-
PDF
- Split View
-
Views
-
Cite
Cite
Hye-Kyoung Kwon, Ryusuke Yokoyama, Kazuhiko Nishitani, A Proteomic Approach to Apoplastic Proteins Involved in Cell Wall Regeneration in Protoplasts of Arabidopsis Suspension-cultured Cells, Plant and Cell Physiology, Volume 46, Issue 6, June 2005, Pages 843–857, https://doi.org/10.1093/pcp/pci089
Close - Share Icon Share
Abstract
To clarify the mechanisms of cell wall construction, we used a proteomic approach to investigate the proteins secreted into cell wall spaces during cell wall regeneration from the protoplasts of Arabidopsis suspension-cultured cells. We focused on cell wall proteins loosely bound to the cell wall architecture and extractable with 1 M KCl solutions from: (i) native suspension cultured cells; (ii) protoplasts that had been allowed to regenerate their cell walls for 1 h; and (iii) protoplasts allowed to regenerate their cell walls for 3 h. We adopted a non-destructive extraction procedure without disrupting cellular integrity, thereby avoiding contamination from cytoplasmic proteins. Using two-dimensional polyacrylamide gel electrophoresis (2-D PAGE) and matrix-assisted laser desorption ionization-time-of-flight/mass spectrometry (MALDI-TOF/MS), we separated, mapped and identified 71 proteins derived from the native cell wall, and 175 and 212 proteins derived from the 1 and 3 h regenerated protoplasts, respectively. Quite different sets of proteins with differing status of their post-translational modifications, including phosphorylation and glycosylation, were identified in the three protein fractions. This indicated dynamic in muro changes in the cell wall proteins during cell wall regeneration in the protoplasts. The analysis revealed a set of enzymes specifically involved in cell wall expansion and construction in suspension-cultured cells. This approach has also determined a set of cell wall proteins that had not been predicted to be localized in cell wall spaces.
Introduction
The plant cell wall is a dynamic structure that plays a critical role not only in determining cell shape and formation of the plant body, but also in interactions with environmental factors including those required for nutrition, response to abiotic stress and biological attack by other organisms (Carpita and McCann 2000). It is made of various types of macromolecules, and undergoes extensive construction and remodeling during cell division and differentiation. Cell wall dynamics are achieved via two distinct steps: (i) the biosynthesis of cell wall components by the actions of membrane-bound enzymes at the plasma membrane and endoplasmic reticulum (ER)–Golgi apparatus; and (ii) the assembly and rearrangement of cell wall structures in muro via the actions of extracellular proteins. Significant progress has been made recently in the identification of membrane-bound enzymes involved in the syntheses of cellulose and matrix polymers (Arioli et al. 1998, Perrin et al. 1999, Hong et al. 2001, Faik et al. 2002, Iwai et al. 2002, Li et al. 2003), as well as with certain cell wall proteins such as expansin and the XTH family of genes (Cosgrove 1997, Nishitani 1997). However, only a limited number of extracellular proteins compared with those predicted based on the genomic database have thus far been characterized, particularly in terms of cell wall dynamics (Girke et al. 2004, Yokoyama and Nishitani 2004).
The functional and structural diversity of the plant cell wall largely mirrors the abundance and diversity of extracellular proteins, which make up approximately 10% of the cell wall mass, and are typically encoded by large gene families (Chivasa et al. 2002, Yokoyama and Nishitani 2004). More than 2,000 putative cell wall-related genes are estimated to occur in the genome database of Arabidopsis thaliana (Arabidopsis Genome Initiative 2000). These genes are considered to encode proteins in categories with differing biological functions (Cassab 1998, Darley et al. 2001, Nishitani 2002, Qin et al. 2003). Reverse genetic approaches based on oligo DNA microarray technology coupled with phenotypic analyses of T-DNA insertion lines have paved the way for understanding the functions of this large number of gene sets involved with the cell wall (Bonetta et al. 2002, Li et al. 2004, Manfield et al. 2004). Despite these genomic approaches, a complete picture of the concerted action of these cell wall proteins in the cell wall construction and remodeling processes is largely unknown.
A proteomic approach has been adopted to gain insight into the function and localization of individual cell wall proteins in muro. This promising technology allows the large-scale analysis of protein expression profiles, including their post-translational modifications, and protein–protein interactions (Pandey and Mann 2000). This is particularly the case for proteins found in specific organs and cellular compartments, or for those expressed under specific biological conditions (Kuster et al. 2001, Martin and Nelson 2001). Using this proteomic approach, lists of proteins expressed in the cell wall of A. thaliana have been determined recently (Rose et al. 2004a, Rose et al. 2004b).
However, even with current proteomic approaches, dynamic aspects of cell wall proteins, particularly those during cell wall construction, remain elusive. To better understand cell wall proteins, we considered the process of cell wall regeneration in the present study. We used protoplasts generated from suspension-cultured Alex cells of Arabidopsis, which is not only a model plant with a complete genome database and abundant transcriptomes, but also provides us with intact cells ready to extract cell wall proteins without damaging cell integrity. Using this ‘intact model cell’ system, we extracted extracellular proteins that were secreted from the protoplasts and bound loosely to its surface during the cell wall regeneration process with 1 M KCl solutions. We then analyzed them comprehensively with the aid of two-dimensional polyacrylamide gel electrophoresis (2-D PAGE) and matrix-assisted laser desorption ionization-time-of-flight/mass spectrometry (MALDI-TOF/MS) technologies. This approach has revealed dynamics of protein profiles that are distinct from those found by conventional cell wall proteome analyses. In addition, this approach also disclosed the presence of a certain set of proteins that had not been identified as being secreted.
Results
Purification of cell wall proteins secreted from suspension-cultured cells of Arabidopsis during cell wall regeneration on protoplasts
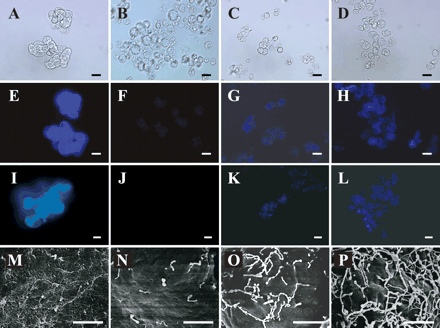
To gain insight into dynamic aspects of cell wall construction, we focused on a set of proteins secreted onto the surface of protoplasts during cell wall regeneration. Fig. 1 shows differential interference contrast images (Fig. 1A–D) and epifluorescent images (Fig. 1E–L) of native cells and protoplasts stained with Calcofluor White M2R (Fig. 1E–H) and aniline blue (Fig. 1I–L), which preferentially stain cellulose and β-1,3 glucan, respectively. Fresh protoplasts were not stained with either Calcofluor White M2R (Fig. 1F) or aniline blue (Fig. 1J). The deposition of glucans, as determined by staining with the two fluorescent compounds, appeared within 1 h of cell wall regeneration, and the extent of deposition increased and spread over the surface during 3 h of regeneration (Fig. 1G, H, K, L).
A close-up examination of the cell surface using a scanning electron microscope (SEM, Fig. 1M–P) showed that the native cell surface was covered with a tangled network of fine fibrils (Fig. 1M), whereas the surface of freshly prepared protoplasts was smooth, and some dumbbell-shaped structures were sparsely deposited onto the surface (Fig. 1N). As cell wall regeneration proceeded, the dumbbell-shaped threads elongated. On the surface of the protoplasts that had been allowed to regenerate cell walls for 3 h, a network of fine threads similar to those found on the native cell surface was observed (Fig. 1M, P). These results indicate that the secretion of cell wall material begins within 1 h of the incubation of protoplasts, and that the formation of β-glucan-containing cell wall structures similar to those found in native cells begins within 3 h of cell wall regeneration. Therefore, we decided to examine proteins on the surface of protoplasts that had been incubated for 1 and 3 h. We also extracted loosely bound cell wall proteins using a 1 M KCl solution according to the procedure examined previously (Robertson et al. 1997, Borderies et al. 2003).
To validate whether cytosolic proteins leak out of the protoplasts during protein extraction processes, we tested for the activity of glucose-6-phosphate dehydrogenase (G6PDH), a typical marker enzyme used to detect contamination by cytoplasmic proteins (Li et al. 1989). Table 1 shows that specific activities of G6PDH in the cell wall protein fraction from the native cells contained 1.6% of the activity of those from the cytosolic extracts. Furthermore, a much lower amount of enzyme activity was detected in the cell wall protein fraction derived from protoplasts compared with those in native cells. From these results, we consider that contamination of intracellular proteins is minimized in the present study.
We also examined the viability of cells and protoplasts that had been extracted with 1 M KCl by staining with fluorescein diacetate (FDA). We counted the viable cells under a fluorescence microscope. This showed that >90% of protoplasts remained viable after the 1 M KCl extraction (data not shown). Judging from the results obtained from the two different tests, we conclude that the cell membrane is not severely damaged during protein extraction, and that contamination of the extracellular protein fraction by cytosolic proteins is negligible.
Characterization of cell wall proteomes
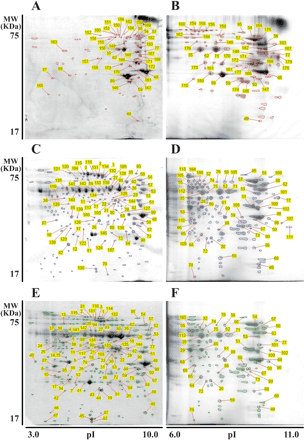
Cell wall proteins were separated by 2-D PAGE using differently immobilized isoelectric point (pI) gradient (IPG) strips for the first-dimension separation (pI 3–10 and pI 6–11) and 12% SDS–PAGE for the second separation. These were visualized with Coomassie brilliant blue (CBB) staining (Fig. 2). Using Image Master 2D-Elite software, we mapped 71 proteins in the 2-D gel derived from the native cultured cells, and 175 and 212 protein spots in those derived from the 1 and 3 h regenerated protoplasts, respectively (Fig. 2, 3). By matching individual spots in different gels derived from the three protein fractions, we identified 327 protein spots (Tables 2–4). This spot matching revealed that 26 of the 71 spots found in native cells are not in the protoplasts, while 75 of the 175 spots, and 104 of the 212 spots are specific to the 1 and 3 h regenerated protoplasts, respectively, as summarized in Fig 3. Consequently, 256 of the 327 spots detected in the present study were specifically expressed during cell wall regeneration on the cell wall surface.
After individual protein spots were picked and in-gel digested with trypsin, protein sequences and genes encoding the proteins were identified with a MALDI-TOF/MS analysis coupled with a database search using the Mascot sequence database search program. The protein spots with their genes predicted were numbered on each of the gels as shown in Fig. 2 and listed in Tables 2–4, respectively. Protein spots with the same numbers are identical. The data in Tables 2–4 indicate that two or more spots are predicted to be encoded by the same gene, indicating the occurrence of post-translational modifications. Consequently, 55 protein spots derived from the native cells were identified as being encoded by a total of 39 independent genes (Table 2). Similarly, 108 protein spots derived from the 1 h regenerated protoplasts were identified as being encoded by 61 independent genes (Table 3), and 116 protein spots from the 3 h regenerated protoplasts were identified as being encoded by 67 independent genes (Table 4). Other spots that were recognized on the gel by staining, but were not identified in terms of genes, were not numbered in Fig. 2.
This analysis has identified several carbohydrate-related enzymes as being localized in the cell wall of intact cells and/or in the protoplasts during cell wall regeneration. These enzymes include expansin, chitinase, xyloglucan endotransglycosylase/hydrolase, β-glucosidase, α-glucosidase, α-xylosidase, β-xylosidase, α-galactosidase, β-galactosidase, β-fructosidase, α-mannosidase, polygalacturonases, fructokinase and β-hexosaminidases (Tables 2–4). Given that protein families have dozens of family members (Yokoyama and Nishitani 2004), it is worth noting that only a single, or at most two, members of each of these protein families is identified in each protein fraction. This implies specific roles for individual proteins in specific aspects of cell wall dynamics in suspension-cultured cells. In addition to these carbohydrate-related enzymes, the present study identified several types of proteases, esterases, kinases and oxidoreductases as cell wall resident proteins.
Furthermore, we have identified a few proteins whose molecular functions have not yet been predicted. For these unknown proteins, we predicted their functions by examining their structural features using domain and motif analyses. These analyses identified a group of proteins that are classified into lectin families with potential domains for interaction with certain polysaccharides or proteins present in the cell wall (Tables 2–4).
Post-translational modifications
As denoted in Tables 2–4, identification using the MALDI-TOF/MS analysis showed that some spots are attributed to a single protein encoded by a single gene, suggesting post-translational modifications. A comparison of the relative positions in the 2-D gel among the protein spots derived from the same gene indicates that modifications would have affected the pI and/or the molecular weight. To characterize these modifications, we assayed for phosphorylated and glycosylated proteins in the 2-D polyacrylamide gel by staining with Pro-Q Diamond dye (Molecular Probes, Eugene, OR, USA) and Pro-Q Emerald dye (Molecular Probe), respectively. Data indicated that of 39 independent proteins found in native cells, 18 proteins (denoted by asterisks in Table 2) were phosphorylated, and 28 proteins (denoted by underlines in Table 2) were glycosylated. Similarly, 22 and 43 proteins of 61 proteins derived from the 1 h regenerated protoplasts were phosphorylated and glycosylated, respectively (Tables 2, 4), while 31 and 45 proteins derived from the 3 h regenerated protoplasts were phosphorylated and glycosylated, respectively. For example, spots 150, 151 and 152 in the native cells (Fig. 2A) are the same α-xylosidase (At1g68560). Whereas they have the same molecular weight, they have a different pI. Spots 95, 96 and 118 from 1 h regenerated extracts (Fig. 2C, D) with different molecular weights and pI represent the same β-galactosidase (At5g63810). α-Mannosidase (At3g26720) was also identified as different spots with different molecular weights and pI in individual steps of cell wall regeneration: native (spots 159 and 160), 1 h (spot 119) and 3 h (spot 33). These results suggest that cell wall proteins undergo various types of post-translational modifications during cell wall regeneration.
Discussion
Proteins potentially involved in cell wall expansion
In this work, we carefully extracted cell wall proteins that were ionically bound to the cell surface using a non-destructive procedure without disrupting cellular integrity, thereby avoiding contamination from cytoplasmic proteins, and analyzed the protein profiles comprehensively. This non-destructive cell wall proteome approach disclosed a set of proteins that are expressed specifically during cell wall regeneration, and are loosely attached to the cell surface during the cell wall regeneration process in Arabidopsis suspension-cultured cells. In the cell wall protein fraction of the native cells, we identified several key proteins implicated in the remodeling of the cellulose/xyloglucan network, in which xyloglucan functions as major load-bearing cross-links between cellulose microfibrils. These proteins include expansin, XTH, α-xylosidase and chitinase.
Expansins are a class of proteins capable of causing cell wall expansion under acidic conditions (Cosgrove 2000). Whereas the mechanism of expansin action still remains elusive, it is implicated in a reversible disruption of hydrogen bonding between cellulose microfibrils and xyloglucan. This results in a loosening of the cell wall and allows turgor-driven deformation of the cellulose/xyloglucan framework (McQueen-Mason and Cosgrove 1994, Cosgrove 2000, Whitney et al. 2000). These proteins are encoded by a large multigene family (McQueen-Mason and Cosgrove 1994), which can be divided into four subfamilies: two expansin subfamilies, EXPA and EXPB; and two expansin-like subfamilies, EXLA and EXLB (Kende et al. 2004). The EXLA1 (At3g45970) identified in this study is a member of the EXLA subfamily. Unlike EXPA and EXPB (Cosgrove 1997), no biological or biochemical function has yet been established for any member of the EXLA or EXLB family (Kende et al. 2004), despite the fact that members of EXLA and EXLB possess both expansin domain I and II. Thus, the present result might imply a possible role for EXLA1 protein in the cell wall construction, particularly in suspension-cultured cells.
The chitinase family in Arabidopsis consists of two distinct subfamilies. One, acidic endochitinase, encodes proteins highly homologous to ‘yieldin’, which is implicated in the modulation of mechanical properties of the cell wall in cowpea plants (Okamoto-Nakazato et al. 2000, Yokoyama and Nishitani 2004). A phylogenetic analysis reveals that the chitinase isoform (At5g24090), which we have identified in the present study, is the one and only yieldin-like protein in Arabidopsis (Table 2; Yokoyama and Nishitani 2004). It is likely that this Arabidopsis yieldin ortholog is involved in the regulation of mechanical properties in the plant cell wall, thereby modulating cell expansion in the suspension-cultured cells.
Two proteins encoded by AtXTH11 (At3g48580) and AtXYL1 (At1g68560), which were found to be present in the apoplastic fraction in both the native cells and protoplasts (Tables 2–4), are implicated in the modification of these xyloglucan molecules. AtXTH11 is a member of the endotransglucosylase/hydrolase (XTH) family of proteins, which can mediate splitting and/or reconnection of the xyloglucan cross-links in the cell wall (Nishitani and Tominaga 1991, Nishitani and Tominaga 1992, Okazawa et al. 1993, Campbell and Braam 1999, Yokoyama and Nishitani 2001, Rose et al. 2002). As such, it is considered essential for cell wall dynamics. In Arabidopsis, 33 genes encoding this family of proteins have been identified in the genome. The present study disclosed that only one of the 33 XTH members is specifically expressed both in the native cell wall and during the cell wall regeneration processes in the protoplasts. This AtXTH11 protein belongs to the group I/II subfamily of the XTH family of proteins, and is predicted to exhibit xyloglucan endotransglucosylase activity exclusively, suggesting its role in the integration of newly secreted cellulose/xyloglucan complexes into the cell wall components during cell wall construction (Ito and Nishitani 1999, Yokoyama and Nishitani 2001).
The AtXYL1 protein, which shows a signature region of family 31 glycosyl hydrolases, is implicated in the degradation of an unsubstituted α-xylosyl residue situated nearest to the non-reducing end of xyloglucan oligosaccharides. Higher levels of the AtXYL1 transcript as well as α-xylosidase activity were observed in younger rosette leaves of Arabidopsis, indicating developmental regulation of their expression (Sampedro et al. 2001). It should be noted that several protein spots for the AtXYL1 protein with differing molecular sizes and pI have been identified. This indicates that this protein undergoes extensive post-translational modifications. It is also interesting that some spots are phosphorylated, while others are not. This implies that kinase-mediated regulation in muro may be involved in the regulation of this enzyme. Thus, the data imply that AtXTH11 and AtXYL1 play roles in the modification of the cellulose xyloglucan network during cell wall regeneration, as well as in the native cell walls. In the present study, we did not examine incorporation of phosphate into individual proteins or kinase activities responsible for the putative phosphorylation reactions. Further investigation is required to demonstrate the in muro phosphorylation of these proteins.
A polygalacturonase (At5g41870), which was also found in the native cells, hydrolyses the α-1,4-d-galacturonan backbone of pectic polysaccharides (Brummell and Harpster 2001). Enzyme activity of the polygalacturonase reportedly increases throughout the expansion period of ripening fruit in many plant species (Hobson 1962, Brummell and Labavitch 1997). A polygalacturonase present in liverwort cell cultures increased in parallel with cell growth until the cessation of the exponential growth phase, at which time the activity declined (Konno et al. 1983). In addition, polygalacturonases extracted from cultured carrot cells degraded pectic polymers from these cultures (Konno et al. 1984). Based on these developmental correlations, it is possible that polygalacturonases act coordinately with other cell wall-modifying enzymes to increase cell wall extensibility (Konno et al. 1983). We also identified other proteins involved in the exo- or endo-type hydrolysis of polysaccharides in the native cells (Table 2). These enzymes may also act coordinately with cell wall-loosening proteins to increase cell wall extensibility, thus permitting cell expansion.
Proteins possibly involved in cell wall regeneration
Whereas expansin, yieldin and polygalacturonase were not detected in the cell wall fraction derived from the protoplasts, several protein spots encoded by AtXTH11 and AtXYL1 were found to be expressed in the protoplasts as well as in native cells. However, given the incomplete construction of the cellulose/xyloglucan network during the first hours of cell wall regeneration on the protoplasts (Fig. 1), it is unlikely that AtXTH11 participates in the remodeling of the pre-existing cellulose/xyloglucan framework as in native cell walls. It is more probable that AtXTH11 plays a role in the integration of newly secreted xyloglucan material into the pre-secreted xyloglucan molecules, thereby assembling the precursor for cellulose/xyloglucan construction (Ito and Nishitani 1999, Nishitani 1997). Accordingly, AtXTH11 may fill dual roles, wall assembly and restructuring, during the cell wall regeneration and expansion processes (Nishitani 1997).
XTH can mediate the depolymerization of xyloglucan molecules by transferring a xyloglucan fragment generated by splitting a xyloglucan donor substrate to a small xyloglucan acceptor substrate. This depolymerization activity of XTH directly depends on the concentration of xyloglucan oligomers in muro. Since AtXYL1 is demonstrated to remove the xylosyl residue at the non-reducing terminus of xyloglucan oligomers (Sampedro et al. 2001), and since the terminal xylosyl substitution is a prerequisite of the acceptor substrate activity of xyloglucan for XTH (Nishitani 1997), it is quite likely that AtXYL1 may suppress the depolymerization of xyloglucan by degrading or inactivating the acceptor substrate (Lorences and Fry 1993). The AtXYL1-mediated suppression of xyloglucan depolymerization would result in the generation of higher molecular weight xyloglucans outside the plasma membrane.
As we have mentioned before the AtXYL1 protein undergoes a variety of post-translational modifications throughout the cell wall assembly and remodeling processes (Tables 2–4, Fig. 2). These modifications in the AtXYL1 protein raise the possibility that differently modified AtXYL1 proteins exhibit differing enzymatic activities on xyloglucan oligomers, and thereby on the regulation of AtXTH11 activity.
Proteins involved in defense and extracellular signaling
The present study revealed that several defense-related proteins, including chitinase (At2g43610) and β-glucanase (At1g66280 and At3g09260), are secreted into the extracellular matrix of the protoplasts. Some defense-related proteins are directly involved in the structural organization of cell walls and take part in an active defense system. These defense proteins may be induced by various stresses that would be generated during protoplast preparation. Alternatively, proteins required for cell wall dynamics during cell wall regeneration in the protoplasts may be mediated by the same molecular components that are also recruited as defense proteins upon defense response. It is worth noting that the β-glucanase (At3g09260), which was found in the protoplast cell wall in the present study, has recently been identified as being a major component of the ER body, a subcellular compartment discovered recently (Matsushima et al. 2003).
The presence of phosphorylated proteins in plant cell walls has recently been reported (Chivasa et al. 2002, Baluska et al. 2003). These proteins may participate in extracellular signaling events. An acid phosphatase (At3g02750), which was identified in the present study, could be involved in such signal mediation. The phosphorylation of cell wall proteins in muro would play an important role in regulating their localizations and enzyme activities, thereby allowing multiple locations and/or functions of the proteins encoded by a single gene. It might be possible that a post-translational modification may participate in the extracellular localization of proteins without any conventional signal peptide via an unknown trafficking mechanism.
Glycolytic enzymes and other unexpected proteins found in extracellular spaces during cell wall regeneration processes
In cell wall-regenerating protoplasts, we identified several glycolytic enzymes that are known to catalyze the initial steps of the glycolytic pathway. These enzymes have never been considered to be secreted outside of the protoplasm. However, some glycolytic enzymes have already been classified as cell wall components in some organisms (Chaffin et al. 1998, Pardo et al. 2000, Chivasa et al. 2002). For example, glyceraldehyde-3-phosphate dehydrogenase (At1g13440 and At3g04120) has been immunologically localized in yeast cell walls (Gozalbo et al. 1998). Although how these glycolytic enzymes actually cross the plasma membrane has not been fully elucidated, some previous substantial data are in support of the idea that some glycolytic enzymes may be secreted out of the protoplasts during the cell wall regeneration process in A. thaliana. It should be noted that in the present study glycolytic enzymes were identified in the apoplasts around the protoplasts only, not in native cells. This means that these glycolytic enzymes may fulfill certain special functions that are unknown, but might be required for the cell wall regeneration of protoplasts.
In addition to glycolytic enzymes, we identified in the extracellular protein fractions a few proteins that have been considered present exclusively in the cytosol or in some intracellular compartments. Although we used a non-destructive method for protein extraction to minimize cross-contamination of cytosolic marker proteins (Table 1), we cannot exclude the possibility of minor contamination from cytoplasmic proteins that cannot be detected by the activity assay of G6PDH or by a simple viability test using fluorescein diacetate. The presence of 60S ribosomal protein L12 in the 1 M KCl extract might imply possible leakage of some cytoplasmic protein from the cortical region of cytoplasm during extraction of the protoplast with 1 M KCl solution. Thus, we should be careful in interpreting localization of proteins that lack a conventional secretory signal peptide. Nevertheless, apart from the 60S ribosomal protein, there is reason to believe that most proteins without known secretory signal peptides are actually secreted. Whereas the conventional theory for protein trafficking cannot explain the mechanism by which proteins without any secretory signal peptide are exported into the extracellular spaces, several proteins that lack signal peptides are reportedly localized in the extracellular matrix (Cleves 1997). For example, an ortholog of the elongation factor 1-α protein that we have identified in the present study was localized in tobacco cell walls by immunogold labeling (Zhu et al. 1994), despite the lack of a secretory signal peptide. A cell wall-bound malate dehydrogenase (Li et al. 1989, Karkonen et al. 2002), which is implicated in the regeneration of NAD(P)H required for cell wall peroxidase-mediated free radical generation associated with lignin polymerization, is another example of this interesting class of extracellular proteins without a signal peptide. Judging from this line of evidence, we conclude that most of the proteins identified in the 1 M KCl fractions have been localized in the apoplastic spaces. However, the real functions of these proteins remain unknown at present, and require further investigation.
Taken together, this proteomic approach using the cell wall regeneration system on protoplasts derived from Arabidopsis suspension-cultured cells has disclosed a set of proteins required for cell wall expansion as well as wall construction processes. Most of these proteins show a differing status of post-translational modifications such as phosphorylation and glycosylation among the three protein fractions. This indicates dynamic changes in muro of these cell wall proteins during the cell wall regeneration of protoplasts. More interestingly, the use of a non-destructive extraction procedure for cell wall proteins allowed us to avoid contamination from cytosolic fractions, thus making it possible to identify a new class of cell wall proteins that were predicted not to be localized in the cell wall spaces. These results strongly vindicate the usefulness of a proteomic approach for cell wall dynamics. Functional analyses for each of these newly identified cell wall proteins, together with their biochemical characterization and expression analyses, should extend our understanding of the whole picture of cell wall dynamism from a genomic point of view, and give an invaluable molecular basis for metabolomic approaches of this complex supermolecule.
Materials and Methods
Plant material
Suspension-cultured Alex cells of A. thaliana L. (Heynh.), ecotype Columbia, established by Mathur et al. (1998), were kindly provided by Dr. Masaaki Umeda from The University of Tokyo. The cell line was maintained in a modified Murashige and Skoog medium (Murashige and Skoog 1962) containing 3% sucrose, B5 vitamins (Gamborg et al. 1976) and 0.5 mg l–1 2,4-dichlorophenoxyacetic acid (2,4-D), adjusted to pH 5.8, at 23°C with constant shaking (120 rpm, 2 inches) in the dark. The suspension-culture was subcultured every 7 d by transfer of 15 ml of culture into 35 ml of fresh medium (Mathur et al. 1998).
Cell wall regeneration from protoplasts
Five-day-old suspension-cultured cells were collected by centrifugation at 190×g for 10 min and incubated for 1 h at 37°C in an enzyme mixture containing 1% Cellulase Onozuka RS (Yakult Pharmaceutical Ind. Co. Ltd., Tokyo, Japan) and 0.1% Pectolyase Y-23 (Kikkoman Shoyu Co. Ltd., Tokyo, Japan). This was supplemented with 120 g l–1 sorbitol and adjusted to pH 5.8 with KOH. After completion of incubation, the protoplasts were collected by centrifugation at 70×g for 10 min and washed twice with 40 ml of 120 g l–1 sorbitol. The purified protoplasts were gently resuspended in a modified cell wall regeneration medium as described by Schirawski et al. (2000) at a final density of about 105 protoplasts ml–1, and incubated for 1 or 3 h at 25°C to regenerate cell walls.
Microscopic observation
For light microscope observation, native cells, fresh protoplasts and protoplasts that had been allowed to regenerate cell walls for 1 and 3 h were fixed in 1% glutaraldehyde and stained with 0.001% Calcofluor White M2R (Sigma, St Louis, MO, USA) or aniline blue in 120 g l–1 sorbitol at 25°C (Smith and McCully 1978). Both epifluorescent images and differential interference contrast images for individual specimens were observed under a fluorescence microscope equipped with a UV fluorescence filter set (excitation filter, 350 nm; barrier filter, 430 nm) and differential interference contrast optics (DMRXP, Leica, Heerbrugg, Switzerland). The images were then photographed with a digital camera (Color Chilled 3CCD CAMERA C-5810, Hamamatsu Photonics K.K., Hamamatsu, Japan).
For electron microscopic observation, native cells and protoplasts were fixed in 2% glutaraldehyde with 120 g l–1 sorbitol and adjusted to pH 6.0 for 16 h. This was followed by an additional 30 min of fixation in 0.2% (v/v) ruthenium tetroxide (RuO4) at room temperature. After completion of fixation, specimens were dehydrated through an ethanol series and n-butyl alcohol, lyophilized (ES-2030, Hitachi Science Systems Ltd., Tokyo, Japan), coated with silver (E-1030, Hitachi Science Systems Ltd.), and observed with an SEM (S-4100, Hitachi Science Systems Ltd.) at 3 kV according to a similar procedure described by Konomi et al. (2000).
G6PDH activity assay
The activity of G6PDH (EC 1.1.1.49) was examined according to the procedure described by Weimar and Rothe (1986), with some modifications. Briefly, protein extracts were mixed with a substrate solution containing 1 mM glucose-6-phosphate, 0.2 mM NADP and 1 mM MgCl2. In 0.1 M Tris, the extracts were adjusted to pH 8.0 and incubated at 25°C for 60 min. During the incubation period, the reduction of NADP in the reaction mixture was measured by its absorbance at 340 nm using a spectrometer (UV-160A, Shimadzu Co., Kyoto, Japan).
Viability assay
The viability of native cells and protoplasts that had been extracted with 1 M KCl was examined by staining with FDA (Ishikawa et al. 1995, Amano et al. 2003). Briefly, cells or protoplasts were mixed with 12 µM FDA in a 120 g l–1 sorbitol solution with gentle shaking and kept for 5 min in the dark at 25°C. Viable cells and protoplasts were counted under a fluorescence microscope equipped with a UV fluorescence filter set (excitation filter, 490 nm; barrier filter, 514 nm).
Protein extraction and separation by two-dimensional gel electrophoresis
To extract extracellular proteins ionically bound to the cell wall, 5-day-old suspension-cultured cells were pelleted by centrifugation at 60×g for 10 min, resuspended in an equal volume of 1 M KCl and kept under a reduced pressure at –70 kP at 25°C for 10 min (Vanacker et al. 1998, Ito and Nishitani 1999, Souza and MacAdam 2001). This was followed by centrifugation to obtain the supernatant. Extracellular proteins on the surface of the protoplasts were also extracted with essentially the same procedure. An exception was that extraction with 1 M KCl was conducted under atmospheric pressure by gently stirring the protoplasts. This extraction procedure was repeated twice. The extracts were combined and reduced to 1/100 volume by ultrafiltration (Amicon Ultra-4 10000MWCO, Millipore, Milford, MA, USA). Proteins were precipitated with the addition of 9 vol of 80% acetone containing 10% trichloroacetic acid and 200 mM dithiothreitol (DTT), pelleted by centrifugation at 20,400×g for 30 min, washed three times with 80% acetone, and resuspended in 2D-PAGE sample buffer (5 M urea, 2 M thiourea, 2% 3-[(3-cholamidopropyl)-dimethylammonio]-1-propanesulfonate (CHAPS), 2% SB3-10, 2 mM tributyl phosphate (TBP), 40 mM Tris base and 0.5% Pharmalyte; pH 3–10 and 6–11). After the protein contents were determined using a modification of the method of Bradford (1976), a 180 µg protein portion was loaded into 13 cm isoelectric focusing (IEF) gel strips (Amersham Pharmacia Biotech, Uppsala, Sweden) with pH ranges of 3–10 or 6–11. These were rehydrated according to the procedure provided by the manufacturer (Rabilloud et al. 1994). Electrofocusing was performed at 17 kV h–1 using a Multiphor II system (Amersham Pharmacia Biotech), equilibrated for 15 min in a buffer (6 M urea, 30% glycerol, 1% SDS, and 0.5 M Tris–HCl, pH 6.8) containing 1% DTT, and for another 10 min in the same buffer containing 30 mM iodoacetamide. After equilibration, the IEF gel strips were loaded on top of 12% polyacrylamide gels (160×160×1 mm) for SDS–PAGE, which was performed using the 2-D gel system (Nihon Eido Co., Ltd, Tokyo, Japan; Laemmli 1970). The gels were stained with CBB.
Protein identification by MALDI-TOF/MS
Proteins separated by the 2D-PAGE were mapped using Image Master 2D-Elite software (Amersham Pharmacia Biotech). All protein spots were picked from the 2D-polyacrylamide gels, washed in 25 mM bicarbonate buffer, destained, and desiccated in the presence of acetonitrile. Each gel piece was then rehydrated in 25 µl of 50 mM bicarbonate buffer containing 200 ng of modified trypsin (Promega, Madison, WI, USA), followed by incubation for 16 h at 37°C. Peptides released by trypsin digestion were extracted from the gel piece with 75 µl of 50% acetonitrile solution containing 5% trifluoroacetic acid followed by evaporation under a reduced pressure using Speed Vac (SC110A, Savant Instruments, Inc., NY, USA) to reduce the concentration to about 20 µl. After the concentrated samples were treated with ZipTip C18 (Milipore, Milford, MA, USA) and mixed with 1 µl of an α-cyano-4-hydroxycinnamic acid (α-CHCA) matrix, they were subjected to MALDI-TOF/MS analysis (Applied Biosystems, Foster City, CA, USA). The mass spectrum of the peptide derived from each protein spot was recorded using MALDI-TOF/MS (Applied Biosystems, Voyager DE-STR). Each peptide mass was matched to the theoretical tryptic digests of proteins predicted based on the complete non-redundant Arabidopsis database using the Mascot sequence database search program (Matrix Science, Boston, MA, USA). We selected sequence results that matched all conditions as follows: (i) the sequence coverage was >30%; (ii) the mass error margin was within 50 ppm; and (iii) the molecular weight search (MOWSE) score was significantly (P < 0.05) greater than 76.
Data analysis
The presence of putative cleavable signal peptides or non-cleavable signal anchors was predicted using PSORT (http://psort.nibb.ac.jp/form.html), Target P (http://www.cbs.dtu.dk/services/TargetP/) and the Signal-P V2.0.b2 program (http://www.cbs.dtu.dk/services/SignalP/; Nielsen et al. 1997). The amino acid sequences of hypothetical proteins were also analyzed using the TMHMM prediction method (http://www.cbs.dtu.dk/services/TMHMM/) for transmembrane helices (Sonnhammer et al. 1998). Homologies to other proteins were searched using BLAST programs (http://www.ncbi.nlm.nih.gov/BLAST/) and TAIR (http://www.arabidopsis.org). The search of protein domains was performed using Pfam (http://www.sanger.ac.uk/software/Pfam) and InterPro (http://www.ebi.au.uk/iterpro).
Characterization of post-translational modifications
Phosphorylated proteins on 2-D polyacrylamide gels were determined using the Pro-Q Diamond phosphoprotein gel stain kit (Molecular Probes; Schulenberg et al. 2003, Hart et al. 2003). Briefly, after the gel was fixed in a solution containing 50% methanol and 10% trichloroacetic acid overnight, it was stained in a Pro-Q Diamond phosphoprotein gel stain solution for 3 h in the dark, followed by destaining with successive washes of 2% acetonitrile in 25 mM sodium acetate, pH 4.0, for 1 h. Protein spots on the Pro-Q-stained gels were determined by staining with CBB. Fluorescent images of the Pro-Q-stained gels were acquired using a Fuji FLA 3000 laser scanner (Fuji Photo Film Co., Ltd., Tokyo, Japan) with a 532 nm excitation and 580 nm emission filter for Pro-Q Diamond dye detection and total protein detection.
Glycosylated proteins on 2-D polyacrylamide gels were determined using the Pro-Q Emerald 488 glycoprotein gel staining kit (Molecular Probes; Hart et al. 2003). Briefly, after 2-D polyacrylamide gels were fixed in a solution containing 50% methanol and 5% acetic acid, and extensively washed in 3% acetic acid, glycans were oxidized by incubation in a solution containing 1% periodic acid and 3% acetic acid for 1 h. After the gels were washed in 3% acetic acid, they were incubated in a Pro-Q Emerald 488 dye solution for 2.5 h in the dark, followed by incubation in 3% acetic acid. The gels were then stained for total protein with a CBB stain. Fluorescence images were acquired on a Fuji FLA 3000 laser scanner (Fuji Photo Film Co., Ltd.) with 473 nm excitation and a 520 nm emission filter for Pro-Q Emerald 488 dye detection and total protein detection.
Acknowledgments
The authors thank Dr. Csaba Koncz (Max-Planck-Institut fur Zuchtungsforschung) and Dr. Masaaki Umeda (The University of Tokyo) for kindly providing the Arabidopsis thaliana suspension-cultured Alex cell line. This work was supported by a Grant-in-Aid for Scientific Research on Priority Areas (15031202) and Scientific Research (B) (15370016) from the Ministry of Education, Culture, Sports, Science, and Technology of Japan, and by the Program for ‘Development of Fundamental Technologies for Controlling the Process of Material Production of Plants’ from the New Energy and Industrial Technology Development Organization, Japan.
Fig. 1 Time course of changes in the cell surface during cell wall regeneration from the protoplasts of Arabidopsis suspension-cultured cells. Cells and protoplasts stained with Calcofluor White M2R (E–H) and aniline blue (I-L) were observed under a differential interference contrast microscope (A–D) and an epifluorescent microscope (E–L). For scanning electron microscopic observation (M–P), cells and protoplasts were fixed in glutaraldehyde and ruthenium tetroxide, and then coated with silver. Native suspension-cultured cells (A, E, I and M), fresh protoplasts (B, F, J and N), protoplasts regenerated for 1 h (C, G, K and O) and those regenerated for 3 h (D, H, L and P) are shown. Bar: 25 µm (A–L); 1.5 µm (M–P)
Fig. 2 2-D PAGE patterns of cell wall proteins extracted with 1 M KCl from native cell walls of suspension-cultured cells (A and B), from protoplast cell walls that were regenerated for 1 h (C and D) and those regenerated for 3 h (E and F). An isoelectric point (pI) gradient strip with pI 3–10 (A, C and E) or pI 6–11 (B, D and F) was loaded with 180 µg of total cell wall proteins. Protein spots disclosed by CBB staining were mapped, identified, and summarized in Tables 2–4, respectively.
Fig. 3 Summary of the number of protein spots identified in the apoplastic fraction derived from suspension-cultured native cells (native), protoplast cell walls that had been regenerated for 1 h (1 hour), and those regenerated for 3 h (3 hours). Numbers in parentheses indicate the total number of protein spots found in each fraction.
Examination of cytosolic contamination in the cell wall protein fraction as assessed by the cytosolic protein marker, glucose-6-phosphate dehydrogenase (G6PDH)
| Fraction | Total protein (mg flask–1) | Specific activity (mU mg–1 protein) | Total activity (mU flask–1) |
| Cytosolic protein from native cells | 4.28 | 375.70 | 1607.4 |
| Cell wall protein from native cells | 0.74 | 5.68 | 4.2 |
| Cytosolic protein from nascent protoplasts | 6.09 | 371.58 | 2264.7 |
| Cell wall protein from 1 h regenerated protoplasts | 0.58 | 1.03 | 0.6 |
| Cell wall protein from 3 h regenerated protoplasts | 0.51 | 4.04 | 2.0 |
| Fraction | Total protein (mg flask–1) | Specific activity (mU mg–1 protein) | Total activity (mU flask–1) |
| Cytosolic protein from native cells | 4.28 | 375.70 | 1607.4 |
| Cell wall protein from native cells | 0.74 | 5.68 | 4.2 |
| Cytosolic protein from nascent protoplasts | 6.09 | 371.58 | 2264.7 |
| Cell wall protein from 1 h regenerated protoplasts | 0.58 | 1.03 | 0.6 |
| Cell wall protein from 3 h regenerated protoplasts | 0.51 | 4.04 | 2.0 |
Cytosolic proteins were extracted with 1 M KCl after cells or protoplasts were disrupted by homogenization, while cell wall proteins were extracted with 1 M KCl without disrupting cellular integrity. A unit of G6PDH activity is given as 1 µmol of substrate turnover per minute.
Examination of cytosolic contamination in the cell wall protein fraction as assessed by the cytosolic protein marker, glucose-6-phosphate dehydrogenase (G6PDH)
| Fraction | Total protein (mg flask–1) | Specific activity (mU mg–1 protein) | Total activity (mU flask–1) |
| Cytosolic protein from native cells | 4.28 | 375.70 | 1607.4 |
| Cell wall protein from native cells | 0.74 | 5.68 | 4.2 |
| Cytosolic protein from nascent protoplasts | 6.09 | 371.58 | 2264.7 |
| Cell wall protein from 1 h regenerated protoplasts | 0.58 | 1.03 | 0.6 |
| Cell wall protein from 3 h regenerated protoplasts | 0.51 | 4.04 | 2.0 |
| Fraction | Total protein (mg flask–1) | Specific activity (mU mg–1 protein) | Total activity (mU flask–1) |
| Cytosolic protein from native cells | 4.28 | 375.70 | 1607.4 |
| Cell wall protein from native cells | 0.74 | 5.68 | 4.2 |
| Cytosolic protein from nascent protoplasts | 6.09 | 371.58 | 2264.7 |
| Cell wall protein from 1 h regenerated protoplasts | 0.58 | 1.03 | 0.6 |
| Cell wall protein from 3 h regenerated protoplasts | 0.51 | 4.04 | 2.0 |
Cytosolic proteins were extracted with 1 M KCl after cells or protoplasts were disrupted by homogenization, while cell wall proteins were extracted with 1 M KCl without disrupting cellular integrity. A unit of G6PDH activity is given as 1 µmol of substrate turnover per minute.
Summary of the proteins identified as 1M KCl extractable from the native cell walls of Arabidopsis suspension cultured cells
| Protein spot number a | Accession no. b | Protein name | Theoretical mol. wt/pI | TMD c | Signal peptide d | 1 h e | 3 h |
| 146*,174 | At3g45970 | Expansin | 29253/8.28 | 2 | + | ||
| 147 | At5g24090 | Chitinase | 33501/9.22 | 0 | + | ||
| 59 | At3g48580 | Xyloglucan endotransglucosylase/hydrolase | 31062/8.62 | 0 | + | † | † |
| 148 | At1g71380 | β-Glucanase | 53638/8.81 | 0 | + | ||
| 149 | At2g44450 | β-Glucosidase | 57211/7.56 | 0 | + | ||
| 150*,151*,152 | At1g68560 | α-Xylosidase | 102961/6.31 | 0 | + | † | † |
| 153,154*,155*, 156 | 102210/6.32 | 0 | + | † | † | ||
| 175* | At5g20950 | β-Xylosidase | 67910/9.4 | 0 | + | † | † |
| 157*,158 | At5g08380 | α-Galactosidase | 46079/6.19 | 0 | + | † | |
| 95* | At5g63810 | β-Galactosidase | 83619/8.58 | 1 | + | † | † |
| 54* | At3g13790 | β-Fructosidase | 66255/9.19 | 1 | + | † | † |
| 159*,160 | At3g26720 | α-Mannosidase | 115547/6.27 | 1 | + | † | † |
| 176 | At5g41870 | Polygalacturonase | 49013/6.89 | 0 | + | ||
| 161,177 | At5g06860 | Polygalacturonase | 37293/8.24 | 0 | + | ||
| 178,179* | At3g54400 | Aspartyl protease | 46259/9.31 | 0 | + | ||
| 162 | At5g67360 | Serine protease | 80024/5.91 | 0 | + | ||
| 163* | At3g02750 | Phosphatase-2C | 54332/5.04 | 0 | – | ||
| 184* | At1g13750 | Phosphoesterase | 68826/6.29 | 0 | + | † | |
| 49* | At4g34870 | Peptidylprolyl isomerase | 18594/8.90 | 0 | – | † | † |
| 57 | At1g30730 | FAD-linked oxidoreductase | 59161/9.23 | 0 | + | † | † |
| 56* | At5g44380 | FAD-linked oxidoreductase | 60920/9.51 | 1 | + | † | † |
| 164 | At5g18860 | Nucleoside hydrolase | 99580/7.59 | 0 | + | † | |
| 165 | At2g02990 | Ribonuclease | 26007/5.08 | 1 | + | ||
| 166 | At3g47800 | Aldose 1-epimerase | 39861/9.12 | 1 | + | ||
| 48* | At4g25900 | Aldose 1-epimerase | 36229/9.65 | 1 | + | † | † |
| 167,168 | At1g03220 | Glycoprotein | 46373/8.98 | 0 | + | ||
| 77 | At1g03230 | Glycoprotein | 46803/9.33 | 0 | + | † | |
| 99*,180* | At1g53070 | Legume lectin | 30416/8.49 | 1 | + | † | † |
| 110 | At3g16530 | Legume lectin | 30547/6.97 | 0 | + | † | |
| 85 | At3g16420 | Jacalin lectin | 32096/5.46 | 0 | – | † | † |
| 87* | At3g16450 | Jacalin lectin | 32003/5.06 | 0 | – | † | † |
| 35*,62*,71 | At1g78850 | Curculin-like lectin | 49818/7.82 | 0 | + | † | † |
| 182,183* | At1g33590 | Leucine-rich repeat protein | 561123/9.10 | 0 | + | ||
| 169 | At1g21670 | Unknown | 77468/9.33 | 0 | + | ||
| 170 | At1g51630 | Unknown | 47454/6.44 | 1 | + | ||
| 171 | At2g32330 | Unknown | 35134/8.61 | 0 | – | ||
| 172 | At2g41800 | Unknown | 40630/9.22 | 0 | + | ||
| 181,173 | At3g08030 | Unknown | 39326/7.17 | 1 | + | ||
| 50 | At5g26260 | Unknown | 839802/9.21 | 0 | + | † | † |
| Protein spot number a | Accession no. b | Protein name | Theoretical mol. wt/pI | TMD c | Signal peptide d | 1 h e | 3 h |
| 146*,174 | At3g45970 | Expansin | 29253/8.28 | 2 | + | ||
| 147 | At5g24090 | Chitinase | 33501/9.22 | 0 | + | ||
| 59 | At3g48580 | Xyloglucan endotransglucosylase/hydrolase | 31062/8.62 | 0 | + | † | † |
| 148 | At1g71380 | β-Glucanase | 53638/8.81 | 0 | + | ||
| 149 | At2g44450 | β-Glucosidase | 57211/7.56 | 0 | + | ||
| 150*,151*,152 | At1g68560 | α-Xylosidase | 102961/6.31 | 0 | + | † | † |
| 153,154*,155*, 156 | 102210/6.32 | 0 | + | † | † | ||
| 175* | At5g20950 | β-Xylosidase | 67910/9.4 | 0 | + | † | † |
| 157*,158 | At5g08380 | α-Galactosidase | 46079/6.19 | 0 | + | † | |
| 95* | At5g63810 | β-Galactosidase | 83619/8.58 | 1 | + | † | † |
| 54* | At3g13790 | β-Fructosidase | 66255/9.19 | 1 | + | † | † |
| 159*,160 | At3g26720 | α-Mannosidase | 115547/6.27 | 1 | + | † | † |
| 176 | At5g41870 | Polygalacturonase | 49013/6.89 | 0 | + | ||
| 161,177 | At5g06860 | Polygalacturonase | 37293/8.24 | 0 | + | ||
| 178,179* | At3g54400 | Aspartyl protease | 46259/9.31 | 0 | + | ||
| 162 | At5g67360 | Serine protease | 80024/5.91 | 0 | + | ||
| 163* | At3g02750 | Phosphatase-2C | 54332/5.04 | 0 | – | ||
| 184* | At1g13750 | Phosphoesterase | 68826/6.29 | 0 | + | † | |
| 49* | At4g34870 | Peptidylprolyl isomerase | 18594/8.90 | 0 | – | † | † |
| 57 | At1g30730 | FAD-linked oxidoreductase | 59161/9.23 | 0 | + | † | † |
| 56* | At5g44380 | FAD-linked oxidoreductase | 60920/9.51 | 1 | + | † | † |
| 164 | At5g18860 | Nucleoside hydrolase | 99580/7.59 | 0 | + | † | |
| 165 | At2g02990 | Ribonuclease | 26007/5.08 | 1 | + | ||
| 166 | At3g47800 | Aldose 1-epimerase | 39861/9.12 | 1 | + | ||
| 48* | At4g25900 | Aldose 1-epimerase | 36229/9.65 | 1 | + | † | † |
| 167,168 | At1g03220 | Glycoprotein | 46373/8.98 | 0 | + | ||
| 77 | At1g03230 | Glycoprotein | 46803/9.33 | 0 | + | † | |
| 99*,180* | At1g53070 | Legume lectin | 30416/8.49 | 1 | + | † | † |
| 110 | At3g16530 | Legume lectin | 30547/6.97 | 0 | + | † | |
| 85 | At3g16420 | Jacalin lectin | 32096/5.46 | 0 | – | † | † |
| 87* | At3g16450 | Jacalin lectin | 32003/5.06 | 0 | – | † | † |
| 35*,62*,71 | At1g78850 | Curculin-like lectin | 49818/7.82 | 0 | + | † | † |
| 182,183* | At1g33590 | Leucine-rich repeat protein | 561123/9.10 | 0 | + | ||
| 169 | At1g21670 | Unknown | 77468/9.33 | 0 | + | ||
| 170 | At1g51630 | Unknown | 47454/6.44 | 1 | + | ||
| 171 | At2g32330 | Unknown | 35134/8.61 | 0 | – | ||
| 172 | At2g41800 | Unknown | 40630/9.22 | 0 | + | ||
| 181,173 | At3g08030 | Unknown | 39326/7.17 | 1 | + | ||
| 50 | At5g26260 | Unknown | 839802/9.21 | 0 | + | † | † |
a Protein spot numbers correspond to the numbers on gel images (Fig. 2A, B). An asterisk indicates that the protein spot is phosphorylated, while underlined spots are glycosylated.
b Accession numbers are based on the NCBI database.
c The number of transmembrane domains (TMDs) as predicted by TMHMM is shown.
d The potential signal peptide as predicted by iPSORT, Signal P and Target P is represented by +.
e † in the 1 h and 3 h columns indicates that proteins encoded by the same gene are also identified in the 1 and 3 h regenerated protoplasts, respectively.
Summary of the proteins identified as 1M KCl extractable from the native cell walls of Arabidopsis suspension cultured cells
| Protein spot number a | Accession no. b | Protein name | Theoretical mol. wt/pI | TMD c | Signal peptide d | 1 h e | 3 h |
| 146*,174 | At3g45970 | Expansin | 29253/8.28 | 2 | + | ||
| 147 | At5g24090 | Chitinase | 33501/9.22 | 0 | + | ||
| 59 | At3g48580 | Xyloglucan endotransglucosylase/hydrolase | 31062/8.62 | 0 | + | † | † |
| 148 | At1g71380 | β-Glucanase | 53638/8.81 | 0 | + | ||
| 149 | At2g44450 | β-Glucosidase | 57211/7.56 | 0 | + | ||
| 150*,151*,152 | At1g68560 | α-Xylosidase | 102961/6.31 | 0 | + | † | † |
| 153,154*,155*, 156 | 102210/6.32 | 0 | + | † | † | ||
| 175* | At5g20950 | β-Xylosidase | 67910/9.4 | 0 | + | † | † |
| 157*,158 | At5g08380 | α-Galactosidase | 46079/6.19 | 0 | + | † | |
| 95* | At5g63810 | β-Galactosidase | 83619/8.58 | 1 | + | † | † |
| 54* | At3g13790 | β-Fructosidase | 66255/9.19 | 1 | + | † | † |
| 159*,160 | At3g26720 | α-Mannosidase | 115547/6.27 | 1 | + | † | † |
| 176 | At5g41870 | Polygalacturonase | 49013/6.89 | 0 | + | ||
| 161,177 | At5g06860 | Polygalacturonase | 37293/8.24 | 0 | + | ||
| 178,179* | At3g54400 | Aspartyl protease | 46259/9.31 | 0 | + | ||
| 162 | At5g67360 | Serine protease | 80024/5.91 | 0 | + | ||
| 163* | At3g02750 | Phosphatase-2C | 54332/5.04 | 0 | – | ||
| 184* | At1g13750 | Phosphoesterase | 68826/6.29 | 0 | + | † | |
| 49* | At4g34870 | Peptidylprolyl isomerase | 18594/8.90 | 0 | – | † | † |
| 57 | At1g30730 | FAD-linked oxidoreductase | 59161/9.23 | 0 | + | † | † |
| 56* | At5g44380 | FAD-linked oxidoreductase | 60920/9.51 | 1 | + | † | † |
| 164 | At5g18860 | Nucleoside hydrolase | 99580/7.59 | 0 | + | † | |
| 165 | At2g02990 | Ribonuclease | 26007/5.08 | 1 | + | ||
| 166 | At3g47800 | Aldose 1-epimerase | 39861/9.12 | 1 | + | ||
| 48* | At4g25900 | Aldose 1-epimerase | 36229/9.65 | 1 | + | † | † |
| 167,168 | At1g03220 | Glycoprotein | 46373/8.98 | 0 | + | ||
| 77 | At1g03230 | Glycoprotein | 46803/9.33 | 0 | + | † | |
| 99*,180* | At1g53070 | Legume lectin | 30416/8.49 | 1 | + | † | † |
| 110 | At3g16530 | Legume lectin | 30547/6.97 | 0 | + | † | |
| 85 | At3g16420 | Jacalin lectin | 32096/5.46 | 0 | – | † | † |
| 87* | At3g16450 | Jacalin lectin | 32003/5.06 | 0 | – | † | † |
| 35*,62*,71 | At1g78850 | Curculin-like lectin | 49818/7.82 | 0 | + | † | † |
| 182,183* | At1g33590 | Leucine-rich repeat protein | 561123/9.10 | 0 | + | ||
| 169 | At1g21670 | Unknown | 77468/9.33 | 0 | + | ||
| 170 | At1g51630 | Unknown | 47454/6.44 | 1 | + | ||
| 171 | At2g32330 | Unknown | 35134/8.61 | 0 | – | ||
| 172 | At2g41800 | Unknown | 40630/9.22 | 0 | + | ||
| 181,173 | At3g08030 | Unknown | 39326/7.17 | 1 | + | ||
| 50 | At5g26260 | Unknown | 839802/9.21 | 0 | + | † | † |
| Protein spot number a | Accession no. b | Protein name | Theoretical mol. wt/pI | TMD c | Signal peptide d | 1 h e | 3 h |
| 146*,174 | At3g45970 | Expansin | 29253/8.28 | 2 | + | ||
| 147 | At5g24090 | Chitinase | 33501/9.22 | 0 | + | ||
| 59 | At3g48580 | Xyloglucan endotransglucosylase/hydrolase | 31062/8.62 | 0 | + | † | † |
| 148 | At1g71380 | β-Glucanase | 53638/8.81 | 0 | + | ||
| 149 | At2g44450 | β-Glucosidase | 57211/7.56 | 0 | + | ||
| 150*,151*,152 | At1g68560 | α-Xylosidase | 102961/6.31 | 0 | + | † | † |
| 153,154*,155*, 156 | 102210/6.32 | 0 | + | † | † | ||
| 175* | At5g20950 | β-Xylosidase | 67910/9.4 | 0 | + | † | † |
| 157*,158 | At5g08380 | α-Galactosidase | 46079/6.19 | 0 | + | † | |
| 95* | At5g63810 | β-Galactosidase | 83619/8.58 | 1 | + | † | † |
| 54* | At3g13790 | β-Fructosidase | 66255/9.19 | 1 | + | † | † |
| 159*,160 | At3g26720 | α-Mannosidase | 115547/6.27 | 1 | + | † | † |
| 176 | At5g41870 | Polygalacturonase | 49013/6.89 | 0 | + | ||
| 161,177 | At5g06860 | Polygalacturonase | 37293/8.24 | 0 | + | ||
| 178,179* | At3g54400 | Aspartyl protease | 46259/9.31 | 0 | + | ||
| 162 | At5g67360 | Serine protease | 80024/5.91 | 0 | + | ||
| 163* | At3g02750 | Phosphatase-2C | 54332/5.04 | 0 | – | ||
| 184* | At1g13750 | Phosphoesterase | 68826/6.29 | 0 | + | † | |
| 49* | At4g34870 | Peptidylprolyl isomerase | 18594/8.90 | 0 | – | † | † |
| 57 | At1g30730 | FAD-linked oxidoreductase | 59161/9.23 | 0 | + | † | † |
| 56* | At5g44380 | FAD-linked oxidoreductase | 60920/9.51 | 1 | + | † | † |
| 164 | At5g18860 | Nucleoside hydrolase | 99580/7.59 | 0 | + | † | |
| 165 | At2g02990 | Ribonuclease | 26007/5.08 | 1 | + | ||
| 166 | At3g47800 | Aldose 1-epimerase | 39861/9.12 | 1 | + | ||
| 48* | At4g25900 | Aldose 1-epimerase | 36229/9.65 | 1 | + | † | † |
| 167,168 | At1g03220 | Glycoprotein | 46373/8.98 | 0 | + | ||
| 77 | At1g03230 | Glycoprotein | 46803/9.33 | 0 | + | † | |
| 99*,180* | At1g53070 | Legume lectin | 30416/8.49 | 1 | + | † | † |
| 110 | At3g16530 | Legume lectin | 30547/6.97 | 0 | + | † | |
| 85 | At3g16420 | Jacalin lectin | 32096/5.46 | 0 | – | † | † |
| 87* | At3g16450 | Jacalin lectin | 32003/5.06 | 0 | – | † | † |
| 35*,62*,71 | At1g78850 | Curculin-like lectin | 49818/7.82 | 0 | + | † | † |
| 182,183* | At1g33590 | Leucine-rich repeat protein | 561123/9.10 | 0 | + | ||
| 169 | At1g21670 | Unknown | 77468/9.33 | 0 | + | ||
| 170 | At1g51630 | Unknown | 47454/6.44 | 1 | + | ||
| 171 | At2g32330 | Unknown | 35134/8.61 | 0 | – | ||
| 172 | At2g41800 | Unknown | 40630/9.22 | 0 | + | ||
| 181,173 | At3g08030 | Unknown | 39326/7.17 | 1 | + | ||
| 50 | At5g26260 | Unknown | 839802/9.21 | 0 | + | † | † |
a Protein spot numbers correspond to the numbers on gel images (Fig. 2A, B). An asterisk indicates that the protein spot is phosphorylated, while underlined spots are glycosylated.
b Accession numbers are based on the NCBI database.
c The number of transmembrane domains (TMDs) as predicted by TMHMM is shown.
d The potential signal peptide as predicted by iPSORT, Signal P and Target P is represented by +.
e † in the 1 h and 3 h columns indicates that proteins encoded by the same gene are also identified in the 1 and 3 h regenerated protoplasts, respectively.
Summary of the proteins identified as 1 M KCl extractable from the 1 h regenerated protoplasts
| Protein spot number a | Accession no. b | Protein name | Theoretical mol. wt/pI | TMD c | Signal peptide d | 3 h e | Native |
| 111 | At2g43610 | Chitinase | 30892/9.54 | 1 | + | ||
| 59 | At3g48580 | Xyloglucan endotransglucosylase/hydrolase | 32204/8.76 | 0 | + | † | † |
| 32 | At1g66280 | β-Glucosidase | 60142/6.74 | 0 | + | ||
| 2*, 3*, 104*, 114* 115*, 116* | At1g68560 | α-Xylosidase | 102210/6.32 | 0 | + | † | † |
| 52* | At5g20950 | β-Xylosidase | 67910/9.49 | 0 | + | † | † |
| 117 | At5g08380 | α-Galactosidase | 46079/6.19 | 0 | + | † | † |
| 95*, 96, 118* | At5g63810 | β-Galactosidase | 83619/8.58 | 1 | + | † | † |
| 51, 54*, 73, 100 | At3g13790 | β-Fructosidase | 66581/9.14 | 1 | + | † | |
| 119* | At3g26720 | α-Mannosidase | 115547/6.27 | 1 | + | † | † |
| 63 | At5g19440 | Cinnamyl-alcohol dehydrogenase | 35914/6.67 | 0 | – | † | |
| 106 | At2g04160 | Serine protease | 81364/9.34 | 0 | + | ||
| 120* | At5g10540 | Oligopeptidase | 79223/5.45 | 0 | – | ||
| 121 | At1g63770 | Aminopeptidase | 99495/5.43 | 0 | – | ||
| 185, 186* | At1g13750 | Phosphoesterase | 68826/6.29 | 0 | + | † | |
| 42* | At4g24340 | Phosphorylase | 36024/8.24 | 1 | + | † | |
| 101 | At3g27950 | GDSL-motif lipase/hydrolase | 40120/8.89 | 0 | + | † | |
| 49 | At4g34870 | Peptidylprolyl isomerase | 18594/8.90 | 0 | – | † | † |
| 69 | At4g38740 | Peptidylprolyl isomerase | 18589/7.68 | 0 | – | † | |
| 57 | At1g30730 | FAD-linked oxidoreductase | 59161/9.23 | 0 | + | † | † |
| 56* | At5g44380 | FAD-linked oxidoreductase | 60920/9.51 | 1 | + | † | † |
| 17, 122, 123* | At3g55440 | Triosephosphate isomerase | 27380/5.39 | 0 | – | † | |
| 102 | At1g74000 | Strictosidine synthase | 34819/9.66 | 0 | + | ||
| 76 | At5g04590 | Sulfite reductase | 72391/8.51 | 0 | – | † | |
| 124 | At4g25100 | Iron superoxide dismutase | 23776/6.06 | 0 | – | ||
| 98, 125 | At5g43940 | Alcohol dehydrogenase | 41528/6.51 | 0 | – | † | |
| 89* | At3g48000 | Aldehyde dehydrogenase | 58951/7.11 | 0 | – | † | |
| 126 | At2g41530 | Esterase | 30759/5.91 | 0 | – | ||
| 10* | At2g31390 | Fructokinase | 35424/5.31 | 0 | – | † | |
| 48, 127 | At4g25900 | Aldose 1-epimerase | 36229/9.65 | 1 | + | † | † |
| 11, 21 | At5g07440 | Glutamate dehydrogenase | 45013/6.07 | 0 | – | † | |
| 41, 12 | At1g78380 | Glutathione transferase | 25691/5.80 | 0 | – | † | |
| 58, 105 | At1g65590 | β-Hexosaminidase | 60375/8.30 | 1 | + | † | |
| 22, 30, 60*, 103 | At1g13440 | Glyceraldehyde-3-phosphate dehydrogenase | 37004/6.67 | 0 | + | † | |
| 31 | At3g04120 | Glyceraldehyde-3-phosphate dehydrogenase | 36897/7.15 | 0 | + | ||
| 75 | At3g02360 | 6-Phosphogluconate dehydrogenase | 53829/7.02 | 0 | + | † | |
| 55, 74, 128* | At4g37870 | Phosphoenolpyruvate carboxykinase | 73929/6.61 | 0 | – | † | |
| 129 | At1g79550 | Phosphoglycerate kinase | 42162/5.49 | 0 | – | ||
| 130 | At1g80050 | Adenine phosphoribosyltransferase | 21241/5.69 | 0 | – | ||
| 1*, 28, 113, 131*, 132*, 133 | At5g17920 | Methionine synthase | 84646/6.09 | 0 | – | † | |
| 134 | At3g52880 | Monodehydroascorbate reductase | 46629/6.41 | 0 | – | † | |
| 135* | At5g03630 | Monodehydroascorbate reductase | 47408/5.18 | 0 | – | † | |
| 64*, 109 | At3g22110 | 20S proteasome α-subunit C | 27475/7.14 | 0 | – | † | |
| 66, 65 | At5g66140 | 20S proteasome α-subunit D2 | 27306/8.96 | 0 | – | † | |
| 20, 72* | At1g56450 | 20S proteasome β-subunit G1 | 27691/6.09 | 0 | – | † | |
| 67, 99, 136, 137 | At1g53070 | Legume lectin | 30416/8.49 | 1 | + | † | † |
| 110 | At3g16530 | Legume lectin | 30547/6.97 | 0 | + | † | |
| 85*, 138, 139* | At3g16420 | Jacalin lectin | 32138/5.46 | 0 | – | † | † |
| 26, 86, 140 | At3g16430 | Jacalin lectin | 32138/5.86 | 0 | – | † | |
| 87* | At3g16450 | Jacalin lectin | 32003/5.06 | 0 | – | † | † |
| 38 | At3g16460 | Jacalin lectin | 72430/5.31 | 0 | – | † | |
| 35*, 62*, 71, 80*, 112, 140*, 141*, 142, 143* | At1g78850 | Curculin-like lectin | 49818/7.82 | 0 | + | † | † |
| 29, 82 | At1g54000 | Myrosinase-associated protein | 43498/6.65 | 1 | + | † | |
| 144 | At1g54010 | Myrosinase-associated protein | 43572/8.33 | 1 | + | † | |
| 107 | At5g12940 | Leucine-rich repeat protein | 40497/9.52 | 1 | + | ||
| 61 | At1g18080 | WD-40 repeat protein | 36124/9.62 | 0 | – | † | |
| 53 | At1g07930 | Elongation factor 1-α | 49781/9.15 | 0 | – | † | |
| 68 | At3g53430 | 60S ribosomal protein L12 | 18073/9.05 | 0 | – | † | |
| 145 | At1g21350 | Unknown | 28413/9.14 | 0 | + | ||
| 70 | At4g09320 | Unknown | 15797/6.84 | 0 | – | ||
| 50 | At5g26260 | Unknown | 39802/9.21 | 0 | + | † | † |
| 108 | At5g26280 | Unknown | 39421/8.46 | 1 | + |
| Protein spot number a | Accession no. b | Protein name | Theoretical mol. wt/pI | TMD c | Signal peptide d | 3 h e | Native |
| 111 | At2g43610 | Chitinase | 30892/9.54 | 1 | + | ||
| 59 | At3g48580 | Xyloglucan endotransglucosylase/hydrolase | 32204/8.76 | 0 | + | † | † |
| 32 | At1g66280 | β-Glucosidase | 60142/6.74 | 0 | + | ||
| 2*, 3*, 104*, 114* 115*, 116* | At1g68560 | α-Xylosidase | 102210/6.32 | 0 | + | † | † |
| 52* | At5g20950 | β-Xylosidase | 67910/9.49 | 0 | + | † | † |
| 117 | At5g08380 | α-Galactosidase | 46079/6.19 | 0 | + | † | † |
| 95*, 96, 118* | At5g63810 | β-Galactosidase | 83619/8.58 | 1 | + | † | † |
| 51, 54*, 73, 100 | At3g13790 | β-Fructosidase | 66581/9.14 | 1 | + | † | |
| 119* | At3g26720 | α-Mannosidase | 115547/6.27 | 1 | + | † | † |
| 63 | At5g19440 | Cinnamyl-alcohol dehydrogenase | 35914/6.67 | 0 | – | † | |
| 106 | At2g04160 | Serine protease | 81364/9.34 | 0 | + | ||
| 120* | At5g10540 | Oligopeptidase | 79223/5.45 | 0 | – | ||
| 121 | At1g63770 | Aminopeptidase | 99495/5.43 | 0 | – | ||
| 185, 186* | At1g13750 | Phosphoesterase | 68826/6.29 | 0 | + | † | |
| 42* | At4g24340 | Phosphorylase | 36024/8.24 | 1 | + | † | |
| 101 | At3g27950 | GDSL-motif lipase/hydrolase | 40120/8.89 | 0 | + | † | |
| 49 | At4g34870 | Peptidylprolyl isomerase | 18594/8.90 | 0 | – | † | † |
| 69 | At4g38740 | Peptidylprolyl isomerase | 18589/7.68 | 0 | – | † | |
| 57 | At1g30730 | FAD-linked oxidoreductase | 59161/9.23 | 0 | + | † | † |
| 56* | At5g44380 | FAD-linked oxidoreductase | 60920/9.51 | 1 | + | † | † |
| 17, 122, 123* | At3g55440 | Triosephosphate isomerase | 27380/5.39 | 0 | – | † | |
| 102 | At1g74000 | Strictosidine synthase | 34819/9.66 | 0 | + | ||
| 76 | At5g04590 | Sulfite reductase | 72391/8.51 | 0 | – | † | |
| 124 | At4g25100 | Iron superoxide dismutase | 23776/6.06 | 0 | – | ||
| 98, 125 | At5g43940 | Alcohol dehydrogenase | 41528/6.51 | 0 | – | † | |
| 89* | At3g48000 | Aldehyde dehydrogenase | 58951/7.11 | 0 | – | † | |
| 126 | At2g41530 | Esterase | 30759/5.91 | 0 | – | ||
| 10* | At2g31390 | Fructokinase | 35424/5.31 | 0 | – | † | |
| 48, 127 | At4g25900 | Aldose 1-epimerase | 36229/9.65 | 1 | + | † | † |
| 11, 21 | At5g07440 | Glutamate dehydrogenase | 45013/6.07 | 0 | – | † | |
| 41, 12 | At1g78380 | Glutathione transferase | 25691/5.80 | 0 | – | † | |
| 58, 105 | At1g65590 | β-Hexosaminidase | 60375/8.30 | 1 | + | † | |
| 22, 30, 60*, 103 | At1g13440 | Glyceraldehyde-3-phosphate dehydrogenase | 37004/6.67 | 0 | + | † | |
| 31 | At3g04120 | Glyceraldehyde-3-phosphate dehydrogenase | 36897/7.15 | 0 | + | ||
| 75 | At3g02360 | 6-Phosphogluconate dehydrogenase | 53829/7.02 | 0 | + | † | |
| 55, 74, 128* | At4g37870 | Phosphoenolpyruvate carboxykinase | 73929/6.61 | 0 | – | † | |
| 129 | At1g79550 | Phosphoglycerate kinase | 42162/5.49 | 0 | – | ||
| 130 | At1g80050 | Adenine phosphoribosyltransferase | 21241/5.69 | 0 | – | ||
| 1*, 28, 113, 131*, 132*, 133 | At5g17920 | Methionine synthase | 84646/6.09 | 0 | – | † | |
| 134 | At3g52880 | Monodehydroascorbate reductase | 46629/6.41 | 0 | – | † | |
| 135* | At5g03630 | Monodehydroascorbate reductase | 47408/5.18 | 0 | – | † | |
| 64*, 109 | At3g22110 | 20S proteasome α-subunit C | 27475/7.14 | 0 | – | † | |
| 66, 65 | At5g66140 | 20S proteasome α-subunit D2 | 27306/8.96 | 0 | – | † | |
| 20, 72* | At1g56450 | 20S proteasome β-subunit G1 | 27691/6.09 | 0 | – | † | |
| 67, 99, 136, 137 | At1g53070 | Legume lectin | 30416/8.49 | 1 | + | † | † |
| 110 | At3g16530 | Legume lectin | 30547/6.97 | 0 | + | † | |
| 85*, 138, 139* | At3g16420 | Jacalin lectin | 32138/5.46 | 0 | – | † | † |
| 26, 86, 140 | At3g16430 | Jacalin lectin | 32138/5.86 | 0 | – | † | |
| 87* | At3g16450 | Jacalin lectin | 32003/5.06 | 0 | – | † | † |
| 38 | At3g16460 | Jacalin lectin | 72430/5.31 | 0 | – | † | |
| 35*, 62*, 71, 80*, 112, 140*, 141*, 142, 143* | At1g78850 | Curculin-like lectin | 49818/7.82 | 0 | + | † | † |
| 29, 82 | At1g54000 | Myrosinase-associated protein | 43498/6.65 | 1 | + | † | |
| 144 | At1g54010 | Myrosinase-associated protein | 43572/8.33 | 1 | + | † | |
| 107 | At5g12940 | Leucine-rich repeat protein | 40497/9.52 | 1 | + | ||
| 61 | At1g18080 | WD-40 repeat protein | 36124/9.62 | 0 | – | † | |
| 53 | At1g07930 | Elongation factor 1-α | 49781/9.15 | 0 | – | † | |
| 68 | At3g53430 | 60S ribosomal protein L12 | 18073/9.05 | 0 | – | † | |
| 145 | At1g21350 | Unknown | 28413/9.14 | 0 | + | ||
| 70 | At4g09320 | Unknown | 15797/6.84 | 0 | – | ||
| 50 | At5g26260 | Unknown | 39802/9.21 | 0 | + | † | † |
| 108 | At5g26280 | Unknown | 39421/8.46 | 1 | + |
a Protein spot numbers correspond to the numbers on gel images (Fig. 2C, D). An asterisk indicates that the protein spot is phosphorylated, while underlined spots are glycosylated.
b Accession numbers are based on the NCBI database.
c The number of transmembrane domains (TMDs) as predicted by TMHMM is shown.
d The potential signal peptide as predicted by iPSORT, Signal P and Target P is represented by +.
e † in the 3 h and native columns indicates that proteins encoded by the same gene are also identified in the 3 h regenerated protoplasts and native cell walls, respectively.
Summary of the proteins identified as 1 M KCl extractable from the 1 h regenerated protoplasts
| Protein spot number a | Accession no. b | Protein name | Theoretical mol. wt/pI | TMD c | Signal peptide d | 3 h e | Native |
| 111 | At2g43610 | Chitinase | 30892/9.54 | 1 | + | ||
| 59 | At3g48580 | Xyloglucan endotransglucosylase/hydrolase | 32204/8.76 | 0 | + | † | † |
| 32 | At1g66280 | β-Glucosidase | 60142/6.74 | 0 | + | ||
| 2*, 3*, 104*, 114* 115*, 116* | At1g68560 | α-Xylosidase | 102210/6.32 | 0 | + | † | † |
| 52* | At5g20950 | β-Xylosidase | 67910/9.49 | 0 | + | † | † |
| 117 | At5g08380 | α-Galactosidase | 46079/6.19 | 0 | + | † | † |
| 95*, 96, 118* | At5g63810 | β-Galactosidase | 83619/8.58 | 1 | + | † | † |
| 51, 54*, 73, 100 | At3g13790 | β-Fructosidase | 66581/9.14 | 1 | + | † | |
| 119* | At3g26720 | α-Mannosidase | 115547/6.27 | 1 | + | † | † |
| 63 | At5g19440 | Cinnamyl-alcohol dehydrogenase | 35914/6.67 | 0 | – | † | |
| 106 | At2g04160 | Serine protease | 81364/9.34 | 0 | + | ||
| 120* | At5g10540 | Oligopeptidase | 79223/5.45 | 0 | – | ||
| 121 | At1g63770 | Aminopeptidase | 99495/5.43 | 0 | – | ||
| 185, 186* | At1g13750 | Phosphoesterase | 68826/6.29 | 0 | + | † | |
| 42* | At4g24340 | Phosphorylase | 36024/8.24 | 1 | + | † | |
| 101 | At3g27950 | GDSL-motif lipase/hydrolase | 40120/8.89 | 0 | + | † | |
| 49 | At4g34870 | Peptidylprolyl isomerase | 18594/8.90 | 0 | – | † | † |
| 69 | At4g38740 | Peptidylprolyl isomerase | 18589/7.68 | 0 | – | † | |
| 57 | At1g30730 | FAD-linked oxidoreductase | 59161/9.23 | 0 | + | † | † |
| 56* | At5g44380 | FAD-linked oxidoreductase | 60920/9.51 | 1 | + | † | † |
| 17, 122, 123* | At3g55440 | Triosephosphate isomerase | 27380/5.39 | 0 | – | † | |
| 102 | At1g74000 | Strictosidine synthase | 34819/9.66 | 0 | + | ||
| 76 | At5g04590 | Sulfite reductase | 72391/8.51 | 0 | – | † | |
| 124 | At4g25100 | Iron superoxide dismutase | 23776/6.06 | 0 | – | ||
| 98, 125 | At5g43940 | Alcohol dehydrogenase | 41528/6.51 | 0 | – | † | |
| 89* | At3g48000 | Aldehyde dehydrogenase | 58951/7.11 | 0 | – | † | |
| 126 | At2g41530 | Esterase | 30759/5.91 | 0 | – | ||
| 10* | At2g31390 | Fructokinase | 35424/5.31 | 0 | – | † | |
| 48, 127 | At4g25900 | Aldose 1-epimerase | 36229/9.65 | 1 | + | † | † |
| 11, 21 | At5g07440 | Glutamate dehydrogenase | 45013/6.07 | 0 | – | † | |
| 41, 12 | At1g78380 | Glutathione transferase | 25691/5.80 | 0 | – | † | |
| 58, 105 | At1g65590 | β-Hexosaminidase | 60375/8.30 | 1 | + | † | |
| 22, 30, 60*, 103 | At1g13440 | Glyceraldehyde-3-phosphate dehydrogenase | 37004/6.67 | 0 | + | † | |
| 31 | At3g04120 | Glyceraldehyde-3-phosphate dehydrogenase | 36897/7.15 | 0 | + | ||
| 75 | At3g02360 | 6-Phosphogluconate dehydrogenase | 53829/7.02 | 0 | + | † | |
| 55, 74, 128* | At4g37870 | Phosphoenolpyruvate carboxykinase | 73929/6.61 | 0 | – | † | |
| 129 | At1g79550 | Phosphoglycerate kinase | 42162/5.49 | 0 | – | ||
| 130 | At1g80050 | Adenine phosphoribosyltransferase | 21241/5.69 | 0 | – | ||
| 1*, 28, 113, 131*, 132*, 133 | At5g17920 | Methionine synthase | 84646/6.09 | 0 | – | † | |
| 134 | At3g52880 | Monodehydroascorbate reductase | 46629/6.41 | 0 | – | † | |
| 135* | At5g03630 | Monodehydroascorbate reductase | 47408/5.18 | 0 | – | † | |
| 64*, 109 | At3g22110 | 20S proteasome α-subunit C | 27475/7.14 | 0 | – | † | |
| 66, 65 | At5g66140 | 20S proteasome α-subunit D2 | 27306/8.96 | 0 | – | † | |
| 20, 72* | At1g56450 | 20S proteasome β-subunit G1 | 27691/6.09 | 0 | – | † | |
| 67, 99, 136, 137 | At1g53070 | Legume lectin | 30416/8.49 | 1 | + | † | † |
| 110 | At3g16530 | Legume lectin | 30547/6.97 | 0 | + | † | |
| 85*, 138, 139* | At3g16420 | Jacalin lectin | 32138/5.46 | 0 | – | † | † |
| 26, 86, 140 | At3g16430 | Jacalin lectin | 32138/5.86 | 0 | – | † | |
| 87* | At3g16450 | Jacalin lectin | 32003/5.06 | 0 | – | † | † |
| 38 | At3g16460 | Jacalin lectin | 72430/5.31 | 0 | – | † | |
| 35*, 62*, 71, 80*, 112, 140*, 141*, 142, 143* | At1g78850 | Curculin-like lectin | 49818/7.82 | 0 | + | † | † |
| 29, 82 | At1g54000 | Myrosinase-associated protein | 43498/6.65 | 1 | + | † | |
| 144 | At1g54010 | Myrosinase-associated protein | 43572/8.33 | 1 | + | † | |
| 107 | At5g12940 | Leucine-rich repeat protein | 40497/9.52 | 1 | + | ||
| 61 | At1g18080 | WD-40 repeat protein | 36124/9.62 | 0 | – | † | |
| 53 | At1g07930 | Elongation factor 1-α | 49781/9.15 | 0 | – | † | |
| 68 | At3g53430 | 60S ribosomal protein L12 | 18073/9.05 | 0 | – | † | |
| 145 | At1g21350 | Unknown | 28413/9.14 | 0 | + | ||
| 70 | At4g09320 | Unknown | 15797/6.84 | 0 | – | ||
| 50 | At5g26260 | Unknown | 39802/9.21 | 0 | + | † | † |
| 108 | At5g26280 | Unknown | 39421/8.46 | 1 | + |
| Protein spot number a | Accession no. b | Protein name | Theoretical mol. wt/pI | TMD c | Signal peptide d | 3 h e | Native |
| 111 | At2g43610 | Chitinase | 30892/9.54 | 1 | + | ||
| 59 | At3g48580 | Xyloglucan endotransglucosylase/hydrolase | 32204/8.76 | 0 | + | † | † |
| 32 | At1g66280 | β-Glucosidase | 60142/6.74 | 0 | + | ||
| 2*, 3*, 104*, 114* 115*, 116* | At1g68560 | α-Xylosidase | 102210/6.32 | 0 | + | † | † |
| 52* | At5g20950 | β-Xylosidase | 67910/9.49 | 0 | + | † | † |
| 117 | At5g08380 | α-Galactosidase | 46079/6.19 | 0 | + | † | † |
| 95*, 96, 118* | At5g63810 | β-Galactosidase | 83619/8.58 | 1 | + | † | † |
| 51, 54*, 73, 100 | At3g13790 | β-Fructosidase | 66581/9.14 | 1 | + | † | |
| 119* | At3g26720 | α-Mannosidase | 115547/6.27 | 1 | + | † | † |
| 63 | At5g19440 | Cinnamyl-alcohol dehydrogenase | 35914/6.67 | 0 | – | † | |
| 106 | At2g04160 | Serine protease | 81364/9.34 | 0 | + | ||
| 120* | At5g10540 | Oligopeptidase | 79223/5.45 | 0 | – | ||
| 121 | At1g63770 | Aminopeptidase | 99495/5.43 | 0 | – | ||
| 185, 186* | At1g13750 | Phosphoesterase | 68826/6.29 | 0 | + | † | |
| 42* | At4g24340 | Phosphorylase | 36024/8.24 | 1 | + | † | |
| 101 | At3g27950 | GDSL-motif lipase/hydrolase | 40120/8.89 | 0 | + | † | |
| 49 | At4g34870 | Peptidylprolyl isomerase | 18594/8.90 | 0 | – | † | † |
| 69 | At4g38740 | Peptidylprolyl isomerase | 18589/7.68 | 0 | – | † | |
| 57 | At1g30730 | FAD-linked oxidoreductase | 59161/9.23 | 0 | + | † | † |
| 56* | At5g44380 | FAD-linked oxidoreductase | 60920/9.51 | 1 | + | † | † |
| 17, 122, 123* | At3g55440 | Triosephosphate isomerase | 27380/5.39 | 0 | – | † | |
| 102 | At1g74000 | Strictosidine synthase | 34819/9.66 | 0 | + | ||
| 76 | At5g04590 | Sulfite reductase | 72391/8.51 | 0 | – | † | |
| 124 | At4g25100 | Iron superoxide dismutase | 23776/6.06 | 0 | – | ||
| 98, 125 | At5g43940 | Alcohol dehydrogenase | 41528/6.51 | 0 | – | † | |
| 89* | At3g48000 | Aldehyde dehydrogenase | 58951/7.11 | 0 | – | † | |
| 126 | At2g41530 | Esterase | 30759/5.91 | 0 | – | ||
| 10* | At2g31390 | Fructokinase | 35424/5.31 | 0 | – | † | |
| 48, 127 | At4g25900 | Aldose 1-epimerase | 36229/9.65 | 1 | + | † | † |
| 11, 21 | At5g07440 | Glutamate dehydrogenase | 45013/6.07 | 0 | – | † | |
| 41, 12 | At1g78380 | Glutathione transferase | 25691/5.80 | 0 | – | † | |
| 58, 105 | At1g65590 | β-Hexosaminidase | 60375/8.30 | 1 | + | † | |
| 22, 30, 60*, 103 | At1g13440 | Glyceraldehyde-3-phosphate dehydrogenase | 37004/6.67 | 0 | + | † | |
| 31 | At3g04120 | Glyceraldehyde-3-phosphate dehydrogenase | 36897/7.15 | 0 | + | ||
| 75 | At3g02360 | 6-Phosphogluconate dehydrogenase | 53829/7.02 | 0 | + | † | |
| 55, 74, 128* | At4g37870 | Phosphoenolpyruvate carboxykinase | 73929/6.61 | 0 | – | † | |
| 129 | At1g79550 | Phosphoglycerate kinase | 42162/5.49 | 0 | – | ||
| 130 | At1g80050 | Adenine phosphoribosyltransferase | 21241/5.69 | 0 | – | ||
| 1*, 28, 113, 131*, 132*, 133 | At5g17920 | Methionine synthase | 84646/6.09 | 0 | – | † | |
| 134 | At3g52880 | Monodehydroascorbate reductase | 46629/6.41 | 0 | – | † | |
| 135* | At5g03630 | Monodehydroascorbate reductase | 47408/5.18 | 0 | – | † | |
| 64*, 109 | At3g22110 | 20S proteasome α-subunit C | 27475/7.14 | 0 | – | † | |
| 66, 65 | At5g66140 | 20S proteasome α-subunit D2 | 27306/8.96 | 0 | – | † | |
| 20, 72* | At1g56450 | 20S proteasome β-subunit G1 | 27691/6.09 | 0 | – | † | |
| 67, 99, 136, 137 | At1g53070 | Legume lectin | 30416/8.49 | 1 | + | † | † |
| 110 | At3g16530 | Legume lectin | 30547/6.97 | 0 | + | † | |
| 85*, 138, 139* | At3g16420 | Jacalin lectin | 32138/5.46 | 0 | – | † | † |
| 26, 86, 140 | At3g16430 | Jacalin lectin | 32138/5.86 | 0 | – | † | |
| 87* | At3g16450 | Jacalin lectin | 32003/5.06 | 0 | – | † | † |
| 38 | At3g16460 | Jacalin lectin | 72430/5.31 | 0 | – | † | |
| 35*, 62*, 71, 80*, 112, 140*, 141*, 142, 143* | At1g78850 | Curculin-like lectin | 49818/7.82 | 0 | + | † | † |
| 29, 82 | At1g54000 | Myrosinase-associated protein | 43498/6.65 | 1 | + | † | |
| 144 | At1g54010 | Myrosinase-associated protein | 43572/8.33 | 1 | + | † | |
| 107 | At5g12940 | Leucine-rich repeat protein | 40497/9.52 | 1 | + | ||
| 61 | At1g18080 | WD-40 repeat protein | 36124/9.62 | 0 | – | † | |
| 53 | At1g07930 | Elongation factor 1-α | 49781/9.15 | 0 | – | † | |
| 68 | At3g53430 | 60S ribosomal protein L12 | 18073/9.05 | 0 | – | † | |
| 145 | At1g21350 | Unknown | 28413/9.14 | 0 | + | ||
| 70 | At4g09320 | Unknown | 15797/6.84 | 0 | – | ||
| 50 | At5g26260 | Unknown | 39802/9.21 | 0 | + | † | † |
| 108 | At5g26280 | Unknown | 39421/8.46 | 1 | + |
a Protein spot numbers correspond to the numbers on gel images (Fig. 2C, D). An asterisk indicates that the protein spot is phosphorylated, while underlined spots are glycosylated.
b Accession numbers are based on the NCBI database.
c The number of transmembrane domains (TMDs) as predicted by TMHMM is shown.
d The potential signal peptide as predicted by iPSORT, Signal P and Target P is represented by +.
e † in the 3 h and native columns indicates that proteins encoded by the same gene are also identified in the 3 h regenerated protoplasts and native cell walls, respectively.
Summary of the proteins identified as 1M KCl extractable from the 3h-regenerated protoplasts
| Protein spot number a | Accession no. b | Protein name | Theoretical mol. wt/pI | TMD c | Signal peptide d | 1 h e | Native |
| 59 | At3g48580 | Xyloglucan endotransglycosylase/hydrolase | 31922/8.78 | 0 | + | † | † |
| 32 | At1g66280 | β-Gucosidase | 60142/6.74 | 0 | + | † | |
| 82* | At3g09260 | β-Glucosidase | 60196/6.45 | 0 | + | ||
| 2*, 3*, 114*, 115* 116*, 187* | At1g68560 | α-Xylosidase | 102210/6.32 | 0 | + | † | † |
| 52* | At5g20950 | β-Xylosidase | 67910/9.49 | 0 | + | † | † |
| 117 | At5g08380 | α-Galactosidase | 46079/6.19 | 0 | + | † | † |
| 95*, 96 | At5g63810 | β-Galactosidase | 83619/8.58 | 1 | + | † | † |
| 13*, 51*, 54*, 73, 100 | At3g13790 | β-Fructosidase | 66581/9.14 | 1 | + | † | |
| 33* | At3g26720 | α-Mannosidase | 115203/6.7 | 1 | + | † | † |
| 63* | At5g19440 | Cinnamyl-alcohol dehydrogenase | 35576/7.15 | 0 | – | † | |
| 42* | At4g24340 | Phosphorylase | 36876/9.04 | 1 | + | † | |
| 8* | At4g23590 | Aminotransferase | 47919/5.76 | 0 | – | ||
| 88 | At3g29636 | Transferase | 22757/8.33 | 0 | – | ||
| 9 | At5g65020 | Annexin | 36358/5.76 | 0 | – | ||
| 81 | At2g05710 | Aconitate hydratase | 98720/5.79 | 0 | – | ||
| 83 | At3g14220 | GDSL-motif lipase/hydrolase | 40565/6.37 | 1 | + | ||
| 101 | At3g27950 | GDSL-motif lipase/hydrolase | 40120/8.89 | 0 | + | † | |
| 49 | At4g34870 | Peptidylprolyl isomerase | 18594/8.90 | 0 | – | † | † |
| 69 | At4g38740 | Peptidylprolyl isomerase | 18355/7.65 | 0 | – | † | |
| 57* | At1g30730 | FAD-linked oxidoreductase | 59161/9.23 | 0 | + | † | † |
| 56* | At5g44380 | FAD-linked oxidoreductase | 60920/9.51 | 1 | + | † | † |
| 36*, 94* | At5g18860 | Nucleoside hydrolase | 99580/7.59 | 0 | + | † | |
| 18 | At1g76680 | 12-Oxophytodienoate reductase | 41526/6.51 | 0 | – | ||
| 17* | At3g55440 | Triosephosphate isomerase | 27380/5.39 | 0 | – | † | |
| 102 | At1g74000 | Strictosidine synthase | 34819/9.66 | 0 | + | † | |
| 76 | At5g04590 | Sulfite reductase | 71933/8.47 | 0 | – | † | |
| 43 | At3g10920 | Manganese superoxide dismutase | 25426/8.83 | 0 | – | ||
| 98, 125 | At5g43940 | Alcohol dehydrogenase | 41528/6.51 | 0 | – | † | |
| 89* | At3g48000 | Aldehyde dehydrogenase | 58951/7.11 | 0 | – | † | |
| 10* | At2g31390 | Fructokinase | 35424/5.31 | 0 | – | † | |
| 48*, 93 | At4g25900 | Aldose 1-epimerase | 36234/10.24 | 1 | + | † | † |
| 11, 21 | At5g07440 | Glutamate dehydrogenase | 45013/6.07 | 0 | – | † | |
| 12, 41 | At1g78380 | Glutathione transferase | 25691/5.80 | 0 | – | † | |
| 78* | At1g04410 | Malate dehydrogenase | 35890/6.11 | 0 | + | ||
| 79* | At1g53240 | Malate dehydrogenase | 36010/8.54 | 0 | – | ||
| 58 | At1g65590 | β-Hexosaminidase | 59996/8.26 | 1 | + | † | |
| 22*, 23*, 30, 31, 60*, 103 | At1g13440 | Glyceraldehyde-3-phosphate dehydrogenase | 37004/6.67 | 0 | + | † | |
| 75 | At3g02360 | 6-Phosphogluconate dehydrogenase | 53560/7.49 | 0 | + | † | |
| 27*, 55*, 74* | At4g37870 | Phosphoenolpyruvate carboxykinase | 73929/6.61 | 0 | – | † | |
| 1*, 28, 131*, 132*, 133 | At5g17920 | Methionine synthase | 84646/6.09 | 0 | – | † | |
| 134 | At3g52880 | Monodehydroascorbate reductase | 46629/6.41 | 0 | – | † | |
| 90 | At3g53880 | Monodehydroascorbate reductase | 46629/6.41 | 0 | – | ||
| 25* | At5g03630 | Monodehydroascorbate reductase | 47507/5.24 | 0 | + | † | |
| 34 | At1g16470 | 20S proteasome α-subunit B | 25685/5.53 | 0 | – | ||
| 64* | At3g22110 | 20S proteasome α-subunit C | 27572/6.60 | 0 | – | † | |
| 65*, 66* | At5g66140 | 20S proteasome α-subunit D2 | 27306/8.96 | 0 | – | ||
| 7 | At4g31300 | 20S proteasome β-subunit A | 25193/5.31 | 0 | – | ||
| 19, 44 | At4g14800 | 20S proteasome β-subunit D2 | 22084/6.21 | 0 | + | † | |
| 91 | At3g60820 | 20S proteasome β-subunit F1 | 24856/6.95 | 0 | – | ||
| 20*, 72* | At1g56450 | 20S proteasome β-subunit G1 | 27691/6.09 | 0 | – | † | |
| 77 | At1g03230 | Glycoprotein | 46803/9.33 | 0 | + | † | |
| 67*, 99* | At1g53070 | Lgume lectin | 30361/8.84 | 1 | + | † | † |
| 84 | At3g15356 | LEgume lectin | 29788/8.91 | 0 | + | ||
| 24 | At2g39310 | Jacalin lectin | 50602/5.15 | 0 | – | ||
| 45 | At3g16420 | Jacalin lectin | 32138/5.46 | 0 | – | ||
| 16, 40, 85*, 97 | At3g16420 | Jacalin lectin | 32138/5.46 | 0 | – | † | † |
| 26, 86 | At3g16430 | Jacalin lectin | 32213/5.86 | 0 | – | † | |
| 46*, 47, 87 | At3g16450 | Jacalin lectin | 32003/5.06 | 0 | – | † | † |
| 4*, 5*, 6, 14*, 15*, 37*, 38*, 39* | At3g16460 | Jacalin lectin | 72430/5.31 | 0 | – | † | |
| 35*, 80*, 62*, 71, 142, 141* | At1g78850 | Curculin-like lectin | 49818/7.82 | 0 | + | † | † |
| 29 | At1g54000 | Myrosinase-associated protein | 43498/6.65 | 1 | + | † | |
| 144 | At1g54010 | Myrosinase-associated protein | 43572/8.33 | 1 | + | † | |
| 61 | At1g18080 | WD-40 repeat protein | 35730/7.81 | 0 | – | † | |
| 53 | At1g07930 | Elongation factor 1-α | 49485/9.6 | 0 | – | † | |
| 68 | At3g53430 | 60S ribosomal protein L12 | 17952/9.57 | 0 | – | † | |
| 70, 92 | At4g09320 | Unknown | 16500/6.79 | 0 | – | ||
| 50 | At5g26260 | Unknown | 39802/9.21 | 0 | + | † | † |
| Protein spot number a | Accession no. b | Protein name | Theoretical mol. wt/pI | TMD c | Signal peptide d | 1 h e | Native |
| 59 | At3g48580 | Xyloglucan endotransglycosylase/hydrolase | 31922/8.78 | 0 | + | † | † |
| 32 | At1g66280 | β-Gucosidase | 60142/6.74 | 0 | + | † | |
| 82* | At3g09260 | β-Glucosidase | 60196/6.45 | 0 | + | ||
| 2*, 3*, 114*, 115* 116*, 187* | At1g68560 | α-Xylosidase | 102210/6.32 | 0 | + | † | † |
| 52* | At5g20950 | β-Xylosidase | 67910/9.49 | 0 | + | † | † |
| 117 | At5g08380 | α-Galactosidase | 46079/6.19 | 0 | + | † | † |
| 95*, 96 | At5g63810 | β-Galactosidase | 83619/8.58 | 1 | + | † | † |
| 13*, 51*, 54*, 73, 100 | At3g13790 | β-Fructosidase | 66581/9.14 | 1 | + | † | |
| 33* | At3g26720 | α-Mannosidase | 115203/6.7 | 1 | + | † | † |
| 63* | At5g19440 | Cinnamyl-alcohol dehydrogenase | 35576/7.15 | 0 | – | † | |
| 42* | At4g24340 | Phosphorylase | 36876/9.04 | 1 | + | † | |
| 8* | At4g23590 | Aminotransferase | 47919/5.76 | 0 | – | ||
| 88 | At3g29636 | Transferase | 22757/8.33 | 0 | – | ||
| 9 | At5g65020 | Annexin | 36358/5.76 | 0 | – | ||
| 81 | At2g05710 | Aconitate hydratase | 98720/5.79 | 0 | – | ||
| 83 | At3g14220 | GDSL-motif lipase/hydrolase | 40565/6.37 | 1 | + | ||
| 101 | At3g27950 | GDSL-motif lipase/hydrolase | 40120/8.89 | 0 | + | † | |
| 49 | At4g34870 | Peptidylprolyl isomerase | 18594/8.90 | 0 | – | † | † |
| 69 | At4g38740 | Peptidylprolyl isomerase | 18355/7.65 | 0 | – | † | |
| 57* | At1g30730 | FAD-linked oxidoreductase | 59161/9.23 | 0 | + | † | † |
| 56* | At5g44380 | FAD-linked oxidoreductase | 60920/9.51 | 1 | + | † | † |
| 36*, 94* | At5g18860 | Nucleoside hydrolase | 99580/7.59 | 0 | + | † | |
| 18 | At1g76680 | 12-Oxophytodienoate reductase | 41526/6.51 | 0 | – | ||
| 17* | At3g55440 | Triosephosphate isomerase | 27380/5.39 | 0 | – | † | |
| 102 | At1g74000 | Strictosidine synthase | 34819/9.66 | 0 | + | † | |
| 76 | At5g04590 | Sulfite reductase | 71933/8.47 | 0 | – | † | |
| 43 | At3g10920 | Manganese superoxide dismutase | 25426/8.83 | 0 | – | ||
| 98, 125 | At5g43940 | Alcohol dehydrogenase | 41528/6.51 | 0 | – | † | |
| 89* | At3g48000 | Aldehyde dehydrogenase | 58951/7.11 | 0 | – | † | |
| 10* | At2g31390 | Fructokinase | 35424/5.31 | 0 | – | † | |
| 48*, 93 | At4g25900 | Aldose 1-epimerase | 36234/10.24 | 1 | + | † | † |
| 11, 21 | At5g07440 | Glutamate dehydrogenase | 45013/6.07 | 0 | – | † | |
| 12, 41 | At1g78380 | Glutathione transferase | 25691/5.80 | 0 | – | † | |
| 78* | At1g04410 | Malate dehydrogenase | 35890/6.11 | 0 | + | ||
| 79* | At1g53240 | Malate dehydrogenase | 36010/8.54 | 0 | – | ||
| 58 | At1g65590 | β-Hexosaminidase | 59996/8.26 | 1 | + | † | |
| 22*, 23*, 30, 31, 60*, 103 | At1g13440 | Glyceraldehyde-3-phosphate dehydrogenase | 37004/6.67 | 0 | + | † | |
| 75 | At3g02360 | 6-Phosphogluconate dehydrogenase | 53560/7.49 | 0 | + | † | |
| 27*, 55*, 74* | At4g37870 | Phosphoenolpyruvate carboxykinase | 73929/6.61 | 0 | – | † | |
| 1*, 28, 131*, 132*, 133 | At5g17920 | Methionine synthase | 84646/6.09 | 0 | – | † | |
| 134 | At3g52880 | Monodehydroascorbate reductase | 46629/6.41 | 0 | – | † | |
| 90 | At3g53880 | Monodehydroascorbate reductase | 46629/6.41 | 0 | – | ||
| 25* | At5g03630 | Monodehydroascorbate reductase | 47507/5.24 | 0 | + | † | |
| 34 | At1g16470 | 20S proteasome α-subunit B | 25685/5.53 | 0 | – | ||
| 64* | At3g22110 | 20S proteasome α-subunit C | 27572/6.60 | 0 | – | † | |
| 65*, 66* | At5g66140 | 20S proteasome α-subunit D2 | 27306/8.96 | 0 | – | ||
| 7 | At4g31300 | 20S proteasome β-subunit A | 25193/5.31 | 0 | – | ||
| 19, 44 | At4g14800 | 20S proteasome β-subunit D2 | 22084/6.21 | 0 | + | † | |
| 91 | At3g60820 | 20S proteasome β-subunit F1 | 24856/6.95 | 0 | – | ||
| 20*, 72* | At1g56450 | 20S proteasome β-subunit G1 | 27691/6.09 | 0 | – | † | |
| 77 | At1g03230 | Glycoprotein | 46803/9.33 | 0 | + | † | |
| 67*, 99* | At1g53070 | Lgume lectin | 30361/8.84 | 1 | + | † | † |
| 84 | At3g15356 | LEgume lectin | 29788/8.91 | 0 | + | ||
| 24 | At2g39310 | Jacalin lectin | 50602/5.15 | 0 | – | ||
| 45 | At3g16420 | Jacalin lectin | 32138/5.46 | 0 | – | ||
| 16, 40, 85*, 97 | At3g16420 | Jacalin lectin | 32138/5.46 | 0 | – | † | † |
| 26, 86 | At3g16430 | Jacalin lectin | 32213/5.86 | 0 | – | † | |
| 46*, 47, 87 | At3g16450 | Jacalin lectin | 32003/5.06 | 0 | – | † | † |
| 4*, 5*, 6, 14*, 15*, 37*, 38*, 39* | At3g16460 | Jacalin lectin | 72430/5.31 | 0 | – | † | |
| 35*, 80*, 62*, 71, 142, 141* | At1g78850 | Curculin-like lectin | 49818/7.82 | 0 | + | † | † |
| 29 | At1g54000 | Myrosinase-associated protein | 43498/6.65 | 1 | + | † | |
| 144 | At1g54010 | Myrosinase-associated protein | 43572/8.33 | 1 | + | † | |
| 61 | At1g18080 | WD-40 repeat protein | 35730/7.81 | 0 | – | † | |
| 53 | At1g07930 | Elongation factor 1-α | 49485/9.6 | 0 | – | † | |
| 68 | At3g53430 | 60S ribosomal protein L12 | 17952/9.57 | 0 | – | † | |
| 70, 92 | At4g09320 | Unknown | 16500/6.79 | 0 | – | ||
| 50 | At5g26260 | Unknown | 39802/9.21 | 0 | + | † | † |
a Protein spot numbers correspond to the numbers on gel images (Fig. 2E, F). An asterisk indicates that the protein spot is phosphorylated, while underlined spots are glycosylated.
b Accession numbers are based on the NCBI database.
c The number of transmembrane domains (TMDs) as predicted by TMHMM is shown.
d The potential signal peptide as predicted by iPSORT, Signal P and Target P is represented by +.
e † in the 1 h and Native columns indicates that proteins encoded by the same gene are also identified in the 1 h regenerated protoplasts and native cell walls, respectively.
Summary of the proteins identified as 1M KCl extractable from the 3h-regenerated protoplasts
| Protein spot number a | Accession no. b | Protein name | Theoretical mol. wt/pI | TMD c | Signal peptide d | 1 h e | Native |
| 59 | At3g48580 | Xyloglucan endotransglycosylase/hydrolase | 31922/8.78 | 0 | + | † | † |
| 32 | At1g66280 | β-Gucosidase | 60142/6.74 | 0 | + | † | |
| 82* | At3g09260 | β-Glucosidase | 60196/6.45 | 0 | + | ||
| 2*, 3*, 114*, 115* 116*, 187* | At1g68560 | α-Xylosidase | 102210/6.32 | 0 | + | † | † |
| 52* | At5g20950 | β-Xylosidase | 67910/9.49 | 0 | + | † | † |
| 117 | At5g08380 | α-Galactosidase | 46079/6.19 | 0 | + | † | † |
| 95*, 96 | At5g63810 | β-Galactosidase | 83619/8.58 | 1 | + | † | † |
| 13*, 51*, 54*, 73, 100 | At3g13790 | β-Fructosidase | 66581/9.14 | 1 | + | † | |
| 33* | At3g26720 | α-Mannosidase | 115203/6.7 | 1 | + | † | † |
| 63* | At5g19440 | Cinnamyl-alcohol dehydrogenase | 35576/7.15 | 0 | – | † | |
| 42* | At4g24340 | Phosphorylase | 36876/9.04 | 1 | + | † | |
| 8* | At4g23590 | Aminotransferase | 47919/5.76 | 0 | – | ||
| 88 | At3g29636 | Transferase | 22757/8.33 | 0 | – | ||
| 9 | At5g65020 | Annexin | 36358/5.76 | 0 | – | ||
| 81 | At2g05710 | Aconitate hydratase | 98720/5.79 | 0 | – | ||
| 83 | At3g14220 | GDSL-motif lipase/hydrolase | 40565/6.37 | 1 | + | ||
| 101 | At3g27950 | GDSL-motif lipase/hydrolase | 40120/8.89 | 0 | + | † | |
| 49 | At4g34870 | Peptidylprolyl isomerase | 18594/8.90 | 0 | – | † | † |
| 69 | At4g38740 | Peptidylprolyl isomerase | 18355/7.65 | 0 | – | † | |
| 57* | At1g30730 | FAD-linked oxidoreductase | 59161/9.23 | 0 | + | † | † |
| 56* | At5g44380 | FAD-linked oxidoreductase | 60920/9.51 | 1 | + | † | † |
| 36*, 94* | At5g18860 | Nucleoside hydrolase | 99580/7.59 | 0 | + | † | |
| 18 | At1g76680 | 12-Oxophytodienoate reductase | 41526/6.51 | 0 | – | ||
| 17* | At3g55440 | Triosephosphate isomerase | 27380/5.39 | 0 | – | † | |
| 102 | At1g74000 | Strictosidine synthase | 34819/9.66 | 0 | + | † | |
| 76 | At5g04590 | Sulfite reductase | 71933/8.47 | 0 | – | † | |
| 43 | At3g10920 | Manganese superoxide dismutase | 25426/8.83 | 0 | – | ||
| 98, 125 | At5g43940 | Alcohol dehydrogenase | 41528/6.51 | 0 | – | † | |
| 89* | At3g48000 | Aldehyde dehydrogenase | 58951/7.11 | 0 | – | † | |
| 10* | At2g31390 | Fructokinase | 35424/5.31 | 0 | – | † | |
| 48*, 93 | At4g25900 | Aldose 1-epimerase | 36234/10.24 | 1 | + | † | † |
| 11, 21 | At5g07440 | Glutamate dehydrogenase | 45013/6.07 | 0 | – | † | |
| 12, 41 | At1g78380 | Glutathione transferase | 25691/5.80 | 0 | – | † | |
| 78* | At1g04410 | Malate dehydrogenase | 35890/6.11 | 0 | + | ||
| 79* | At1g53240 | Malate dehydrogenase | 36010/8.54 | 0 | – | ||
| 58 | At1g65590 | β-Hexosaminidase | 59996/8.26 | 1 | + | † | |
| 22*, 23*, 30, 31, 60*, 103 | At1g13440 | Glyceraldehyde-3-phosphate dehydrogenase | 37004/6.67 | 0 | + | † | |
| 75 | At3g02360 | 6-Phosphogluconate dehydrogenase | 53560/7.49 | 0 | + | † | |
| 27*, 55*, 74* | At4g37870 | Phosphoenolpyruvate carboxykinase | 73929/6.61 | 0 | – | † | |
| 1*, 28, 131*, 132*, 133 | At5g17920 | Methionine synthase | 84646/6.09 | 0 | – | † | |
| 134 | At3g52880 | Monodehydroascorbate reductase | 46629/6.41 | 0 | – | † | |
| 90 | At3g53880 | Monodehydroascorbate reductase | 46629/6.41 | 0 | – | ||
| 25* | At5g03630 | Monodehydroascorbate reductase | 47507/5.24 | 0 | + | † | |
| 34 | At1g16470 | 20S proteasome α-subunit B | 25685/5.53 | 0 | – | ||
| 64* | At3g22110 | 20S proteasome α-subunit C | 27572/6.60 | 0 | – | † | |
| 65*, 66* | At5g66140 | 20S proteasome α-subunit D2 | 27306/8.96 | 0 | – | ||
| 7 | At4g31300 | 20S proteasome β-subunit A | 25193/5.31 | 0 | – | ||
| 19, 44 | At4g14800 | 20S proteasome β-subunit D2 | 22084/6.21 | 0 | + | † | |
| 91 | At3g60820 | 20S proteasome β-subunit F1 | 24856/6.95 | 0 | – | ||
| 20*, 72* | At1g56450 | 20S proteasome β-subunit G1 | 27691/6.09 | 0 | – | † | |
| 77 | At1g03230 | Glycoprotein | 46803/9.33 | 0 | + | † | |
| 67*, 99* | At1g53070 | Lgume lectin | 30361/8.84 | 1 | + | † | † |
| 84 | At3g15356 | LEgume lectin | 29788/8.91 | 0 | + | ||
| 24 | At2g39310 | Jacalin lectin | 50602/5.15 | 0 | – | ||
| 45 | At3g16420 | Jacalin lectin | 32138/5.46 | 0 | – | ||
| 16, 40, 85*, 97 | At3g16420 | Jacalin lectin | 32138/5.46 | 0 | – | † | † |
| 26, 86 | At3g16430 | Jacalin lectin | 32213/5.86 | 0 | – | † | |
| 46*, 47, 87 | At3g16450 | Jacalin lectin | 32003/5.06 | 0 | – | † | † |
| 4*, 5*, 6, 14*, 15*, 37*, 38*, 39* | At3g16460 | Jacalin lectin | 72430/5.31 | 0 | – | † | |
| 35*, 80*, 62*, 71, 142, 141* | At1g78850 | Curculin-like lectin | 49818/7.82 | 0 | + | † | † |
| 29 | At1g54000 | Myrosinase-associated protein | 43498/6.65 | 1 | + | † | |
| 144 | At1g54010 | Myrosinase-associated protein | 43572/8.33 | 1 | + | † | |
| 61 | At1g18080 | WD-40 repeat protein | 35730/7.81 | 0 | – | † | |
| 53 | At1g07930 | Elongation factor 1-α | 49485/9.6 | 0 | – | † | |
| 68 | At3g53430 | 60S ribosomal protein L12 | 17952/9.57 | 0 | – | † | |
| 70, 92 | At4g09320 | Unknown | 16500/6.79 | 0 | – | ||
| 50 | At5g26260 | Unknown | 39802/9.21 | 0 | + | † | † |
| Protein spot number a | Accession no. b | Protein name | Theoretical mol. wt/pI | TMD c | Signal peptide d | 1 h e | Native |
| 59 | At3g48580 | Xyloglucan endotransglycosylase/hydrolase | 31922/8.78 | 0 | + | † | † |
| 32 | At1g66280 | β-Gucosidase | 60142/6.74 | 0 | + | † | |
| 82* | At3g09260 | β-Glucosidase | 60196/6.45 | 0 | + | ||
| 2*, 3*, 114*, 115* 116*, 187* | At1g68560 | α-Xylosidase | 102210/6.32 | 0 | + | † | † |
| 52* | At5g20950 | β-Xylosidase | 67910/9.49 | 0 | + | † | † |
| 117 | At5g08380 | α-Galactosidase | 46079/6.19 | 0 | + | † | † |
| 95*, 96 | At5g63810 | β-Galactosidase | 83619/8.58 | 1 | + | † | † |
| 13*, 51*, 54*, 73, 100 | At3g13790 | β-Fructosidase | 66581/9.14 | 1 | + | † | |
| 33* | At3g26720 | α-Mannosidase | 115203/6.7 | 1 | + | † | † |
| 63* | At5g19440 | Cinnamyl-alcohol dehydrogenase | 35576/7.15 | 0 | – | † | |
| 42* | At4g24340 | Phosphorylase | 36876/9.04 | 1 | + | † | |
| 8* | At4g23590 | Aminotransferase | 47919/5.76 | 0 | – | ||
| 88 | At3g29636 | Transferase | 22757/8.33 | 0 | – | ||
| 9 | At5g65020 | Annexin | 36358/5.76 | 0 | – | ||
| 81 | At2g05710 | Aconitate hydratase | 98720/5.79 | 0 | – | ||
| 83 | At3g14220 | GDSL-motif lipase/hydrolase | 40565/6.37 | 1 | + | ||
| 101 | At3g27950 | GDSL-motif lipase/hydrolase | 40120/8.89 | 0 | + | † | |
| 49 | At4g34870 | Peptidylprolyl isomerase | 18594/8.90 | 0 | – | † | † |
| 69 | At4g38740 | Peptidylprolyl isomerase | 18355/7.65 | 0 | – | † | |
| 57* | At1g30730 | FAD-linked oxidoreductase | 59161/9.23 | 0 | + | † | † |
| 56* | At5g44380 | FAD-linked oxidoreductase | 60920/9.51 | 1 | + | † | † |
| 36*, 94* | At5g18860 | Nucleoside hydrolase | 99580/7.59 | 0 | + | † | |
| 18 | At1g76680 | 12-Oxophytodienoate reductase | 41526/6.51 | 0 | – | ||
| 17* | At3g55440 | Triosephosphate isomerase | 27380/5.39 | 0 | – | † | |
| 102 | At1g74000 | Strictosidine synthase | 34819/9.66 | 0 | + | † | |
| 76 | At5g04590 | Sulfite reductase | 71933/8.47 | 0 | – | † | |
| 43 | At3g10920 | Manganese superoxide dismutase | 25426/8.83 | 0 | – | ||
| 98, 125 | At5g43940 | Alcohol dehydrogenase | 41528/6.51 | 0 | – | † | |
| 89* | At3g48000 | Aldehyde dehydrogenase | 58951/7.11 | 0 | – | † | |
| 10* | At2g31390 | Fructokinase | 35424/5.31 | 0 | – | † | |
| 48*, 93 | At4g25900 | Aldose 1-epimerase | 36234/10.24 | 1 | + | † | † |
| 11, 21 | At5g07440 | Glutamate dehydrogenase | 45013/6.07 | 0 | – | † | |
| 12, 41 | At1g78380 | Glutathione transferase | 25691/5.80 | 0 | – | † | |
| 78* | At1g04410 | Malate dehydrogenase | 35890/6.11 | 0 | + | ||
| 79* | At1g53240 | Malate dehydrogenase | 36010/8.54 | 0 | – | ||
| 58 | At1g65590 | β-Hexosaminidase | 59996/8.26 | 1 | + | † | |
| 22*, 23*, 30, 31, 60*, 103 | At1g13440 | Glyceraldehyde-3-phosphate dehydrogenase | 37004/6.67 | 0 | + | † | |
| 75 | At3g02360 | 6-Phosphogluconate dehydrogenase | 53560/7.49 | 0 | + | † | |
| 27*, 55*, 74* | At4g37870 | Phosphoenolpyruvate carboxykinase | 73929/6.61 | 0 | – | † | |
| 1*, 28, 131*, 132*, 133 | At5g17920 | Methionine synthase | 84646/6.09 | 0 | – | † | |
| 134 | At3g52880 | Monodehydroascorbate reductase | 46629/6.41 | 0 | – | † | |
| 90 | At3g53880 | Monodehydroascorbate reductase | 46629/6.41 | 0 | – | ||
| 25* | At5g03630 | Monodehydroascorbate reductase | 47507/5.24 | 0 | + | † | |
| 34 | At1g16470 | 20S proteasome α-subunit B | 25685/5.53 | 0 | – | ||
| 64* | At3g22110 | 20S proteasome α-subunit C | 27572/6.60 | 0 | – | † | |
| 65*, 66* | At5g66140 | 20S proteasome α-subunit D2 | 27306/8.96 | 0 | – | ||
| 7 | At4g31300 | 20S proteasome β-subunit A | 25193/5.31 | 0 | – | ||
| 19, 44 | At4g14800 | 20S proteasome β-subunit D2 | 22084/6.21 | 0 | + | † | |
| 91 | At3g60820 | 20S proteasome β-subunit F1 | 24856/6.95 | 0 | – | ||
| 20*, 72* | At1g56450 | 20S proteasome β-subunit G1 | 27691/6.09 | 0 | – | † | |
| 77 | At1g03230 | Glycoprotein | 46803/9.33 | 0 | + | † | |
| 67*, 99* | At1g53070 | Lgume lectin | 30361/8.84 | 1 | + | † | † |
| 84 | At3g15356 | LEgume lectin | 29788/8.91 | 0 | + | ||
| 24 | At2g39310 | Jacalin lectin | 50602/5.15 | 0 | – | ||
| 45 | At3g16420 | Jacalin lectin | 32138/5.46 | 0 | – | ||
| 16, 40, 85*, 97 | At3g16420 | Jacalin lectin | 32138/5.46 | 0 | – | † | † |
| 26, 86 | At3g16430 | Jacalin lectin | 32213/5.86 | 0 | – | † | |
| 46*, 47, 87 | At3g16450 | Jacalin lectin | 32003/5.06 | 0 | – | † | † |
| 4*, 5*, 6, 14*, 15*, 37*, 38*, 39* | At3g16460 | Jacalin lectin | 72430/5.31 | 0 | – | † | |
| 35*, 80*, 62*, 71, 142, 141* | At1g78850 | Curculin-like lectin | 49818/7.82 | 0 | + | † | † |
| 29 | At1g54000 | Myrosinase-associated protein | 43498/6.65 | 1 | + | † | |
| 144 | At1g54010 | Myrosinase-associated protein | 43572/8.33 | 1 | + | † | |
| 61 | At1g18080 | WD-40 repeat protein | 35730/7.81 | 0 | – | † | |
| 53 | At1g07930 | Elongation factor 1-α | 49485/9.6 | 0 | – | † | |
| 68 | At3g53430 | 60S ribosomal protein L12 | 17952/9.57 | 0 | – | † | |
| 70, 92 | At4g09320 | Unknown | 16500/6.79 | 0 | – | ||
| 50 | At5g26260 | Unknown | 39802/9.21 | 0 | + | † | † |
a Protein spot numbers correspond to the numbers on gel images (Fig. 2E, F). An asterisk indicates that the protein spot is phosphorylated, while underlined spots are glycosylated.
b Accession numbers are based on the NCBI database.
c The number of transmembrane domains (TMDs) as predicted by TMHMM is shown.
d The potential signal peptide as predicted by iPSORT, Signal P and Target P is represented by +.
e † in the 1 h and Native columns indicates that proteins encoded by the same gene are also identified in the 1 h regenerated protoplasts and native cell walls, respectively.
Abbreviations
- CBB
Coomassie brilliant blue
- 2-D PAGE
two-dimensional polyacrylamide gel electrophoresis
- DTT
dithiothreitol
- FDA
fluorescein diacetate
- G6PDH
glucose-6-phosphate dehydrogenase
- IEF
isoelectric focusing
- MALDI-TOF/MS
matrix-assisted laser desorption ionization-time-of-flight/mass spectrometry
- SEM
scanning electron microscope
- xyloglucan endotransglucosylase/hydrolase
XTH.
References
Amano, T., Hirasawa, K., O’Donohue, M.J., Pernolle, J.C. and Shioi, Y. (
Arabidopsis Genome Initiative (
Arioli, T., Peng, L., Betzner, A.S., Burn, J., Wittke, W., et al. (
Baluska, F., Samaj, J., Wojtaszek, P., Volkmann, D. and Menzel, D. (
Bonetta, D.T., Facette, M., Raab, T.K. and Somerville, C.R. (
Borderies, G., Jamet, E., Lafitte, C., Rossignol, M., Jauneau, A., Boudart, G., Monsarrat, B., Esquerre-Tugaye, M.T., Boudet, A. and Pont-Lezica, R. (
Bradford, M.M. (
Brummell, D.A. and Harpster, M.H. (
Brummell, D.A. and Labavitch, J.M. (
Campbell, P. and Braam, J. (
Carpita, N.C. and McCann, M.C. (
Cassab, G.I. (
Chaffin, W.L., Lopez-Ribot, J.L., Casanova, M., Gozalbo, D. and Martinez, J.P. (
Chivasa, S., Ndimba, B.K., Simon, W.J., Robertson, D., Yu, X.L., Knox, J.P., Bolwell, P. and Slabas, A.R. (
Cosgrove, D.J. (
Darley, C.P., Forrester, A.M. and McQueen-Mason, S.J. (
Faik, A., Price, N.J., Raikhel, N.V. and Keegstra, K. (
Gamborg, O.L., Murashige, T., Thorpe, T.A. and Vasil, I.K. (
Girke, T., Lauricha, J., Tran, H., Keegstra, K. and Raikhel, N. (
Gozalbo, D., Gil-Navarro, I., Azorin, I., Renau-Piqueras, J., Martinez, J.P. and Gil, M.L. (
Hart, C., Schulenberg, B., Steinberg, T.H., Leung, W.Y. and Patton, W.F. (
Hong, Z.L., Zhang, Z.M., Olson, J.M. and Verma, D.P.S. (
Ishikawa, M., Robertson, A.J. and Gusta, L.V. (
Ito, H. and Nishitani, K. (
Iwai, H., Masaoka, N., Ishii, T. and Satoh, S. (
Karkonen, A., Koutaniemi, S., Mustonen, M., Syrjanen, K., Brounow, G., Kilpelainen, I., Teeri, T.H. and Simola, L.K. (
Kende, H., Bradford, K., Brummell, D., Cho, H.-T., Cosgrove, D., et al. (
Konno, H., Yamasaki, Y. and Katoh, K. (
Konno, H., Yamasaki, Y. and Katoh, K. (
Konomi, M., Ishiguro, J. and Osumi, M. (
Kuster, B., Mortensen, P., Andersen, J.S. and Mann, M. (
Laemmli, U.K. (
Li, J., Burton, R.A., Harvey, A.J., Hrmova, M., Wardak, A.Z., Stone, B.A. and Fincher, G.B. (
Li, X., Cordero, I., Caplan, J., Molhoj, M. and Reiter, W.D. (
Li, Z.C., McClure, J.W. and Hagerman, A.E. (
Lorences, E.P. and Fry, S.C. (
Manfield, I.W., Orfila, C., McCartney, L., Harholt, J., Bernal, A.J., Scheller, H.V., Gilmartin, P.M., Mikkelsen, J.D., Knox, J.P. and Willats, W.G.T. (
Martin, D.B. and Nelson, P.S. (
Mathur, J., Szabados, L., Schaefer, S., Grunenberg, B., Lossow, A., Jonas-Straube, E., Schell, J., Koncz, C. and Koncz-Kalman, Z. (
Matsushima, R., Kondo, M., Nishimura, M. and Hara-Nishimura, I. (
McQueen-Mason, S. and Cosgrove, D.J. (
Murashige, T. and Skoog, F. (
Nielsen, H., Engelbrecht, J., Brunak, S. and von Heijne, G. (
Nishitani, K. (
Nishitani, K. (
Nishitani, K. and Tominaga, R. (
Nishitani, K. and Tominaga, R. (
Okamoto-Nakazato, A., Takahashi, K., Kido, N., Owaribe, K. and Katou, K. (
Okazawa, K., Sato, Y., Nakagawa, T., Asada, I., Tomita, E. and Nishitani, K. (
Pardo, M., Ward, M., Bains, S., Molina, M., Blackstock, W., Gil, C. and Nombela, C. (
Perrin, R.M., DeRocher, A.E., Bar-Peled, M., Zeng, W., Norambuena, L., Orellana, A., Raikhel, N.V. and Keegstra, K. (
Qin, Q., Bergmann, C.W., Rose, J.K.C., Saladie, M., Kolli, V.S.K., Albersheim, P., Darvill, A.G. and York, W.S. (
Rabilloud, T., Valette, C. and Lawrence, J.J. (
Robertson, D., Mitchell, G.P., Gilroy, J.S., Gerrish, C., Bolwell, G.P. and Slabas, A.R. (
Rose, J.K.C., Bashi, S., Giovannoni, J.J., Jahn, M.M. and Saravanan, R.S. (
Rose, J.K.C., Braam, J., Fry, S.C. and Nishitani, K. (
Rose, J.K.C., Lee, S.J., Lee, J.M., Kim, B.D., Catala, C., Saravanan, R.S., Yamane, H. and Gee, C.T. (
Sampedro, J., Sieiro, C., Revilla, G., Gonzalez-Villa, T. and Zarra, I. (
Schirawski, J., Planchais, S. and Haenni, A.L. (
Schulenberg, B., Aggeler, R., Beechem, J.M., Capaldi, R.A. and Patton, W.F. (
Smith, M.M. and McCully, M.E. (
Sonnhammer, E.L., von Heijine, G. and Krogh, A. (
Souza, I.R.P. and MacAdam, J.W. (
Vanacker, H., Carver, T.L.W. and Foyer, C.H. (
Weimar, M.and Rothe, G.M. (
Whitney, S.E.C., Gidley, M.J. and McQueen-Mason, S.J. (
Yokoyama, R. and Nishitani, K. (
Yokoyama, R. and Nishitani, K. (