (Duxbury Press, Belmont, 1995).
Acknowledgements
We thank Mark Welch for screening and automation assistance, Andreas Crameri and Pim Stemmer for advice on library design and assembly parameters, Troy Obrero and Walker Lutringer for DNA sequencing, Ajoy Roy for statistical advice, and Allan Svendsen and Bo Hammer for critically reading the manuscript.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Ness, J., Kim, S., Gottman, A. et al. Synthetic shuffling expands functional protein diversity by allowing amino acids to recombine independently. Nat Biotechnol 20, 1251–1255 (2002). https://doi.org/10.1038/nbt754
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt754
This article is cited by
-
Identification of Thermus aquaticus DNA polymerase variants with increased mismatch discrimination and reverse transcriptase activity from a smart enzyme mutant library
Scientific Reports (2019)
-
Trends in extracellular serine proteases of bacteria as detergent bioadditive: alternate and environmental friendly tool for detergent industry
Archives of Microbiology (2019)
-
Creating a more robust 5-hydroxymethylfurfural oxidase by combining computational predictions with a novel effective library design
Biotechnology for Biofuels (2018)
-
Highly active enzymes by automated combinatorial backbone assembly and sequence design
Nature Communications (2018)
-
Methods for the directed evolution of proteins
Nature Reviews Genetics (2015)