-
PDF
- Split View
-
Views
-
Cite
Cite
Monika S. Doblin, Isaac Kurek, Deborah Jacob-Wilk, Deborah P. Delmer, Cellulose Biosynthesis in Plants: from Genes to Rosettes, Plant and Cell Physiology, Volume 43, Issue 12, 15 December 2002, Pages 1407–1420, https://doi.org/10.1093/pcp/pcf164
Close - Share Icon Share
Abstract
Modern techniques of gene cloning have identified the CesA genes as encoding the probable catalytic subunits of the plant CelS, the cellulose synthase enzyme complex visualized in the plasma membrane as rosettes. At least 10 CesA isoforms exist in Arabidopsis and have been shown by mutant analyses to play distinct role/s in the cellulose synthesis process. Functional specialization within this family includes differences in gene expression, regulation and, possibly, catalytic function. Current data points towards some CesA isoforms potentially being responsible for initiation or elongation of the recently identified sterol β-glucoside primer within different cell types, e.g. those undergoing either primary or secondary wall cellulose synthesis. Different CesA isoforms may also play distinct roles within the rosette, and there is some circumstantial evidence that CesA genes may encode the catalytic subunit of the mixed linkage glucan synthase or callose synthase. Various other proteins such as the Korrigan endocellulase, sucrose synthase, cytoskeletal components, Rac13, redox proteins and a lipid transfer protein have been implicated to be involved in synthesizing cellulose but, apart from CesAs, only Korrigan has been definitively linked with cellulose synthesis. These proteins should prove valuable in identifying additional CelS components.
(Received April 22, 2002; Accepted July 9, 2002)
Introduction
Understanding the biosynthesis of wall polysaccharide components has attracted considerable interest in light of the fundamental importance of these molecules not just to plant function, but also to man. Unfortunately, very little is known of the mechanism(s) and regulation of the biosynthetic steps that control polysaccharide biosynthesis, deposition and assembly, or the interaction of these components to provide cells with a functional wall. Furthermore, manipulation of polysaccharide quantity and quality has been hampered by the lack of cloned genes for plant glycosyltransferases (GTs). As recently as 1995, not one single enzyme involved in plant cell wall biosynthesis had been purified to homogeneity, nor had a single gene coding for such an enzyme been identified and cloned. Fortunately, since that time, a number of GTs have been cloned using traditional biochemical or, more modern, in silico, molecular and genetic techniques. By far the most significant of these has been the cloning of the CesA genes of cotton and Arabidopsis, presumed to encode catalytic subunits of cellulose synthase (CelS), the enzyme complex responsible for the synthesis of cellulose. [We refer to CelS as the entire synthase complex, and will use the accepted term CesA when referring just to the catalytic subunits within that complex.] Identification of these genes has led to remarkable progress in the field, and this review will focus on this recent work. Due to space limitations, we will not cover very interesting recent work on cellulose synthesis in bacteria (Ausmees et al. 1999, Ausmees et al. 2001, Nakai et al. 1999, Römling et al. 2000, Zogaj et al. 2001). The reader is also referred to a number of reviews that give the background on cellulose synthesis (Brown 1996, Delmer 1999, Brown and Saxena 2000, Saxena and Brown 2000, Richmond and Somerville 2000, Dhugga 2001). However, for the general reader, a few words about our current understanding of the nature of the CelS complex may be helpful and are provided below.
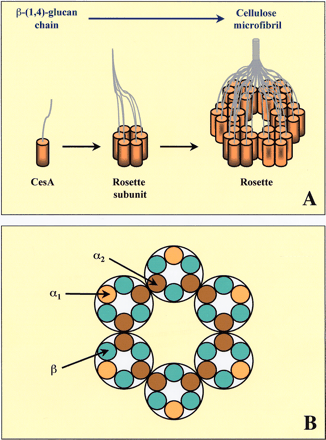
Structures responsible for cellulose synthesis have been identified by electron microscopy in freeze-fractured plasma membranes of many organisms (Brown 1996, Kimura et al. 1999a). Linearly arranged terminal complexes (TCs) in single or multiple rows are observed in bacteria, D. discoideum and some algae, or hexagonal structures with six-fold symmetry, termed rosettes, are observed in mosses, ferns, algae and vascular plants (Brown 1996, Delmer 1999, Tsekos 1999). Although TCs and rosettes reside in the plasma membrane, Haigler and Brown (1986) revealed that during synthesis, rosettes are assembled in the Golgi and then transported to the plasma membrane. Biochemical studies indicate that the higher plant CelS complex is a large (>500 kDa), integral membrane, multi-subunit enzyme utilising uridine diphospho-glucose (UDP-Glc) as substrate (Delmer 1999). Assumed to be included within each complex are a specific number of obligatory CesA catalytic subunits that utilize UDP-Glc as substrate for glucan chain elongation, as well as other components that may be involved in providing the substrate, in initiating or terminating chain elongation or that may be involved in regulating the activity of the complex. Based on our current understanding of synthesis, UDP-Glc is thought to bind to an active site on the cytoplasmic face of the plasma membrane with the polysaccharide being extruded through the membrane, presumably through a pore-type structure, into the wall (Delmer 1999, Brown and Saxena 2000). Each rosette (~25 nm in diameter comprising six subunits) is believed to contain a number of synthetic units, possibly six per subunit, each of which polymerizes a glucan chain that associates with adjacent chains of the same rosette to form elementary microfibrils (Fig. 1A).
Identification of CesA Genes
The road towards identifying components of the plant CelS complex has been a long and arduous one primarily because of the inability to assay this enzyme in vitro despite high levels of CelS activity in vivo. When incubated with UDP-Glc, isolated cell membranes from numerous plant species produce limited amounts of (1,4)-β-glucan, with the predominant product instead being callose, a linear (1,3)-β-linked polymer of D-Glc with occasional (1,6)-β-linked branches, that is normally found in only small amounts in specialized cell-types and wall structures (Delmer 1987, Delmer 1999). Rosette disassembly occurs concomitantly with the loss of CelS activity upon cell rupture. Studies on the stability of the rosettes within the plasma membrane of Funaria hygrometrica have shown that rosette number is reduced after 4–5 min in the presence of the vesicle transport inhibitors monensin and cytochalasin (Rudolph et al. 1989), indicating that rosettes have a short half-life and are subject to rapid degradation. Thus, early attempts to purify the enzyme were, and continue to be, severely hampered.
Due to these difficulties, researchers looked towards simpler systems, notably the bacterium Acetobacter xylinum, to gain insight into the mechanism of cellulose synthesis. A cellulose-synthesis operon [bcsA–D, later renamed AxCesA1-D1 by Delmer (1999)] was identified by genetic complementation of A. xylinum insertion mutants lacking CelS activity and by purification and partial sequencing of the AxCesA1 catalytic subunit (reviewed by Ross et al. 1991). These advances in the bacterial cellulose synthesis field did not, however, lead to the immediate identification of an homologous plant gene. Rather, these plant genes were identified using a purely molecular approach that targeted a plant tissue highly enriched in cellulose, the cotton fiber, wherein two cDNA clones (CelA1 and CelA2, renamed GhCesA1 and GhCesA2) were identified (Pear et al. 1996). The full-length ORF of GhCesA1 encodes a 974 amino-acid polypeptide of ~110 kDa and, like the bacterial CesA proteins, is predicted to be a membrane-bound protein with eight transmembrane helices (TMHs), two at the N-terminus and six at the C-terminus, that border a central, cytoplasmic domain. However, GhCesA1 and GhCesA2 differ from the bacterial CesAs in that they contain two large “plant-specific” insertions within the central domain: one a conserved “plant conserved region” (CR-P) and the other, a “hypervariable region” (HVR) (Pear et al. 1996, Delmer 1999). Furthermore, the plant CesA proteins have an extended N-terminal region and a shorter C-terminal region in comparison with the bacterial CesAs.
The identification of the cotton CesA genes as being homologous to the bacterial CesA genes and, thus, encoding potential catalytic subunits of CelS, was based on three lines of evidence. First, the encoded protein sequences contained three regions within the central domain that had a low (50–60%) level of similarity to the bacterial CesA proteins. Overall, DNA identity is less than 30%, which accounts for the lack of detection of plant homologues in earlier heterologous screens. Furthermore, included in these regions were small and more highly conserved regions that contained the “D,D,D,QXXRW” motif (Saxena et al. 1995). Enzymes having this signature have since been classified as members of the glycosyltransferase family 2 (GT family 2) (Campbell et al. 1997, Henrissat et al. 2001; http://afmb.cnrs-mrs.fr/~cazy/CAZY/index.html). The critical nature of the amino acids comprising the D,D,D,QXXRW motif for substrate binding and catalysis has been shown experimentally by numerous site-directed mutagenesis studies and by resolution of the crystal structure of a member of GT family 2 (Nagahashi et al. 1995, Saxena et al. 1995, Saxena et al. 2001, Saxena and Brown 1997, Saxena and Brown 2000, Charnock and Davies 1999, Charnock et al. 2001). The D,D,D,QXXRW motif has now become a robust sequence characteristic of the polysaccharide synthase members of this GT family, examples of which include all known chitin synthases, hyaluronan synthases and cellulose synthases. Secondly, Pear et al. (1996) demonstrated that the DNA segment encoding the central region of GhCesA1 bound the substrate UDP-Glc; binding did not occur under the same conditions when a fusion protein with the region containing the first conserved D residue was deleted. Thirdly, expression of GhCesA1 and GhCesA2 correlated with the timing of cellulose biosynthesis. Northern blot analysis showed that their expression was lowest during the stage of primary wall deposition in the fiber, but rose to much higher levels in the transition stage from primary to secondary wall synthesis at ~17 days post-anthesis (dpa) and reaching maximal levels at 24 dpa, when the rate of cellulose synthesis is highest in vivo. Taken together, these data point towards the cotton CesA genes encoding functional homologues of the bacterial CesA genes, and thus the presumed CelS catalytic subunits.
Experimental evidence for this proposed role has now been provided by the isolation of a number of cellulose-deficient mutants in Arabidopsisthaliana, the genetic lesions for which have been shown to be in CesA genes (AtCesA1: rsw1, Arioli et al. 1998a; AtCesA3: ixr1, Scheible et al. 2001; AtCesA4: irx5, Taylor and Turner 2001; AtCesA6: procuste, Fagard et al. 2000; ixr2, Scheible et al. 2001, Desprez et al. 2002; AtCesA7: irx3, Taylor et al. 1999; AtCesA8: irx1, Taylor et al. 2000). In addition, the disassembly of rosettes within the rsw1 mutant (Arioli et al. 1998a), the significantly lower number of rosettes (20% compared with wild type) in a cellulose-deficient brittle culm mutant line of barley (Kimura et al. 1999b), and the specific labelling of these structures by polyclonal antisera directed towards a recombinant cotton CesA polypeptide (Kimura et al. 1999a), confirms that the rosettes observed in freeze-fracture experiments are the sites of cellulose synthesis in plants.
Discovery of the CesA/Csl superfamily
Sequence database searches with the cotton CesA genes revealed that a relatively large number of Arabidopsis and other plant sequences exist that are related to different extents to CesA (Richmond and Somerville 2000, Richmond and Somerville 2001, Hazen et al. 2002). In Arabidopsis, these genes have been divided into the “true” CesA family and six distinct groups of cellulose synthase-like (Csl) genes – the CslA, B, C, D, E and G families (Richmond and Somerville 2001; http://cellwall.stanford.edu/). Each Csl family is characterized by specific sequence features, but all members contain the D,D,D,QXXRW motif and belong to GT family 2 (http://afmb.cnrs-mrs.fr/~cazy/CAZY/index.html). Recent completion of the Arabidopsis genome sequencing project has revealed the presence of at least 10, but possibly 12, CesA genes (a singleton EST and genome survey sequence do not match any of the other 10 CesA sequences; http://cellwall.stanford.edu/). The Arabidopsis CslA, B, C, D, E and G families comprise 9, 6, 5, 6, 1 and 3 members, respectively, making a total of 30 genes.
A comprehensive survey of the almost completed rice (Oryza sativa) genome has revealed that monocots as well as dicots possess multiple CesA and Csl family members (Hazen et al. 2002). Interestingly, no rice CslB or CslG genes have been identified to date. Instead, rice appears to encode several members of two new Csl families, CslF and CslH. The CslF family is most closely related to the CslDs, and the CslH family to the CslBs (Hazen et al. 2002). At present, the rice CslA, C, D, E, F and H families encode 10, 9, 4, 5, 7 and 2 members, respectively, which is 37 genes in total, but these numbers are expected to increase upon completion of genome sequencing.
Origin and evolution of the plant CesA/Csl supergene family
Recent data suggest that initially, CesA and Csl-type genes may have originated from endosymbiotic transfer from cyanobacteria to plants (Nobles et al. 2001). A specific search of cyanobacterial genomes for sequences containing a D,D,D,QXXRW motif revealed CesA and Csl-type sequences in Anabaena sp. Pasteur Culture Collection (PCC) 7120 and Nostoc punctiforme (Nobles et al. 2001). The occurrence of cellulose in these strains, as evidenced by X-ray diffraction, electron microscopy of microfibrils and immuno-gold labeling with cellobiohydrolase I, an enzyme that specifically binds cellulose, provides good correlatory evidence that the CesA-like genes are involved in cellulose biosynthesis (Nobles et al. 2001). Phylogenetic analyses placed the cyanobacterial protein sequences in three distinct clades, the first containing vascular plant CesAs, a second containing CesA genes from gram-negative bacteria, and a third clade containing a single Bacillus subtilis sequence with a D,D,D,QXXRW motif (Nobles et al. 2001). The Anabaena and Nostoc proteins group with the plant CesA sequences largely because of the presence of a CR-P-type insertion between the first and second conserved D residues, a sequence feature observed only in the plant CesAs and the single D. discoideum CesA protein, not in bacterial CesA proteins (Delmer 1999, Blanton et al. 2000). That the cyanobacterial insertions are more similar to the plant CR-P region than to the D. discoideum insertion indicates a distinct phylogenetic relationship between the cyanobacterial and vascular plant CesAs. This situation closely resembles the relationship of 16S rRNAs from cyanobacteria and chloroplasts, suggesting that vascular plant CesAs originated in the cyanobacterial symbiont and were subsequently transferred to the host plant nuclear genome, an occurrence evidenced by numerous other nuclear-encoded plant genes (Nobles et al. 2001, Rujan and Martin 2001). By analogy, the other cyanobacterial genes that group with either the bacterial CesAs or the D,D,D,QXXRW motif-containing B. subtilis sequence may also have undergone similar horizontal gene transfer events. Plant Csl families A and C are most similar to the bacterial CesAs (Richmond and Somerville 2001; http://cellwall.stanford.edu/). It is therefore tempting to speculate that the cyanobacterial sequences grouping with the bacterial CesAs evolved into the plant Csls, and the cyanobacterial sequences grouping with the B. subtilis sequence are the progenitors of other types of family 2 GTs.
Such an hypothesis of plant CesA and Csl origin is impossible to prove but, for it to be true, consistencies must be found within the context of our current understanding of plant evolution. For example, it predicts that taxa further down the evolutionary ladder than higher plants, such as ferns, mosses and algae, should have examples of both CesA and Csl genes and that their CesA proteins carry a plant CR-P-type insertion rather than one more similar to D. discoideum. Elucidation of a partial CesA sequence isolated from the unicellular alga Mesotaenium caldariourum UTEX 41 is consistent with the second of these predictions, as it contains a plant CR-P-type insertion (Roberts et al. 2000). Furthermore, phylogenetic analyses using this sequence demonstrated not only the expected close relationship of the algal sequence to vascular plant CesAs, but also a branch point closer to the cyanobacterial/vascular plant divergence point (Nobles et al. 2001). Obviously, the characterization of additional CesA genes from other lineages basal to the higher plants will be necessary to add strength to the cyanobacterial origin hypothesis.
Regardless of whether the hypothesis of a cyanobacterial origin of the CesA/Csl superfamily is true, several major genome evolution events such as segmental or complete genome duplications within ancestral species would need to have occurred to account for the large numbers of CesA/Csl genes that we see in higher plant species today. That such events have taken place has been documented within many plant species (Wendel 2000, Schmidt 2002) and has been specifically demonstrated for the CesA genes of cotton and maize (Cronn et al. 1999, Holland et al. 2000). For example, analysis of map location of maize CesAs using restriction-fragment-length polymorphisms determined that closely related CesAs (i.e. those found to pair in sequence cladograms) were found at different chromosomal locations that reflect ancient duplications (Holland et al. 2000).
There appears to have been an enormous diversification of the CesA/Csl gene superfamily within the plant lineage in comparison to other cellulose-synthesising organisms such as bacteria and fungi. Two major questions arise from the discovery of such a large number of CesA and Csl genes in plants. First, why are there so many Csl gene families? Unfortunately, no explicit function has yet been assigned to any of the Csl genes to help answer this question, but the commonly held view is that the Csl genes are involved in the synthesis of the many other non-cellulosic polysaccharides found in plant cell walls and in secretions such as root cap or stylar mucilage (Cutler and Somerville 1997, Arioli et al. 1998b, Delmer 1999, Richmond and Somerville 2000, Richmond and Somerville 2001, Vergara and Carpita 2001). In other words, the sequence divergence of the Csl genes is thought to reflect their functional divergence. Since all the Csl polypeptides have the D,D,D,QXXRW motif and have been classified as family 2 GTs, it is possible that they are exclusively involved in the synthesis of β-linked polysaccharides such as callose, xyloglucan, glucuronoarabinoxylan, mixed-linkage (1→3)(1→4)-β-D-glucan, glucomannan, galactomannan or galactoglucomannan, and possibly cellulose. A role for CesA/Csls in the synthesis of α-linked polysaccharides, such as the pectic components homogalacturonan, rhamnogalacturonan I and rhamnogalacturonan II, is not currently favoured because of their GT family 2 classification. Nevertheless, such a possibility cannot be discounted at present, since it is not yet known what sequence features of the GT family 2 members control substrate and/or enzymatic specificity.
The second question is why are there so many members of each of the CesA and Csl gene families within extant plant species? Is it merely a consequence of the multiple gene duplication events that have occurred throughout plant genome evolution? Gene duplications are predicted to initially produce functionally redundant genes which can then further evolve over time by mutation. It appears that having multiple functional copies of CesA and Csl genes has been selectively advantageous to plants, since no CesA or Csl pseudogenes have been detected within the Arabidopsis genome. It can therefore be envisaged then that despite their sequence differences, gene family members could encode functionally identical isoforms that are interchangeable and thus capable of compensating for the loss of another isoform if mutated. Alternatively, plant genomes may contain multiple CesA and Csl genes because either some or all of the encoded isoforms perform distinct functions. It is generally regarded that gene duplications allow family members to evolve a unique function, either through having a different cell-type-specific expression pattern, regulation or possibly an altered or additional activity, or a combination of these. An altered catalytic function could mean that some family members participate in synthesis of different polysaccharides, but catalytic differences may also be more subtle. For example, isoforms may utilize the same donor but a different acceptor molecule in the synthesis of the same polysaccharide. Thus, having multiple genes may be a requirement for synthesis of some types of plant polysaccharides. At least in the case of the CesA gene family, the necessity for multiple GT genes is not seen in other organisms: one CesA isoform is clearly sufficient for synthesis of cellulose within D. discoideum and various bacterial species. A possible explanation for this is that cellulose synthesis in plants occurs within the context of rosettes rather than single or multiple row TCs. More than one CesA isoform may therefore be required for rosette assembly.
Phylogenetic analyses using CesA and Csl sequences derived from various plant species have shown that orthologous genes (genes of different species) are more similar than paralogous genes (genes of the same species), implying that CesA/Csl genes diverged in sequence before the divergence of the monocot and dicot lineages (Fagard et al. 2000, Holland et al. 2000, Doblin et al. 2001, Vergara and Carpita 2001; http://cellwall.stanford.edu/). This implies that the role/s that paralogous CesA/Csl genes play must have been established early in higher plant evolution. What the roles are of the CesA family members is discussed in more detail in the following section.
Evidence for a specialized role for different CesA family members
(a) CesA expression and regulation
The currently available data addressing CesA function indicate that significant functional specialization has occurred within this gene family. There is evidence for all types of functional differences discussed above – gene expression, regulation and catalytic function. Various types of CesA expression analyses (standard and electronic Northern analysis, in situ hybridization, RT-PCR, tissue prints, massively parallel signature sequencing, microarrays and promoter–reporter gene fusions) have been conducted in both monocot and dicot species (cotton, Pear et al. 1996; Populus sp., Wu et al. 2000; maize, Holland et al. 2000, Dhugga 2001; rice, Vergara and Carpita 2001; tobacco, Doblin et al. 2001; Arabidopsis, Taylor et al. 1999, Fagard et al. 2000, Richmond and Somerville 2001, M. Doblin, R. Eshed, P. Hogan, D. Jacob-Wilk, A. Roberts, J. Schultz and D. Delmer, unpublished data). Most studies have been conducted in Arabidopsis and these have yielded a number of generalizations regarding CesA expression. First, CesA family members are expressed to varying levels. For example, electronic Northern analysis of Arabidopsis ESTs indicates that AtCesA1 (RSW1) is much more highly expressed than AtCesA9 (currently represented by 38 and one EST, respectively; http://cellwall.stanford.edu). Promoter–GUS analyses have shown that this is, not only, because AtCesA1is expressed in many more cells and cell-types than AtCesA9, but also, because it is more highly expressed in some of these cells (M. Doblin, R. Eshed, P. Hogan, D. Jacob-Wilk, A. Roberts, J. Schultz and D. Delmer, unpublished data). In contrast, the expression of AtCesA7 (IRX3) is limited to the xylem but appears to be expressed to a similar level in that cell type compared with the cells in which AtCesA1 is expressed (Turner and Somerville 1997, Taylor et al. 1999, M. Doblin, R. Eshed, P. Hogan, D. Jacob-Wilk, A. Roberts, J. Schultz and D. Delmer, unpublished data). Secondly, CesAgenes are expressed in multiple tissues that relate to their expression in specific cell types common to these tissues. Epidermal cells of all plant organs are affected in the AtCesA1 mutant rsw1, for example (Arioli et al. 1998a, Williamson et al. 2001a). Similarly, the AtCesA7 mutant irx3shows a collapsed xylem phenotype throughout the plant, within leaves, hypocotyls, stem and root (Turner and Somerville 1997, Taylor et al. 1999, M. Doblin, R. Eshed, P. Hogan, D. Jacob-Wilk, A. Roberts, J. Schultz and D. Delmer, unpublished data). Thirdly, multiple CesA genes are expressed in the same cell-type. AtCesAs 1, 3 and 6 all exhibit a very similar expression pattern, being expressed in cells undergoing expansion in tissues such as the root and hypocotyl among others (Arioli et al. 1998a, Fagard et al. 2000, Scheible et al. 2001, Williamson et al. 2001a, M. Doblin, R. Eshed, P. Hogan, D. Jacob-Wilk, A. Roberts, J. Schultz and D. Delmer, unpublished data). AtCesAs 4, 7 and 8 also exhibit a similar overlapping expression pattern, being expressed only within xylem cells (Holland et al. 2000, Taylor et al. 2000, Taylor and Turner 2001, M. Doblin, R. Eshed, P. Hogan, D. Jacob-Wilk, A. Roberts, J. Schultz and D. Delmer, unpublished data). AtCesAs 4, 7 and 8 are therefore thought to be involved in secondary wall cellulose synthesis and AtCesAs 1, 3 and 6, in the formation of cellulose in the primary wall.
Preliminary microarray analyses in Arabidopsis have also indicated that CesA gene expression is differentially regulated (Richmond and Somerville 2001). Comparison of transcript levels in leaves versus etiolated seedlings has shown that expression of AtCesAs2, 3 and 5 is higher in leaves than in etiolated seedlings, whereas transcript levels for all the other CesA genes used in the analysis were the same (AtCesA1) or lower (AtCesAs 4, 6 and 7; Richmond and Somerville 2001). That AtCesAs 1, 3 and 6 do not show similar levels of transcript even though they are expressed in the same spatial pattern indicates that each gene’s expression is regulated by different factors. In contrast, the similar results obtained with AtCesAs 4 and 7 support the notion that these two genes may be coordinately regulated. Regulation of the CesA genes is likely to be highly complex in light of other microarray results obtained with various treatments. AtCesA1, for example, is the only gene of four (AtCesAs 1, 3, 4, 6) that is down-regulated three-fold on treatment of light-grown seedlings by salt stress, whereas a five-fold increase in AtCesA1 and AtCesA3 transcripts is observed in plants treated with ethylene in the light (Richmond and Somerville 2001).
Whilst CesA genes are clearly modulated at the transcriptional level, there is some evidence indicating that post-transcriptional regulation is also likely to be important. First, an epitope-tagged version of GhCesA1 expressed in Arabidopsis and tobacco under the control of the CaMV35S promoter was unable to be detected on Western blots of membrane proteins in spite of high transcript levels (Delmer 1999). It was speculated that this was due to the steady-state level of CesA protein being low and/or that it is subject to rapid degradation: CesA protein might be targeted for fast removal if present in excess of the normal in vivo stoichiometric ratio of CelS components. It is possible that the functional CesA protein was not coextracted with the plasma membrane fraction because it remained associated with glucan chains or aggregated artefactually in vitro. From another study of CesA expression, it appears that the former explanation is more likely. Nakagawa and Sakurai (1998) were not able to detect CesA protein in membrane fractions of tobacco BY2 cells using a generic polyclonal antibody generated towards a conserved plant CesA peptide near the third conserved D residue. Rather, they were only able to detect CesA protein after treatment with the cellulose synthesis inhibitor 2,6-dichlorobenzonitrile (DCB; Montezinos and Delmer 1980), suggesting that DCB may cause an elevation in the steady-state level of CesA, possibly by stabilising the CesA protein in some manner (Nakagawa and Sakurai 1998). Our recent experiments following CesA degradation in cultured ovules estimate the half-life of GhCesA1 to be ~30 min (I. Kurek and D. Delmer, unpublished data). Thus, a high rate of CesA turnover appears to contribute to the low levels of CesA protein observed in plant cells in vivo. Analyses at both the RNA and protein level will therefore be required to determine what factors are responsible for the modulation of CesA expression within different plant tissues in response to differing physiological and environmental conditions. This will necessitate probe and antibody generation for the specific detection of CesAs, a considerable feat given the relatively large numbers of CesA family members within a plant species.
Several questions remain regarding CesA function. Are all CesA isoforms functionally unique or do some isoforms function in an analogous manner? Phylogenetic analyses tend to group CesA proteins known to be involved in primary wall formation separately from those involved in secondary wall formation (Fagard et al. 2000, Holland et al. 2000, Vergara and Carpita 2001; http://cellwall.stanford.edu/) indicating that CesAs may not be functionally analogous. These results are supported by experiments with promoter–GUS fusions in Arabidopsis where AtCesAs 1, 2, 3, 5 and 6 are all expressed in young expanding tissues, whereas AtCesAs4, 7 and 8 are expressed only in vascular tissues (Holland et al. 2000, M. Doblin, R. Eshed, P. Hogan, D. Jacob-Wilk, A. Roberts, J. Schultz and D. Delmer, unpublished data). These results also relate to the additional question of how many different CesA isoforms are required for cellulose production within a cell. Promoter-GUS fusion experiments indicate that up to five isoforms (AtCesA1, 2, 3, 5 and 6) are likely to be expressed within the same cell types undergoing primary wall synthesis and that AtCesA1, 3 and 6, in addition to AtCesA4, 7 and 8, are expressed in developing vascular tissues. However, only two CesAs, AtCesA9 and AtCesA10, are expressed in cells at the base of the rosette leaves at the site of stem inflorescence attachment (M. Doblin, R. Eshed, P. Hogan, D. Jacob-Wilk, A. Roberts, J. Schultz and D. Delmer, unpublished data).
(b) Rosette assembly
The observation that multiple CesA genes have overlapping expression patterns may mean that these CesAs are functionally redundant and thus play identical roles in cellulose biosynthesis. However, mutant analysis indicates that this is not the case. All six of the single mutants of ArabidopsisCesA genes described earlier have observable phenotypes that relate to an impairment in cellulose production, despite at least two other active CesA isoforms being co-expressed in the same cells. If different CesA isoforms are randomly assembled into rosettes, as would be predicted if they are entirely functionally identical, then defective components are likely to affect the ability of a rosette to function normally, leading to reduced cellulose production (Taylor et al. 2000, Perrin 2001). Such mutations could therefore be expected to be dominant or semi-dominant, i.e., only one mutant gene copy would be required for a cellulose-deficient phenotype to be observed, on the proviso that at least some of the mutant protein is incorporated into rosettes and that CesA turnover is slow enough for a reduction in synthesis rate to be manifested. In both rsw1 and procustemutants, it is only when the mutation is in the homozygous state not the heterozygous state that an impairment in cellulose production is observed (Arioli et al. 1998a, Fagard et al. 2000). Furthermore, if these CesA isoforms are interchangeable, then the total CesA pool size should be the limiting factor in cellulose production, rather than the amount of each CesA isoform. Plants transheterozygous for the rsw1 and procustemutations do not show a severe cellulose-deficient phenotype and thus are not equivalent to the homozygous mutants. In addition, a failure to complement the rsw1 mutant upon introduction of a wild-type copy of AtCesA3 (Burn et al. 2002) provides more robust proof that another primary CesA isoform cannot effectively compensate for the loss of another. This strongly suggests that the presence of each of these CesA isoforms is critical for cellulose synthesis, and argues a case for a non-random incorporation of CesA proteins into rosettes (Fagard et al. 2000). A similar situation is observed in the secondary wall CesA mutants. Only homozygous and not heterozygous irx1, irx3 and irx5 mutants have a cellulose-deficient phenotype implying that AtCesA8, AtCesA7 and AtCesA4, respectively, also have a role in cellulose synthesis that is distinct from the other two isoforms (Taylor et al. 2000, Taylor and Turner 2001). Therefore, at least three primary wall and three secondary wall CesA isoforms are not functionally redundant but, rather, play distinct roles in the cellulose biosynthesis mechanism.
It seems certain that more than one CesA isoform is required per cell on the basis of the mutant analyses. But whilst two or more isoforms may be required for normal levels of cellulose production within each cell, this does not necessarily mean that all of them are assembled into the same rosette (Perrin 2001). CesA proteins do, nonetheless, appear to be essential for the assembly process, as evidenced by the observed disruption of rosettes in rsw1 (Arioli et al. 1998a). Some evidence has accumulated indicating that at least two CesA isoforms exist within the same rosette and, furthermore, that their association within a rosette is via a direct interaction. Taylor et al. (2000) used a His-tagged version of AtCesA7 to complement irx3 and showed that binding of AtCesA8 protein to the nickel matrix was dependent on the epitope-tagged version of AtCesA7, providing some evidence for a direct interaction between these two proteins.
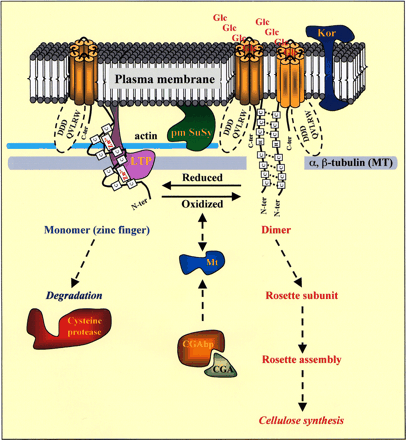
Other data suggest that the in vivo association of CesA polypeptides is mediated by the N-terminal portion of the CesA protein. All plant CesAs identified to date contain two putative zinc fingers at their N-terminus that show high similarity to the RING finger motif known to mediate protein–protein interactions (Delmer 1999, http://cellwall.stanford.edu/). Yeast two-hybrid experiments have shown that the zinc-finger domain of GhCesA1 is able to interact with itself to form homodimers or heterodimers with the zinc-finger domain of GhCesA2 in a redox-dependent manner (Kurek et al. 2002). Bands on native PAGE gels corresponding to the size of CesA monomer, dimer and tetramer were detected by a zinc-finger GhCesA1 polyclonal antibody in membrane extracts of yeast cells expressing the full-length GhCesA1 cDNA, suggesting that higher-order assembly of CesA isoforms occurs under oxidative conditions (Fig. 2; Kurek et al. 2002). These results have led us to consider a model in which dimerization of CesA subunits represents the first step in rosette assembly, with the functional specificity of rosettes (e.g. primary wall versus secondary wall synthesis) being determined by the varying combinations of CesA homo- and/or heterodimers. The disrupted-rosette phenotype of the rsw1 mutant at non-permissive temperatures suggests that different parts of the CesA protein also play a role in rosette assembly. The rsw1 mutant allele encodes a V→A-549 substitution within the cytosolic catalytic domain of the protein (Arioli et al. 1998a). Taken together, these data suggest that the zinc-binding domain is essential for dimerization but different CesA protein regions are also important for tetramerization and rosette formation. A third possible site of interaction of CesA subunits may be via transmembrane domains that form β-sheets, similar to the model of regulation of the outer membrane phospholipase A (OMPLA) of Escherichia coli by reverse dimerization (Saxena and Brown 2000).
If we consider rosette assembly purely from a theoretical perspective, at least two different types of interaction can be envisaged, one between CesA polypeptides within a rosette subunit and one between rosette subunits (Fig. 1B). If rosettes spontaneously self-assemble via interaction between CesA polypeptides alone, as in the model proposed by Scheible et al. (2001), at least two CesA isoforms, α and β, are required. In their model, Scheible et al. (2001) make no distinction between the α isoforms that interact with two β isoforms (α1) and those that interact with an α isoform as well two β isoforms (α2, Fig. 1B). For spontaneous self-assembly, the α2–α2 binding site would need to be “blocked” in some manner for a rosette rather than a linear or multiple linear TC array to be assembled. Alternatively, a rosette could self-assemble via CesA interactions alone if a third type of CesA isoform were present which lacks the α2 binding site. This type of model explains the need for three different types of CesA isoforms within a cell-type, as evidenced by CesA mutant analyses. In primary walls, for example, AtCesA1 could encode the α2-type of CesA isoform, based on the rosette-disassociation phenotype observed in the rsw1 mutant (Arioli et al. 1998a). AtCesAs 3 and 6 could encode the α1- and β-type CesA isoforms, since they are expressed in the same cell type as AtCesA1. The different positions of the primary-wall CesA proteins within the rosette may also explain why the AtCesA3 and AtCesA6 isoforms are targets of isoxaben, another type of cellulose synthesis inhibitor, whilst AtCesA1 is insensitive despite high levels of sequence identity in the C-terminal region of CesA proteins where isoxaben is thought to bind (Scheible et al. 2001, Desprez et al. 2002). Furthermore, since the procuste mutant phenotype is only observed in the roots of dark-grown seedlings, it can be envisaged that another primary wall CesA isoform such as AtCesA2 or AtCesA5 may substitute for the AtCesA6 isoform in rosettes within aerial parts of the plant. However, this model of rosette structure does not adequately explain how rosette assembly would occur in cells in which there are apparently less than three CesA isoforms expressed, nor how homodimerization would be abrogated. Clearly, further characterization of rosette components and their interactions, as well as the unequivocal identification of which CesA proteins are assembled into a rosette in a particular cell-type is required before any conclusions can be drawn as to the merits of this model.
(c) Possible functional roles for CesA isoforms
One explanation of the requirement for multiple CesA isoforms for proper cellulose synthesis relates to the hypothesis that initiation of cellulose synthesis may require the formation of a primer, although, until recently, evidence for this has been limited. However, far more convincing proof of this hypothesis has recently been reported by Peng et al. (2002). When cotton-fiber membranes are incubated with 14C-labeled sitosterol-β-glucoside (SG) and unlabeled UDP-Glc, labeled sterolcellodextrins (SCDs) are produced, indicating that SG can serve as a primer for glucan chain elongation. Incubation of yeast membranes expressing GhCesA1 (it’s apparent ortholog in Arabidopsis is AtCesA8) with the same components resulted in synthesis of SG3, whereas yeast with vector only or expressing a mutated form of GhCesA1 in which the first conserved D residue is replaced by an A residue did not carry out this reaction. This demonstrates that the GT responsible for SCD synthesis in cotton is likely to be a CesA, and that the GhCesA1 isoform is capable of catalyzing this reaction. Additional evidence that the generation of SCDs is related to cellulose production comes from the observation that DCB inhibits SG synthesis in vivo: SG and SCD levels are reduced upon treatment of cultured cotton ovules with DCB, but addition of SG into the culture medium rescues the cellulose-deficient phenotype (Peng et al. 2002).
Based on these data, a model has been proposed in which SG serves as a primer for (1,4)-β-glucan elongation catalyzed by CesA proteins (Peng et al. 2002, Read and Bacic 2002). Additional evidence (outlined in the following section) suggests that CDs are cleaved from the SCD primer by a Ca2+-activated membrane-bound cellulase that could be analogous to the Korrigan (Kor) cellulase of Arabidopsis, known to be required for cellulose synthesis, and that the CDs are transferred to another CesA protein for further elongation (Nicol et al. 1998, Mølhøj et al. 2001, Williamson et al. 2001b, Peng et al. 2002, Read and Bacic 2002). Earlier experiments with Agrobacterium tumefaciens had suggested that a similar cellulose synthesis mechanism involving lipid-linked intermediates might exist in bacteria (Matthysse et al. 1995a, Matthysse et al. 1995b) where a membrane-anchored endocellulase encoded by one of the A. tumefaciens CelS operon genes (celC, AgtCesC) has significant sequence similarity to KOR, and is required for cellulose biosynthesis in this species. In this case, the cellulase was hypothesized to act as a transferase, catalyzing the transfer of cellobiose or larger units from the lipids to the growing cellulose chain in the final polymerization step. Although the model differs somewhat from that proposed by Peng et al. (2002) for plants, some rough similarities in the mechanism of primer synthesis for cellulose production may operate in plants and bacteria.
Such a synthesis mechanism in plants could easily explain the requirement for multiple CesA isoforms: some may be required for SCD primer formation, catalyzing SG elongation, and others could function in elongation of the CD primer after cleavage from SG (Perrin 2001, Scheible et al. 2001, Read and Bacic 2002). This model is attractive by virtue of its consistency with a number of other data. First, sterol attachment is at the reducing end of the Glc moiety, leaving the non-reducing end free for addition of Glc residues (Peng et al. 2002): the majority of data point towards elongation occurring at the non-reducing end of (1,4)-β-glucan chains (Koyama et al. 1997, Delmer 1999, Charnock et al. 2001). Secondly, SG synthesis is observed to occur on the inner face of plant plasma membranes (Cantatore et al. 2000) where cellulose synthesis is thought to take place and a SGT enzyme responsible for synthesis of sterol-β-glucosides has been found associated with the plasma membrane (Elbein and Forsee 1975). Thirdly, a difference in substrate binding and catalytic residues for the different CesA isoforms does not need to be invoked since each CesA polypeptide can utilize the same substrate, UDP-Glc, to catalyze the formation of a (1,4)-β-glycosidic linkage, the only difference being in the acceptor to which a Glc residue is attached. Fourthly, the model does not require that CesA polypeptides contain two catalytic centres. Models of Glc polymerization with multiple substrate binding sites have been proposed in order to account for the ~180° rotation of adjacent Glc residues in (1,4)-β-glucan chains (Albersheim et al. 1997, Carpita and Vergara 1998, Buckeridge et al. 1999), but the crystal structure of the at family 2 bacterial SpsA synthase clearly argues for a single glycosyl-transfer site (Charnock and Davies 1999, Charnock et al. 2001). Evidence of CesA homo- and heterodimerization (Taylor et al. 2000, Kurek et al. 2002) does, however, support the possibility that two CesA polypeptides may be involved in chain extension. Each CesA polypeptide may donate a catalytic site and in combination, add a cellobiose unit to the growing chain (Perrin 2001, Vergara and Carpita 2001). Furthermore, dimerization of CesA isoforms is thought to be more sterically favourable compared with a single CesA polypeptide for the formation of a pore (Vergara and Carpita 2001). The current model of CesA topology predicts that the eight TMHs interact to form a central channel through which the nascent glucan chain could be secreted, similar to the models predicted for H+ and Ca2+ ATPases (Delmer 1999). How a rosette produces 36 chains for elementary fibril formation by extrusion through a CesA dimer does present a difficulty, since such a mechanism is likely to produce only 18 chains. More recent estimates of microfibril size, however, suggest that 18 chains could produce a ~3.5 nm fibril, the size initially estimated for a 36-chain elementary microfibril (Ha et al. 1998). Unfortunately, there is no evidence as yet verifying that glucan chains pass through a pore formed by one or more CesAs, rather than a pore formed by another type of protein, as may be the case in A. xylinum (Saxena et al. 1994).
Given that six of possibly 12 ArabidopsisCesA genes have now been specifically linked with defects in cellulose synthesis, it seems likely that all CesA proteins are involved in cellulose production, rather than in synthesis of another polysaccharide. However, there is some circumstantial evidence that other CesA genes may be involved in either callose and specifically in the case of monocots, mixed-linkage glucan synthesis. Early work in the purification of the CelS enzyme indicated that a single synthase complex could be responsible for synthesizing both cellulose and callose, and that the wound-activated CalS enzyme could be a deregulated form of CelS (Delmer 1987). The CelS and callose synthase (CalS) enzymes both reside in the plasma membrane and utilize UDP-Glc as substrate (Delmer 1987). In addition, these enzymes appear to have opposing modes of regulation, with both glucans rarely being made by the same cell at the same time without apparent plasma membrane disruption (Delmer 1987). There is already a precedent that a bacterial member of GT family 2, CrdS, synthesizes (1,3)-β-glucan (Stasinopoulos et al. 1999). Direct evidence of a CesA gene product being deregulated to make a (1,3)-β-glucan comes from the observation that the D. discoideum CelS enzyme synthesizes (1,3)-β-glucan in vitro even though this polymer is not made in vivo, and that this activity is not detected upon disruption of its CesA gene (Blanton et al. 2000).
Massively parallel signature sequencing analysis of CesA genes in maize suggests that ZmCesA5 may encode the Golgi-localized mixed-linkage (1→3)(1→4)-β-D-glucan synthase that is found specifically in grasses (Dhugga 2001). ZmCesA5 is preferentially expressed in the endosperm and in expanding tissues such as coleoptiles, tissues rich in mixed-link glucan (Dhugga 2001). Its expression pattern is therefore consistent with a possible role in the synthesis of mixed-link glucan. Such a role for a CesA protein is conceivable, because this enzyme is likely to be a family 2 GT based upon the structure of its synthesis product and consequently, could be similar in sequence to CesA proteins. The mixed-link glucan enzyme also has a number of biochemical features that make it similar to CelS: its substrate is UDP-Glc, it is associated with sucrose synthase (see below), and membrane disruption causes a switch to CalS activity (Gibeaut and Carpita 1994, Becker et al. 1995, Vergara and Carpita 2001). Furthermore, phylogenetic analyses of cereal and dicot CesAs group ZmCesA5 into a separate subclass with other cereal CesAs that is most closely related to known primary-wall CesAs of maize and Arabidopsis (Vergara and Carpita 2001). Whilst again consistent with ZmCesA5 playing a role in mixed-link glucan synthesis, proof of its involvement remains to be demonstrated.
Other genes involved in cellulose biosynthesis
To date, no other genes encoding definitive CelS subunits have been identified, although a number of candidates have been proposed. Sucrose synthase (SuSy), an enzyme catalyzing the formation of UDP-Glc from sucrose, has been implicated to associate with the CelS complex or at least with the plasma membrane, possibly via interaction with actin, where it can channel substrate to CelS. The role of SuSy has been reviewed recently by Haigler et al. (2001) and will not be covered here except to add that recent protein–protein interaction screens with plant-specific regions of GhCesA1 have not yielded evidence for a direct interaction with SuSy, favouring the alternate concept that it might interact with actin at the plasma membrane. The cytoskeleton has long been viewed as playing a role in cellulose biosynthesis, with many studies indicating that the cortical microtubule (MT) network is involved in aligning the orientation of cellulose microfibrils as they are deposited into the wall (reviewed in Baskin 2001). However, studies in both cotton fibers and Zinnia tracheary elements using compounds that disrupt either MTs or actin microfilaments lead to a disorganized pattern of cellulose deposition, indicating that actin microfilaments are also involved in this process (Seagull 1990). Actin may therefore play a role in setting the pattern of cortical MTs, which in turn directs the pattern of cellulose microfibril deposition (Delmer and Amor 1995). The most recent alignment model, called “templated incorporation”, proposes that a scaffold of wall polysaccharides and plasma-membrane proteins in contact with MTs directs the orientation of the microfibril as it is deposited into the wall (Baskin 2001). A novel CelS purification method suggests, however, that the CelS enzyme is able to interact with tubulin directly. Eluate from an anti-tubulin column loaded with solubilized plasma membranes from azuki bean were found to contain 10 nm granules that produced (1,4)-β-glucan in the presence of UDP-Glc and Mg2+, with synthesis being inhibited by DCB (Mizuno 1996). Dissociation of these granular structures released eight polypeptides, including α- and β-tubulin (50 and 51 kDa; Mizuno 1996) and polypeptides at 120 kDa, 93 kDa and 65 kDa, which possibly represent CesA, SuSy and Kor subunits, respectively. In addition, an 18-kDa polypeptide was identified that may be the same as the uncharacterized 18-kDa protein in cotton fiber extracts that can be specifically labeled with a photoreactive analogue of DCB (Delmer et al. 1987). Since the inhibitory effect of DCB can be partly rescued by addition of SG (Peng et al. 2002), this DCB target protein may be involved in SG synthesis, but it’s precise role needs to be clarified. Further evidence for a direct interaction between tubulin and the CelS enzyme has been obtained from protein–protein interaction experiments (D. Jacob-Wilk and D. Delmer, unpublished data). The zinc-binding domain of GhCesA1 was used as bait against a 17 dpa cotton-fiber two-hybrid library in yeast. One of the clones found to activate reporter gene expression contained the last 32 amino acids of α-tubulin (GenBank accession no. AF106568). Expression of this gene has previously been found to increase during cotton fiber development (Whittaker and Triplett 1999). Together, these data indicate a direct interaction between the CelS complex and the MT network.
Whilst the cytoskeleton is unlikely to play a role in cellulose synthesis in prokaryotes, there may be other CelS enzyme components that are shared between bacteria and plants. This reasoning is based upon the commonality of the CesA and endocellulase proteins and the similarities in the primer synthesis mechanism observed between these systems. Early studies by several laboratories explored the possibility that a nucleotide-type activator similar to cyclic diguanylic acid (c-di-GMP) was required by the plant CelS enzyme. This molecule was initially identified as the allosteric activator of the A. xylinum CelS enzyme (Ross et al. 1991) but has also recently been implicated to be a CelS regulator in other cellulose-producing bacteria such as E. coli and Rhizobium leguminosarum, a bacterium with a very similar CelS operon compared with A. tumefaciens (Ausmees et al. 1999, Ausmees et al. 2001, Römling et al. 2000, Zogaj et al. 2001). AxCesB, the second gene of the CelS operon of A. xylinum, is thought to encode a 90 kDa regulatory CelS subunit that binds c-di-GMP (Saxena et al. 1994). A. tumefaciens does not encode a similar gene, but an antibody raised against the diguanyl cyclase of A. xylinum has been reported to cross-react with an A. tumefaciens polypeptide of similar molecular weight (Mayer et al. 1991). There have been some indications that c-di-GMP may also exist and function in plants. Using the same diguanyl cyclase antibody, Mayer et al. (1991) showed that a variety of plant species contained immunoreactive polypeptides. Further experiments showed that an 83-kDa polypeptide detected in cotton fiber membrane extracts can be labelled with good specificity and affinity by [32P]c-di-GMP, indicating that plants may contain a similar c-di-GMP-dependent CelS enzyme (Amor et al. 1991). This protein has not been able to be purified for further characterization in subsequent studies, however. A diguanylate cyclase gene has not been identified in analyses of the Arabidopsis genome (The Arabidopsis Genome Initiative 2000), hence the likelihood of c-di-GMP being the specific plant CelS activator now appears remote.
Instead, a different type of activated nucleotide may be a regulator of cellulose synthesis in plants. Expression of Rac13, a gene encoding a plant-specific GTPase, increases in the transition phase from primary to secondary wall synthesis in cotton fibers (Delmer et al. 1995). The timing of Rac13 expression coincides with the production of hydrogen peroxide (H2O2) which has been shown to stimulate the synthesis of cellulose during cotton fiber differentiation (Potikha et al. 1999). A dominant-active form of Rac13 constitutively activates H2O2 production in soybean and Arabidopsis cell cultures, implying that it may be involved in the differentiation of cotton fibers via H2O2 (Potikha et al. 1999). As mentioned previously, Kurek et al. (2002) have shown that CesA protein dimerization is regulated by redox state, with dimerization and higher order assembly favoured under oxidative conditions. Therefore, an oxidative environment favours rosette formation and cellulose synthesis. Redox regulation of CesA implicates other proteins which modulate redox state as being involved in cellulose biosynthesis, and a number of candidates have now been identified in our laboratory. A metallothionein (Mt) was found to interact with the zinc-binding domain of GhCesA1 in a yeast two-hybrid system, and its interaction further verified at the biochemical level (D. Wilk and D. Delmer, unpublished data). Mts are small cysteine-rich proteins that bind and exchange metal ions, particularly zinc. The Mt/t system has at least two functions, to sequester zinc, a consequence of gene regulation, and to release it by events that signal its requirement (Maret 2000). This regulatory process is a direct function of the cluster structure of Mt (Vasak and Kagi 1994). The cluster unit operates via a mechanism in which the cysteine sulfur bonds coordinating zinc ions are oxidized and reduced with concomitant release and binding of zinc, respectively (Maret and Vallee 1997). Thus, Mt is a redox protein and could be involved in the regulation of CelS complex formation or activation or deactivation by release or binding of zinc ions to the zinc-binding domain of GhCesA proteins, respectively (Fig. 2). A thioredoxin clone was also identified in the same two-hybrid screen, suggesting that Mt may not be the only protein involved in the redox regulation of CesAs (D. Wilk and D. Delmer, unpublished data).
As a follow-up to the decades-old suggestion that rosettes have short half-lives (Rudolph et al. 1989), we have recently shown that under reduced conditions (monomer state, Fig. 2), recombinant GhCesA1 zinc finger is specifically degraded by cysteine protease extracted from 24 dpa cotton fibers, suggesting that regulation of CesA redox state is also a means of controlling CesA turnover. There is also evidence to suggest that the rsw1-like rosette disassembly and non-crystalline glucan accumulation effects observed upon treatment of cotton ovules and Arabidopsis seedlings with the cellulose synthesis inhibitor CGA 325′615 are due to the inability of CesA proteins to be oxidized (H2O2 completely reverses the effects of CGA), indicating that this inhibitor may target a protein that is involved in CesA oxidation (Peng et al. 2001, Kurek et al. 2002).
Also found to interact with the GhCesA1 zinc-binding domain was a putative non-specific lipid transfer protein (LTP, GenBank accession no. AF228333). This gene was also previously found to be differentially expressed in developing cotton fibers (Ma et al. 1995) and promotor–GUS analyses in tobacco plants indicate that this gene is specifically expressed in trichomes (Liu et al. 2000). Due to its sequence homology to LTPs, this gene was suggested to be involved in cuticle deposition. However, to our knowledge, there is no supporting data showing the involvement of this fiber-specific putative LTP in the binding or synthesis of cuticular materials. Our laboratory is currently investigating the function of this protein and its possible involvement in cellulose biosynthesis.
A vast array of other genes are likely to be directly and indirectly involved in the cellulose synthesis process. One example of an indirectly associated gene is CYT1 encoding a mannose-1-phosphate guanylyltransferase. An Arabidopsiscyt1 mutant shows a five-fold reduction in cellulose content (Lukowitz et al. 2001). The effect on cellulose synthesis is likely to be due to pleiotropic secondary effects, since the encoded enzyme is required for the production of GDP-mannose, GDP-fucose, ascorbic acid, glycosylphosphatidylinositol membrane anchors, and the core N-glycan. There is some evidence to suggest that CesA proteins are N-glycosylated (Delmer 1999) and we also note that Korrigan has been shown to require such glycosylation for activity, indicating a potentially more specific requirement for CYT1 in cellulose synthesis. Characterization of the genetic lesions responsible for other cellulose-deficient mutants such as tbr (trichome birefringence), impaired in its ability to synthesize secondary wall cellulose in trichomes and other cell types (Potikha and Delmer 1995), and brittle culm lines of barley, also specifically affected in their ability to produce secondary wall cellulose (Kimura et al. 1999b), may shed more light on which other proteins are required for cellulose synthesis in plants.
Open questions for the future
The SG primer model of cellulose synthesis is still highly speculative and far from proven. A number of important questions with respect to the cellulose synthesis mechanism need to be answered. First, are SGs the primer used in plants in vivo and, if so, how widespread are they in cellulose synthesis within eukaryotes? It is possible that we have discovered a primer that substitutes very well in our in vitro system for some other natural primer such as a protein or lipid that normally works in the in vivo situation. If SGs are the in vivo primer in plants, other cellulose-synthesizing organisms may still use a different type of glycoside as primer, as would necessarily be the case in bacteria since prokaryotes lack sterols. Secondly, do CesA proteins catalyze the elongation of CDs as well as the elongation of the SG primer in vivo? Our work to date demonstrates that a single CesA in yeast can begin SG elongation but not complete it. Thus, only the latter activity has been demonstrated, and only for one CesA protein. It remains to be determined whether CesAs catalyze an elongation reaction from CDs and whether or not they are capable of utilizing multiple types of acceptors. Also, it is not clear at this stage if the same CesA, either in vitro or in its native environment, with other required proteins such as Kor, might able to complete the elongation or whether it might require cooperative interaction with other CesA isoforms. Thirdly, is the true function of the Kor-type protein to cleave SCD and, if so, how does this cleavage take place? The GT that attaches a Glc moiety to sterols is located in the cytoplasm, yet the catalytic domain of the Kor-type protein implicated in SCD cleavage is predicted to be located on the other side of the plasma membrane. Thus, the SCD must flip in the membrane for the Glc chain to present itself for cleavage in the wall space. Verification of a Type II membrane topology for the Kor protein would be beneficial in answering this question. A fourth and related question is how are CDs elongated by CesA protein/s? The catalytic site of a CesA protein is likely to be located in the cytoplasm where its substrate UDP-Glc is located. Thus, the CD primer must re-traverse the plasma membrane, this time without an attached hydrophobic moiety, to be accessible to the CesA catalytic site for subsequent elongation. Based on the size of cellobiose (1.03 nm), an oligosaccharide of at least ten Glc residues is probably required to span the 5–7 nm width of the plasma membrane (Delmer 1999). Is it possible that the cleaved hydrophilic CD spans the membrane and presents its nonreducing end to the cytoplasmic face of the plasma membrane? One way in which this could occur is if the CD is delivered to the proposed pore-type structure used for glucan chain transfer through the plasma membrane. Such an elaborate biosynthesis mechanism need not be evoked, however, if it were shown that Kor has a type I rather than a type II membrane topology.
Despite the speculative nature of the model, it has generated a number of predictions that can be tested, the results of which should prove useful in resolving what parts of the proposed cellulose synthesis mechanism are correct and which ones are not. For example, if Kor plays the role ascribed to it above, a kor mutant should accumulate SCDs. Sato et al. (2001) mention in their discussion that the ArabidopsisKor mutant acs1 does in fact accumulate lipid-linked (1,4)-β-glucans whose chain length varies from one to six Glc residues. That data generated in both cotton and Arabidopsis conform to the proposed model strengthens the case that this mechanism of cellulose synthesis is universal amongst plants, at least in vitro. Other genes are also predicted as being involved in the cellulose synthesis mechanism by the SG primer model, such as the UDP-Glc:sterol glucosyltransferases that synthesize SG. There are two and three genes within Arabidopsis and cotton, respectively, that are highly related to the purified and cloned oat enzyme (Warnecke et al. 1997), and their possible role is currently being investigated (M. Doblin, W. Scheible, J. Schultz, C. Somerville and D. Delmer, unpublished data).
Status of in vitro cellulose synthesis in plants
One of the achievements in the work by Peng et al. (2002) was the establishment of conditions that led to significant synthesis of SG, SCDs and non-crystalline cellulose in vitro. Thus, there is reason to believe that this work can now be continued and optimized further in a variety of plant systems. This would be an important breakthrough, as it could lead ultimately to the ability to purify active rosettes and to identify all the components of the CelS complex. One of the keys to improvement in cellulose synthesis in vitro was the substitution of MOPS buffer for Tris, something that had also been noted previously. In a series of papers, Malcolm Brown and his colleagues reported on their attempts to improve the in vitro assay for plant cellulose biosynthesis (Okuda et al. 1993, Kudlicka et al. 1995, Kudlicka and Brown 1997). What has become clear from their work is that the conditions of the assay as well as membrane extraction and enzyme solubilization are crucial in obtaining any cellulose synthesis in vitro (defined as the ability to produce acetic-nitric acid-resistant (1,4)-β-glucan). For example, synthesis is significantly improved if in the extraction buffer MOPS is used rather than Tris and when a cocktail of protease inhibitors showing a broad range of specificity is added to slow the degradation of the labile CelS complex. Solubilization with low concentrations of digitonin (0.1–0.5%) helped in keeping rosettes intact and addition of effectors such as Mg2+, Ca2+ and cellobiose in the assay were also found to be requirements. Under these conditions, the quantity of in vitro cellulose synthesized from cotton fiber membranes was found to increase from 4% to 32% of total glucan product (Kudlicka et al. 1995). The improvements made by Brown were extremely useful in our studies. We have shown that the use of MOPS buffer, or at least not Tris buffer, is necessary for cellulose synthesis in vitro because it is required for the synthesis of SCDs that start glucan chain elongation (Peng et al. 2002). There is now potential for even greater levels of cellulose synthesis in vitro because some of the molecular components of rosettes have been identified. In vitro assays involving the addition of SG, expression of a second or possibly third cotton CesA gene with and without KOR, etc. should prove useful in establishing what the roles of different CelS components are.
One last point worth mentioning is that even if the SG primer model is shown to be incorrect in vivo, the fact that SG is a good primer in vitro could be extremely beneficial in future CelS purification efforts where loss of the native primer may occur early in the purification procedure. Furthermore, using DCB-habituated cells may also be advantageous in such efforts as CesA protein levels are elevated in these cells compared with untreated or unhabituated cells (Nakagawa and Sakurai 1998).
Conclusions
To date, the CelS components identified to be common to all cellulose-synthesizing organisms are the CesA subunit and a cellulase. The encoding genes, as well as their mutants, provide us with some powerful tools to probe the process of cellulose synthesis. Furthermore, the identification of these obligatory components provides hope that additional improvements can be made to the in vitro cellulose synthesis assay in plant systems so that in the future, a functional CelS complex will be able to be reconstituted. These improvements will be crucial in our ability to further dissect the mechanism of cellulose synthesis in plants. Such advances are likely to be the key to paving the way towards future manipulations of cellulose in plant cell walls for human benefit.
Corresponding author: E-mail, msdoblin@unimelb.edu.au; Fax, +61 3 9347 1071.
Current address: University of Melbourne, School of Botany, Royal Parade, Parkville 3010, Victoria, Australia.
Fig. 1 A model for the structure of the rosette. (A) Six subunits, possibly containing six CesA polypeptides, interact to form a rosette, a single CesA enzyme complex. Each CesA polypeptide is shown to be involved in the synthesis of one β-(1,4)-glucan chain. The CesA protein has eight predicted TMHs which could potentially form a pore in the plasma membrane through which the nascent chain is extruded into the wall. Once the 36 chains emerge from the rosette, they coalesce to form an elementary cellulose microfibril. (B) In this modified rosette structure model of Scheible et al. (2001), at least two types of CesA polypeptides, α and β, are required for spontaneous rosette assembly. Two different types of α isoform can be distinguished, α1 which interacts with two β isoforms only, and α2 interacting with another α2 isoform and two β isoforms.
Fig. 2 Redox regulation of CesA. The zinc-finger domain at the N-terminus of CesA proteins can exist in either a reduced or oxidized form. Under reduced conditions, this domain coordinates two zinc ions and can interact with either lipid transfer protein (LTP), metallothionein (Mt), microtubules (MT) or cysteine protease. Under oxidized conditions, the CesA protein can dimerize with itself or another CesA protein. Proteins such as the Korrigan cellulase (Kor), the plasma membrane-associated form of sucrose synthase (pm-SuSy), actin and the herbicide CGA binding protein (CGAbp) have all been implicated as playing a role in cellulose synthesis.
Abbreviations
- CalS
callose synthase enzyme complex
- CD
cellodextrin
- c-di-GMP
cyclic diguanylic acid
- Csl
cellulose synthase-like
- CelS
cellulose synthase enzyme complex
- CesA (formerly CelA)
cellulose synthase catalytic subunit
- CGAbp
cellulose synthesis inhibitor CGA 325′615 binding protein
- CR-P
plant conserved region
- dpa
days post anthesis
- DCB
2,6-dichlorobenzonitrile
- Glc
glucose
- GT
glycosyltransferase
- HVR
hypervariable region
- Kor
Korrigan endocellulase
- LTP
lipid transfer protein
- Mt
metallothionein
- MT
microtubule
- SCD
sterol cellodextrin
- SG
sitosterol-β-glucoside
- SuSy
sucrose synthase
- TC
terminal complex
- TMH
transmembrane helix, UDP-Glc, uridine diphospho-glucose.
References
Albersheim, P., Darvill, A., Roberts, K., Staehelin, L.A. and Varner, J.E. (
Arioli, T., Peng, L., Betzner, A.S., Burn, J., Wittke, W., Herth, W., Camilleri, C., Höfte, H., Plazinski, J., Birch, R., Cork, A., Glover, J., Redmond, J. and Williamson, R.E. (
Ausmees, N., Jonsson, H., Höglund, S., Ljunggren, H. and Lindberg, M. (
Ausmees, N., Mayer, R., Weinhouse, H., Volman, G., Amikan, D., Benziman, M. and Lindberg, M. (
Blanton, R.L., Fuller, D., Iranfar, N., Grimson, M.J. and Loomis, W.F. (
Burn, J.E., Hocart, C.H., Birch, R.J., Cork, A.C. and Williamson, R.E. (
Desprez, T., Vernhettes, S., Fagard, M., Refrégier, G., Desnos, T., Aletti, E., Py, N., Pelletier, S. and Höfte, H. (
Doblin, M.S., De Melis, L., Newbigin, E., Bacic, A. and Read, S.M. (
Fagard, M., Desnos, T., Desprez, T., Goubet, F., Refregier, G., Mouille, G., McCann, M., Rayon, C., Vernhettes, S. and Höfte, H. (
Ha, M.-A., Apperley, D.C., Evans, B.W., Huxham, I.M., Jardine, W.G., Viëtor, R.J., Reis, D., Vian, B. and Jarvis, M.C. (
Haigler, C.H., Ivanova-Datcheva, M., Hogan, P.S., Salnikov, VV., Hwang, S., Martin, K. and Delmer, D.P. (
Holland, N., Holland, D., Helentjaris, T., Dhugga, K.S., Xoconostle-Cazares, B. and Delmer, D.P. (
Kawagoe, Y. and Delmer, D.P. (
Kimura, S., Laosinchai, W., Itoh, T., Cui, X., Linder, C.R. and Brown, R.M. Jr. (
Koyama, M., Helbert, W., Imai, R., Sugiyama, J. and Henrissat, B. (
Kudlicka, K., Brown, R.M., Li, L., Lee, J.H., Shin, H. and Shigenori, K. (
Kurek, I., Kawagoe, Y., Jacob-Wilk, D., Doblin, M. and Delmer, D.P. (
Lukowitz, W., Nickle, T.C., Meinke, D.W., Last, R.L., Conklin, P.L. and Somerville, C.R. (
Mayer, R., Ross, P., Weinhouse, H., Amikam, D., Volman, G., Ohana, P., Calhoon, R.D., Wong, H.C., Emerick, A.W. and Benziman, M (
Nagahashi, S., Sudoh, M., Ono, N., Sawada, R., Yamaguchi, E., Uchida, Y., Mio, T., Takagi, M., Atisawa, M. and Yamada-Okabe, H. (
Nakai, T., Tonouchi, N., Konishi, T., Kojima, Y., Tsuchida, T., Yoshinaga, F., Sakai, F. and Hayashi, T. (
Nicol, F., His, I., Jauneau, A., Vernhettes, S., Canut, H. and Höfte, H. (
Pear, J.R., Kawagoe, Y., Schreckengost, W.E., Delmer, D.P. and Stalker, D.M. (
Peng, L., Xiang, F., Roberts, E., Kawagoe, Y., Greve, L.C., Kreuz, K. and Delmer, D.P. (
Potikha, T.S., Collins, C.C., Johnson, D.I., Delmer, D.P. and Levine, A. (
Roberts, A., Roberts, E. and Delmer, D.P. (
Sato, S., Kato, T., Kakegawa, K., Ishii, T., Liu, Y.-G., Awano, T., Takabe, K., Nishiyama, Y., Kuga, S., Sato, S., Nakamura, Y., Tabata, S. and Shibata, D. (
Saxena, I.M., Brown, R.M. Jr., Fevre, M., Geremia, R.A. and Henrissat, B. (
Scheible, W.-R., Eshed, R., Richmond, T., Delmer, D. and Somerville, C. (
Taylor, N.G., Scheible, W.-R., Cutler, S., Somerville, C.R. and Turner, S.R. (
Taylor, N.G. and Turner, S.R. (
Vasak, M. and Kagi, J.H.R. (
Warnecke, D.C., Baltrusch, M., Buck, F., Wolter, F.P. and Heinz, E. (
Williamson, R.E., Burn, J.E., Birch, R., Baskin, T.I., Arioli, T., Betzner, A.S. and Cork, A. (