Abstract
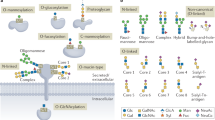
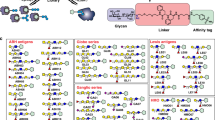
In comparison with genomics and proteomics, the advancement of glycomics has faced unique challenges in the pursuit of developing analytical and biochemical tools and biological readouts to investigate glycan structure-function relationships. Glycans are more diverse in terms of chemical structure and information density than are DNA and proteins. This diversity arises from glycans' complex nontemplate-based biosynthesis, which involves several enzymes and isoforms of these enzymes. Consequently, glycans are expressed as an 'ensemble' of structures that mediate function. Moreover, unlike protein-protein interactions, which can be generally viewed as 'digital' in regulating function, glycan-protein interactions impinge on biological functions in a more 'analog' fashion that can in turn 'fine-tune' a biological response. This fine-tuning by glycans is achieved through the graded affinity, avidity and multivalency of their interactions. Given the importance of glycomics, this review focuses on areas of technologies and the importance of developing a bioinformatics platform to integrate the diverse datasets generated using the different technologies to allow a systems approach to glycan structure-function relationships.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Lowe, J.B. & Marth, J.D. A genetic approach to mammalian glycan function. Annu. Rev. Biochem. 72, 643–691 (2003).
Varki, A. et al. Essentials of Glycobiology (Cold Spring Harber Laboratory Press, New York, 1999).
Taylor, M.E. & Drickamer, K. Introduction to glycobiology (Oxford University Press, Oxford and New York, 2003).
Raman, R., Sasisekharan, V. & Sasisekharan, R. Structural insights into biological roles of protein-glycosaminoglycan interactions. Chem. Biol. 12, 267–277 (2005).
Hwang, H.Y., Olson, S.K., Esko, J.D. & Horvitz, H.R. Caenorhabditis elegans early embryogenesis and vulval morphogenesis require chondroitin biosynthesis. Nature 423, 439–443 (2003).
Inatani, M., Irie, F., Plump, A.S., Tessier-Lavigne, M. & Yamaguchi, Y. Mammalian brain morphogenesis and midline axon guidance require heparan sulfate. Science 302, 1044–1046 (2003).
Lin, X. Functions of heparan sulfate proteoglycans in cell signaling during development. Development 131, 6009–6021 (2004).
Haltiwanger, R.S. & Lowe, J.B. Role of glycosylation in development. Annu. Rev. Biochem. 73, 491–537 (2004).
Cipollo, J.F., Awad, A.M., Costello, C.E. & Hirschberg, C.B. N-glycans of Caenorhabditis elegans are specific to developmental stages. J. Biol. Chem. 280, 26063–26072 (2005).
Sasisekharan, R., Shriver, Z., Venkataraman, G. & Narayanasami, U. Roles of heparan-sulphate glycosaminoglycans in cancer. Nat. Rev. Cancer 2, 521–528 (2002).
Liu, D., Shriver, Z., Venkataraman, G., El Shabrawi, Y. & Sasisekharan, R. Tumor cell surface heparan sulfate as cryptic promoters or inhibitors of tumor growth and metastasis. Proc. Natl. Acad. Sci. USA 99, 568–573 (2002).
Fuster, M.M., Brown, J.R., Wang, L. & Esko, J.D. A disaccharide precursor of sialyl Lewis X inhibits metastatic potential of tumor cells. Cancer Res. 63, 2775–2781 (2003).
Ishida, H. et al. A novel β1,3-N-acetylglucosaminyltransferase (β3Gn-T8), which synthesizes poly(N-acetyllactosamine), is dramatically upregulated in colon cancer. FEBS Lett. 579, 71–78 (2005).
Iwai, T. et al. Core 3 synthase is downregulated in colon carcinoma and profoundly suppresses the metastatic potential of carcinoma cells. Proc. Natl. Acad. Sci. USA 102, 4572–4577 (2005).
Dube, D.H. & Bertozzi, C.R. Glycans in cancer and inflammation–potential for therapeutics and diagnostics. Nat. Rev. Drug Discov. 4, 477–488 (2005).
Casu, B., Guerrini, M. & Torri, G. Structural and conformational aspects of the anticoagulant and anti-thrombotic activity of heparin and dermatan sulfate. Curr. Pharm. Des. 10, 939–949 (2004).
Petitou, M. & van Boeckel, C.A. A synthetic antithrombin III binding pentasaccharide is now a drug! What comes next? Angew. Chem. Int. Edn Engl. 43, 3118–3133 (2004).
Shriver, Z., Liu, D. & Sasisekharan, R. Emerging views of heparan sulfate glycosaminoglycan structure/activity relationships modulating dynamic biological functions. Trends Cardiovasc. Med. 12, 71–77 (2002).
Kinjo, Y. et al. Recognition of bacterial glycosphingolipids by natural killer T cells. Nature 434, 520–525 (2005).
Guo, Y. et al. Structural basis for distinct ligand-binding and targeting properties of the receptors DC-SIGN and DC-SIGNR. Nat. Struct. Mol. Biol. 11, 591–598 (2004).
Crocker, P.R. Siglecs in innate immunity. Curr. Opin. Pharmacol. 5, 431–437 (2005).
Rudd, P.M., Wormald, M.R. & Dwek, R.A. Sugar-mediated ligand-receptor interactions in the immune system. Trends Biotechnol. 22, 524–530 (2004).
Rudd, P.M., Elliott, T., Cresswell, P., Wilson, I.A. & Dwek, R.A. Glycosylation and the immune system. Science 291, 2370–2376 (2001).
Collins, B.E. & Paulson, J.C. Cell surface biology mediated by low affinity multivalent protein-glycan interactions. Curr. Opin. Chem. Biol. 8, 617–625 (2004).
Crocker, P.R. Siglecs: sialic-acid-binding immunoglobulin-like lectins in cell-cell interactions and signalling. Curr. Opin. Struct. Biol. 12, 609–615 (2002).
Fry, E.E. et al. The structure and function of a foot-and-mouth disease virus-oligosaccharide receptor complex. EMBO J. 18, 543–554 (1999).
Ganesh, V.K., Smith, S.A., Kotwal, G.J. & Murthy, K.H. Structure of vaccinia complement protein in complex with heparin and potential implications for complement regulation. Proc. Natl. Acad. Sci. USA 101, 8924–8929 (2004).
Liu, J. et al. Characterization of a heparan sulfate octasaccharide that binds to herpes simplex virus type 1 glycoprotein d. J. Biol. Chem. 277, 33456–33467 (2002).
Mahdavi, J. et al. Helicobacter pylori SabA adhesin in persistent infection and chronic inflammation. Science 297, 573–578 (2002).
Miller, S.I., Ernst, R.K. & Bader, M.W. LPS, TLR4 and infectious disease diversity. Nat. Rev. Microbiol. 3, 36–46 (2005).
Shriver, Z., Raguram, S. & Sasisekharan, R. Glycomics: a pathway to a class of new and improved therapeutics. Nat. Rev. Drug Discov. 3, 863–873 (2004).
Sasisekharan, R. & Venkataraman, G. Heparin and heparan sulfate: biosynthesis, structure and function. Curr. Opin. Chem. Biol. 4, 626–631 (2000).
Sugahara, K. et al. Recent advances in the structural biology of chondroitin sulfate and dermatan sulfate. Curr. Opin. Struct. Biol. 13, 612–620 (2003).
Blixt, O. et al. Chemoenzymatic synthesis of 2-azidoethyl-ganglio-oligosaccharides GD3, GT3, GM2, GD2, GT2, GM1, and GD1a. Carbohydr. Res. 340, 1963–1972 (2005).
Hanson, S., Best, M., Bryan, M.C. & Wong, C.H. Chemoenzymatic synthesis of oligosaccharides and glycoproteins. Trends Biochem. Sci. 29, 656–663 (2004).
Homeister, J.W., Daugherty, A. & Lowe, J.B. α(1,3)fucosyltransferases FucT-IV and FucT-VII control susceptibility to atherosclerosis in apolipoprotein E−/− mice. Arterioscler. Thromb. Vasc. Biol. 24, 1897–1903 (2004).
Smithson, G. et al. Fuc-TVII is required for T helper 1 and T cytotoxic 1 lymphocyte selectin ligand expression and recruitment in inflammation, and together with Fuc-TIV regulates naive T cell trafficking to lymph nodes. J. Exp. Med. 194, 601–614 (2001).
Martin, L.T., Marth, J.D., Varki, A. & Varki, N.M. Genetically altered mice with different sialyltransferase deficiencies show tissue-specific alterations in sialylation and sialic acid 9-O-acetylation. J. Biol. Chem. 277, 32930–32938 (2002).
Comelli, E.M., Amado, M., Head, S.R. & Paulson, J.C. Custom microarray for glycobiologists: considerations for glycosyltransferase gene expression profiling. Biochem. Soc. Symp. 69, 135–142 (2002).
An, H.J., Peavy, T.R., Hedrick, J.L. & Lebrilla, C.B. Determination of N-glycosylation sites and site heterogeneity in glycoproteins. Anal. Chem. 75, 5628–5637 (2003).
Cipollo, J.F., Costello, C.E. & Hirschberg, C.B. The fine structure of Caenorhabditis elegans N-glycans. J. Biol. Chem. 277, 49143–49157 (2002).
Dell, A. & Morris, H.R. Glycoprotein structure determination by mass spectrometry. Science 291, 2351–2356 (2001).
Morelle, W., Page, A. & Michalski, J.C. Electrospray ionization ion trap mass spectrometry for structural characterization of oligosaccharides derivatized with 2-aminobenzamide. Rapid Commun. Mass Spectrom. 19, 1145–1158 (2005).
Morelle, W. et al. Fragmentation characteristics of permethylated oligosaccharides using a matrix-assisted laser desorption/ionization two-stage time-of-flight (TOF/TOF) tandem mass spectrometer. Rapid Commun. Mass Spectrom. 18, 2637–2649 (2004).
Kameyama, A. et al. Detection of oligosaccharides labeled with cyanine dyes using matrix-assisted laser desorption/ionization mass spectrometry. Anal. Chem. 76, 4537–4542 (2004).
Goldberg, D., Sutton-Smith, M., Paulson, J. & Dell, A. Automatic annotation of matrix-assisted laser desorption/ionization N-glycan spectra. Proteomics 5, 865–875 (2005).
Rudd, P.M. et al. A high-performance liquid chromatography based strategy for rapid, sensitive sequencing of N-linked oligosaccharide modifications to proteins in sodium dodecyl sulphate polyacrylamide electrophoresis gel bands. Proteomics 1, 285–294 (2001).
Royle, L. et al. An analytical and structural database provides a strategy for sequencing O-glycans from microgram quantities of glycoproteins. Anal. Biochem. 304, 70–90 (2002).
Ethier, M., Saba, J.A., Ens, W., Standing, K.G. & Perreault, H. Automated structural assignment of derivatized complex N-linked oligosaccharides from tandem mass spectra. Rapid Commun. Mass Spectrom. 16, 1743–1754 (2002).
Joshi, H.J. et al. Development of a mass fingerprinting tool for automated interpretation of oligosaccharide fragmentation data. Proteomics 4, 1650–1664 (2004).
Lohmann, K.K. & von der Lieth, C.W. GlycoFragment and GlycoSearchMS: web tools to support the interpretation of mass spectra of complex carbohydrates. Nucleic Acids Res. 32, W261–W266 (2004).
Tang, H., Mechref, Y. & Novotny, M.V. Automated interpretation of MS/MS spectra of oligosaccharides. Bioinformatics 21 (Suppl. 1), i431–i439 (2005).
Park, Y. & Lebrilla, C.B. Application of Fourier transform ion cyclotron resonance mass spectrometry to oligosaccharides. Mass Spectrom. Rev. 24, 232–264 (2005).
Hakansson, K. et al. Electron capture dissociation and infrared multiphoton dissociation MS/MS of an N-glycosylated tryptic peptic to yield complementary sequence information. Anal. Chem. 73, 4530–4536 (2001).
McFarland, M.A. et al. Structural characterization of the GM1 ganglioside by infrared multiphoton dissociation, electron capture dissociation, and electron detachment dissociation electrospray ionization FT-ICR MS/MS. J. Am. Soc. Mass Spectrom. 16, 752–762 (2005).
Froesch, M. et al. Coupling of fully automated chip electrospray to Fourier transform ion cyclotron resonance mass spectrometry for high-performance glycoscreening and sequencing. Rapid Commun. Mass Spectrom. 18, 3084–3092 (2004).
Zamfir, A. et al. Fully automated chip-based mass spectrometry for complex carbohydrate system analysis. Anal. Chem. 76, 2046–2054 (2004).
Guerrini, M., Bisio, A. & Torri, G. Combined quantitative 1H and 13C nuclear magnetic resonance spectroscopy for characterization of heparin preparations. Semin. Thromb. Hemost. 27, 473–482 (2001).
Manzi, A.E. et al. Exploring the glycan repertoire of genetically modified mice by isolation and profiling of the major glycan classes and nano-NMR analysis of glycan mixtures. Glycobiology 10, 669–689 (2000).
Lopez, M. et al. Microheterogeneity of the oligosaccharides carried by the recombinant bovine lactoferrin expressed in Mamestra brassicae cells. Glycobiology 7, 635–651 (1997).
Venkataraman, G., Shriver, Z., Raman, R. & Sasisekharan, R. Sequencing complex polysaccharides. Science 286, 537–542 (1999).
Guerrini, M. et al. A novel computational approach to integrate NMR spectroscopy and capillary electrophoresis for structure assignment of heparin and heparan sulfate oligosaccharides. Glycobiology 12, 713–719 (2002).
Sharon, N. & Lis, H. Lectins (Kluwer Academic Publishers, Boston, 2004).
Plante, O.J., Palmacci, E.R. & Seeberger, P.H. Automated solid-phase synthesis of oligosaccharides. Science 291, 1523–1527 (2001).
Hirabayashi, J. Lectin-based structural glycomics: glycoproteomics and glycan profiling. Glycoconj. J. 21, 35–40 (2004).
Feizi, T., Fazio, F., Chai, W. & Wong, C.H. Carbohydrate microarrays—a new set of technologies at the frontiers of glycomics. Curr. Opin. Struct. Biol. 13, 637–645 (2003).
Galanina, O.E., Mecklenburg, M., Nifantiev, N.E., Pazynina, G.V. & Bovin, N.V. GlycoChip: multiarray for the study of carbohydrate-binding proteins. Lab Chip 3, 260–265 (2003).
Blixt, O. et al. Printed covalent glycan array for ligand profiling of diverse glycan binding proteins. Proc. Natl. Acad. Sci. USA 101, 17033–17038 (2004).
von der Lieth, C.W., Bohne-Lang, A., Lohmann, K.K. & Frank, M. Bioinformatics for glycomics: status, methods, requirements and perspectives. Brief. Bioinform. 5, 164–178 (2004).
Cooper, C.A., Harrison, M.J., Wilkins, M.R. & Packer, N.H. GlycoSuiteDB: a new curated relational database of glycoprotein glycan structures and their biological sources. Nucleic Acids Res. 29, 332–335 (2001).
Kanehisa, M., Goto, S., Kawashima, S., Okuno, Y. & Hattori, M. The KEGG resource for deciphering the genome. Nucleic Acids Res. 32, D277–D280 (2004).
Aoki, K.F. et al. Application of a new probabilistic model for recognizing complex patterns in glycans. Bioinformatics 20 (Suppl. 1), I6–I14 (2004).
Bohne-Lang, A., Lang, E., Forster, T. & von der Lieth, C.W. LINUCS: linear notation for unique description of carbohydrate sequences. Carbohydr. Res. 336, 1–11 (2001).
Kikuchi, N. et al. The carbohydrate sequence markup language (CabosML): an XML description of carbohydrate structures. Bioinformatics 21, 1717–1718 (2005).
Lutteke, T., Frank, M. & von der Lieth, C.W. Data mining the protein data bank: automatic detection and assignment of carbohydrate structures. Carbohydr. Res. 339, 1015–1020 (2004).
Li, J. et al. The Molecule Pages database. Nature 420, 716–717 (2002).
Acknowledgements
This work was supported by the US National Institute of General Medical Sciences Glue Grant U54 GM62116. We thank the CFG members, the members of Bioinformatics Core (B) of CFG for the work described in this review and V. Sasisekharan for critical reading of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
G.V. is presently employed by Momenta Pharmaceuticals. R.S. is a consultant to Momenta Pharmaceuticals.
Supplementary information
Rights and permissions
About this article
Cite this article
Raman, R., Raguram, S., Venkataraman, G. et al. Glycomics: an integrated systems approach to structure-function relationships of glycans. Nat Methods 2, 817–824 (2005). https://doi.org/10.1038/nmeth807
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth807
This article is cited by
-
Chemoenzymatic synthesis of genetically-encoded multivalent liquid N-glycan arrays
Nature Communications (2023)
-
GproDIA enables data-independent acquisition glycoproteomics with comprehensive statistical control
Nature Communications (2021)
-
Post-synthesis of boric acid–functionalized magnetic covalent organic framework as an affinity probe for the enrichment of N-glycopeptides
Microchimica Acta (2021)
-
Genetically encoded multivalent liquid glycan array displayed on M13 bacteriophage
Nature Chemical Biology (2021)
-
CCMRD: a solid-state NMR database for complex carbohydrates
Journal of Biomolecular NMR (2020)