Abstract
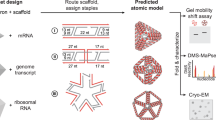
The organization of biological materials into versatile three-dimensional assemblies could be used to build multifunctional therapeutic scaffolds for use in nanomedicine. Here, we report a strategy to design three-dimensional nanoscale scaffolds that can be self-assembled from RNA with precise control over their shape, size and composition. These cubic nanoscaffolds are only ∼13 nm in diameter and are composed of short oligonucleotides, making them amenable to chemical synthesis, point modifications and further functionalization. Nanocube assembly is verified by gel assays, dynamic light scattering and cryogenic electron microscopy. Formation of functional RNA nanocubes is also demonstrated by incorporation of a light-up fluorescent RNA aptamer that is optimally active only upon full RNA assembly. Moreover, we show that the RNA nanoscaffolds can self-assemble in isothermal conditions (37 °C) during in vitro transcription, which opens a route towards the construction of sensors, programmable packaging and cargo delivery systems for biomedical applications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Seeman, N. C. An overview of structural DNA nanotechnology. Mol. Biotechnol. 37, 246–257 (2007).
Aldaye, F. A., Palmer, A. L. & Sleiman, H. F. Assembling materials with DNA as the guide. Science 321, 1795–1799 (2008).
Lin, C., Liu, Y. & Yan, H. Designer DNA nanoarchitectures. Biochemistry 48, 1663–1674 (2009).
Chen, J. H. & Seeman, N. C. The electrophoretic properties of a DNA cube and its substructure catenanes. Electrophoresis 12, 607–611 (1991).
Goodman, R. P. et al. Reconfigurable, braced, three-dimensional DNA nanostructures. Nature Nanotech. 3, 93–96 (2008).
He, Y. et al. Hierarchical self-assembly of DNA into symmetric supramolecular polyhedra. Nature 452, 198–201 (2008).
Erben, C. M., Goodman, R. P. & Turberfield, A. J. A self-assembled DNA bipyramid. J. Am. Chem. Soc. 129, 6992–6993 (2007).
Andersen, F. F. et al. Assembly and structural analysis of a covalently closed nano-scale DNA cage. Nucleic Acids Res. 36, 1113–1119 (2008).
Shih, W. M., Quispe, J. D. & Joyce, G. F. A 1.7-kilobase single-stranded DNA that folds into a nanoscale octahedron. Nature 427, 618–621 (2004).
Zimmermann, J., Cebulla, M. P., Monninghoff, S. & von Kiedrowski, G. Self-assembly of a DNA dodecahedron from 20 trisoligonucleotides with C(3h) linkers. Angew. Chem. Int. Ed. 47, 3626–3630 (2008).
Erben, C. M., Goodman, R. P. & Turberfield, A. J. Single-molecule protein encapsulation in a rigid DNA cage. Angew. Chem. Int. Ed. 45, 7414–7417 (2006).
Bhatia, D. et al. Icosahedral DNA nanocapsules by modular assembly. Angew. Chem. Int. Ed. 48, 4134–4137 (2009).
Rothemund, P. W. Folding DNA to create nanoscale shapes and patterns. Nature 440, 297–302 (2006).
Andersen, E. S. et al. Self-assembly of a nanoscale DNA box with a controllable lid. Nature 459, 73–76 (2009).
Ke, Y. et al. Scaffolded DNA origami of a DNA tetrahedron molecular container. Nano Lett. 9, 2445–2447 (2009).
Dietz, H., Douglas, S. M. & Shih, W. M. Folding DNA into twisted and curved nanoscale shapes. Science 325, 725–730 (2009).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Kim, D. H. & Rossi, J. J. Strategies for silencing human disease using RNA interference. Nature Rev. Genet. 8, 173–184 (2007).
Famulok, M., Hartig, J. S. & Mayer, G. Functional aptamers and aptazymes in biotechnology, diagnostics and therapy. Chem. Rev. 107, 3715–3743 (2007).
Joyce, G. F. Directed evolution of nucleic acid enzymes. Annu. Rev. Biochem. 73, 791–836 (2004).
Davidson, E. A. & Ellington, A. D. Engineering regulatory RNAs. Trends Biotechnol. 23, 109–112 (2005).
Wendell, D. et al. Translocation of double-stranded DNA through membrane-adapted phi29 motor protein nanopores. Nature Nanotech. 4, 765–772 (2009).
Khaled, A., Guo, S., Li, F. & Guo, P. Controllable self-assembly of nanoparticles for specific delivery of multiple therapeutic molecules to cancer cells using RNA nanotechnology. Nano Lett. 5, 1797–1808 (2005).
Guo, P. RNA nanotechnology: engineering, assembly and applications in detection, gene delivery and therapy. J. Nanosci. Nanotechnol. 5, 1964–1982 (2005).
Jaeger, L. & Leontis, N. B. Tecto-RNA: one-dimensional self-assembly through tertiary interactions. Angew. Chem. Int. Ed. 39, 2521–2524 (2000).
Chworos, A. et al. Building programmable jigsaw puzzles with RNA. Science 306, 2068–2072 (2004).
Shu, D., Moll, W.-D., Deng, Z., Mao, M. & Guo, P. Bottom-up assembly of RNA arrays and superstructures as potential parts in nanotechnology. Nano Lett. 4, 1717–1723 (2004).
Jaeger, L. & Chworos, A. The architectonics of programmable RNA and DNA nanostructures. Curr. Opin. Struct. Biol. 16, 531–543 (2006).
Afonin, K. A., Cieply, D. J. & Leontis, N. B. Specific RNA self-assembly with minimal paranemic motifs. J. Am. Chem. Soc. 130, 93–102 (2008).
Severcan, I., Geary, C., Verzemnieks, E., Chworos, A. & Jaeger, L. Square-shaped RNA particles from different RNA folds. Nano Lett. 9, 1270–1277 (2009).
Koyfman, A. Y. et al. Controlled spacing of cationic gold nanoparticles by nanocrown RNA. J. Am. Chem. Soc. 127, 11886–11887 (2005).
Nasalean, L., Baudrey, S., Leontis, N. B. & Jaeger, L. Controlling RNA self-assembly to form filaments. Nucleic Acids Res. 34, 1381–1392 (2006).
Severcan, I. et al. A polyhedron made of tRNAs. Nature Chem. 2, 772–779 (2010).
Bindewald, E., Grunewald, C., Boyle, B., O'Connor, M. & Shapiro, B. A. Computational strategies for the automated design of RNA nanoscale structures from building blocks using NanoTiler. J. Mol. Graph. Model 27, 299–308 (2008).
Seiffert, J. & Huhle, A. A full-automatic sequence design algorithm for branched DNA structures. J. Biomol. Struct. Dyn. 25, 453–466 (2008).
Seeman, N. C. Nucleic acid junctions and lattices. J. Theor. Biol. 99, 237–247 (1982).
Mathews, D. H., Sabina, J., Zuker, M. & Turner, D. H. Expanded sequence dependence of thermodynamic parameters improves prediction of RNA secondary structure. J. Mol. Biol. 288, 911–940 (1999).
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Bernhart, S. H. et al. Partition function and base pairing probabilities of RNA heterodimers. Algorithms Mol. Biol. 1, 3 (2006).
Hofacker, I. L. Fast folding and comparison of RNA secondary structures. Monatshefte f Chemie 125, 167–188 (1994).
Freier, S. M. et al. Improved free-energy parameters for predictions of RNA duplex stability. Proc. Natl Acad. Sci. USA 83, 9373–9377 (1986).
SantaLucia, J. Jr. A unified view of polymer, dumbbell and oligonucleotide DNA nearest-neighbor thermodynamics. Proc. Natl Acad. Sci. USA 95, 1460–1465 (1998).
Sugimoto, N., Nakano, S., Yoneyama, M. & Honda, K. Improved thermodynamic parameters and helix initiation factor to predict stability of DNA duplexes. Nucleic Acids Res. 24, 4501–4505 (1996).
Kato, T., Goodman, R. P., Erben, C. M., Turberfield, A. J. & Namba, K. High-resolution structural analysis of a DNA nanostructure by cryoEM. Nano Lett. 9, 2747–2750 (2009).
Ludtke, S. J., Baldwin, P. R. & Chiu, W. EMAN: semiautomated software for high-resolution single-particle reconstructions. J. Struct. Biol. 128, 82–97 (1999).
Baugh, C., Grate, D. & Wilson, C. 2.8 Å crystal structure of the malachite green aptamer. J. Mol. Biol. 301, 117–128 (2000).
Duxbury, D. F. The photochemistry and photophysics of triphenylmethane dyes in solid and liquid media. Chem. Rev. 93, 381–433 (1993).
Afonin, K. A., Danilov, E. O., Novikova, I. V. & Leontis, N. B. TokenRNA: a new type of sequence-specific, label-free fluorescent biosensor for folded RNA molecules. Chembiochem 9, 1902–1905 (2008).
Marky, L. A. & Breslauer, K. J. Calculating thermodynamic data for transitions of any molecularity from equilibrium melting curves. Biopolymers 26, 1601–1620 (1987).
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005).
Lander, G. C. et al. Appion: an integrated, database-driven pipeline to facilitate EM image processing. J. Struct. Biol. 166, 95–102 (2009).
Acknowledgements
The authors thank V. A. Piunova for assistance with DLS, K. Kahn for helping with Kd curve analysis, and C. Potter and B. Carragher for their invaluable scientific input regarding cryo-EM and single-particle reconstruction. Cryo-EM imaging was performed at National Resources for Automated Molecular Microscopy, which is supported by the National Institutes of Health (NIH) through the National Center for Research Resources P41 program (RR17573). This research was supported (in part) by the Intramural Research Program of the NIH, the National Cancer Institute, the Center for Cancer Research (B.A.S.) and by NIH R01 GM079604 (L.J.). This project has been funded in whole or in part with federal funds from the National Cancer Institute, NIH, under contract HHSN261200800001E. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does any mention of trade names, commercial products or organizations imply endorsement by the US government. K.A. and L.J. wish to dedicate this work to Sts. Cyril and Methodius.
Author information
Authors and Affiliations
Contributions
K.A.A. and L.J. conceived and designed the experiments. E.B., B.A.S., K.A.A. and L.J. contributed to the sequence and 3D model design. K.A.A. and A.J.Y. performed self-assembly PAGE and fluorescence experiments. K.A.A. performed DLS experiments. K.A.A. and L.J. analysed the data. N.V. and E.J. performed cryo-EM characterization and reconstruction. K.A.A., B.A.S. and L.J. co-wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 2964 kb)
Rights and permissions
About this article
Cite this article
Afonin, K., Bindewald, E., Yaghoubian, A. et al. In vitro assembly of cubic RNA-based scaffolds designed in silico. Nature Nanotech 5, 676–682 (2010). https://doi.org/10.1038/nnano.2010.160
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nnano.2010.160
This article is cited by
-
Self-assembling biomolecules for biosensor applications
Biomaterials Research (2023)
-
Structure, folding and flexibility of co-transcriptional RNA origami
Nature Nanotechnology (2023)
-
Nucleic acid-based scaffold systems and application in enzyme cascade catalysis
Applied Microbiology and Biotechnology (2023)