Abstract
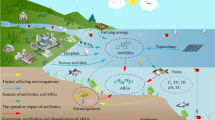
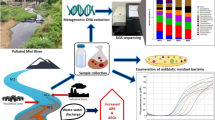
Antibiotic residues in marine sediments of fish farms negatively influence microbial ecologic systems. The microbial degradation of antibiotic residues was experimentally examined in the marine sediments of Uranouchi Bay, to which one of five antibiotics was added. After incubation reducing physical factors, ampicillin, doxycycline, oxytetracycline, and thiamphenicol were significantly degraded, while josamycin maintained most of the initial amounts. The isolates resistant to ampicillin, josamycin, oxytetracycline, or thiamphenicol degraded each antibiotic in wide ranges of degrees, whereas the isolates degrading doxycycline were not obtained. Microbial degradation may contribute to the disappearance of ampicillin, doxycycline, oxytetracycline, and thiamphenicol in the fish farm. In contrast, the disappearance of josamycin would depend on physical factors, but the bacteria degrading josamycin at least exist in the marine sediments. Phylogenetic analysis using 16S rDNA sequences demonstrated that the antibiotic-resistant isolates formed several clusters in the Gram-positive bacterial group, the Flavobacterium-Cytophaga-Bacteroides group, and the proteobacteria subdivisions. The antibiotic-resistant bacterial population would be composed of various species including ubiquitous coastal bacterial groups. Several species of antibiotic resistant bacteria show antibiotic degradation activities, and appear to contribute to the disappearance of antibiotics in Uranouchi Bay.
Similar content being viewed by others
References
Alderman D, Michel C. Chemotherapy in aquaculture today. In: Michel C, Alderman DJ (eds). Chemotherapy in Aquaculture. From Theory to Reality. Office International des Epizootics, Paris. 1992; 3–24.
Jacobsen P, Berglind L. Persistence of oxytetracycline in sediments from fish farms. Aquaculture 1988; 70: 365–370.
Samuelsen OB. Degradation of oxytetracycline in seawater at two different temperatures and light intensities, and the persistence of oxytetracycline in the sediments from a fish farm. Aquaculture 1989; 83: 7–16.
Björklund H, Bondesteam J, Bylund G. Residues of oxytetracycline in wild fish and sediments from fish farm. Aquaculture 1990; 86: 359–367.
Samuelsen OB, Lunestad BT, Ervik A, Fjelde S. Stability of antibacterial agents in an artificial marine aquaculture sediment studied under laboratory conditions. Aquaculture 1994; 126: 283–290.
Rogstad B, Hormazabal V, Ellingsen OF, Rasmussen KE. Pharmacokinetic study of oxytetracycline in fish. I. Absorption. distribution and accumulation in rainbow trout in freshwater. Aquaculture 1991; 96: 219–226.
Lalumera GM, Calamari D, Galli P, Castiglioni S, Crosa G, Fanelli R. Preliminary investigation on the environmental occurrence and effects of antibiotics used in aquaculture in Italy. Chemosphere 2004; 54: 661–668.
Brown JH, Higuera-Ciapara I. Antibiotic residues in farmed shrimp-a developing problem? In: Michel C, Alderman DJ (eds). Chemotherapy in Aquaculture. From Theory to Reality. Office International des Epizootics, Paris, 1992; 394–403.
Aoki T. Present and future problems concerning the development of resistance in aquaculture. In: Michel C, Alderman DJ (eds). Chemotherapy in Aquaculture. From Theory to Reality. Office International des Epizootics. Paris. 1992; 254–262.
Schmidt AS, Bruun MS, Dalsgaard I, Pedersen K, Larsen JL. Occurrence of antimicrobial resistance in fish-pathogenic and environmental bacteria associated with four Danish rainbow trout farms. Appl. Environ. Microbiol. 2000; 66: 4908–4915.
Arvanitidou M, Katsouyannopoulos V, Tsakris A. Antibiotic resistance patterns of enterococci isolated from coastal bathing waters. J. Med. Microbiol. 2001; 50: 1001–1005.
Tendencia EA, de la Pena LD. Antibiotic resistance of bacteria from shrimp ponds. Aquaculture 2001; 195: 193–204.
Miranda CD, Zemelman R. Antimicrobial multiresistance, in bacteria isolated from freshwater Chilean salmon farms. Sci. Total Environ. 2002; 293: 207–218.
Miranda CD, Zemelman R. Bacterial resistance to oxytetracycline in Chilean salmon farming. Aquaculture 2002; 212: 31–47.
Chelossi E, Vezzulli L, Milano A, Branzoni M, Fabiano M, Riccardi G, Banat IM. Antibiotic resistance of benthic bacteria in fish-farm and control sediments of the Western Mediterranean. Aquaculture 2003; 219: 83–97.
Rhodes G, Huys G, Swings J, Mcgann P, Hiney M, Smith P, Pickup RW. Distribution of oxytetracycline resistance plasmids between aeromonads in hospital and aquaculture environments: implication of Tn1721 in dissemination of the tetracycline resistance determinant tet A. Appl. Environ. Microbiol. 2000; 66: 3883–3890.
Schmidt AS, Bruun MS, Dalsgaard I, Larsen JL. Incidence, distribution, and spread of tetracycline resistance determinants and integron-associated antibiotic resistance genes among motile aeromonads from a fish farming environment. Appl. Environ. Microbiol. 2001; 67: 5675–5682.
Martinez JL, Baquero F. Interactions among strategies associated with bacterial infection: pathogenicity, epidemicity, and antibiotic resistance. Clin. Microbiol. Rev. 2002; 15: 647–679.
Sørum H, L’Abee-Lund TM, Solberg A, Wold A. Integron-containing IncU R plasmids pRASI and pAr-32 from the fish pathogen Aeromonas salmonicida. Antimicrob. Agents Chemother. 2003; 47: 1285–1290.
Lunestad BT. Fate and effects of antibacterial agents in aquatic environments. In: Michel C, Alderman DJ (eds). Chemotherapy in Aquaculture. From Theory to Reality. Office International des Epizootics, Paris. 1992; 152–161.
Stock I, Heisig P, Wiedemann B. Expression of betalactamases in Yersinia enterocolitica strains of biovars 2, 4 and 5. J. Med. Microbiol. 1999; 48: 1023–1027.
Tzelepi E, Arvanitidou M, Mavroidi A, Tsakris A. Antibiotic susceptibilities of Yersinia enterocolitica and Y. intermedia isolates from aquatic environments. J. Med. Microbiol. 1999; 48: 157–160.
Nitzan Y, Deutsch EB, Pechatnikov I. Diffusion of betalactam antibiotics through oligomeric or monomeric porin channels of some gram-negative bacteria. Curr. Microbiol. 2002; 45: 446–455.
Petersen A, Dalsgaard A. Antimicrobial resistance of intestinal Aeromonas spp. and Enterococcus spp. in fish cultured in integrated broiler-fish farms in Thailand. Aquaculture 2003; 219: 71–82.
Munekage Y, Naing KM. Uranouchiwan ni okeru suisannyou kouseibussitsu no bunnpu to zannryuusei ni kannsuru kennkyuu. Kaigan Gakkai Ronnbunn 2005; 52: 926–930 (in Japanese).
Knox JH, Jurand J. Mechanism of reversed-phase separation of tetracyclines by high-performance liquid chromatography. J. Chromatogr. 1979; 186: 776–782.
Oka H, Matsumoto K, Harada K, Kadowaki S, Suzuki M. Improvement of chemical analysis of antibiotics, VIII. Application of prepacked C18 cartridge for the analysis of tetracycline residues in animal liver. Chromatography 1985; 325: 265–274.
Alderman DJ, Smith P. Development of draft protocols of standard reference methods for antimicrobial agent susceptibility testing of bacteria associated with fish diseases. Aquaculture 2001; 196: 211–243.
Russell WC, Newman C, Williamson DH. A simple cytochemical technique for demonstration of DNA in cells infected with mycoplasms and viruses. Nature 1974; 253: 461–462.
Porter KG, Feig YS. The use of DAPI for identifying and counting aquatic microflora. Limnol. Oceanogr. 1980; 25: 943–948.
Maidak BL, Olsen GJ, Larsen N, Overbeek R, Mccaughey MJ, Woese C. The RDP (Ribosomal Database Project). Nucleic Acids Res. 1997; 25: 109–111.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J. Mol. Biol. 1990; 215: 403–410.
Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987; 4: 406–425.
Thompson JD, Higgins DG, Gibson TJ. CLUSTALW: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 1994; 22: 4673–4680.
Samuelsen OB, Solheim E, Lunestad BT. Fate and microbiological effects of furazolidone in a marine aquaculture sediment. Sci. Total Environ. 1991; 108: 275–283.
Björklund H, Rabergh CMI, Bylund G. Residues of oxolinic acid and oxytetracycline in fish and sediments from fish farms. Aquaculture 1991; 97: 85–96.
Hancock RE. The bacterial outer membrane as a drug barrier. Trends Microbiol. 1997; 5: 37–42.
Hawkey PM. The origins and molecular basis of antibiotic resistance. Br. Med. J. 1998; 317: 657–660.
Alonso A, Sanchez P, Martinez JL. Environment selection of antibiotic resistance genes. Environ. Microbiol 2001; 3: 1–9.
Chandrasekaran K. Industrial enzymes from marine microorganisms: the Indian scenario. J. Mar. Biotechnol. 1997; 5: 86–89.
Cheung TK, Ho PL, Woo PC, Yuen KY, Chau PY. Cloning and expression of class A beta-lactamase gene blaA (BPS) in Burkholderia pseudomallei. Antimicrob. Agents Chemother. 2002; 46: 1132–1135.
Fang H, Edlund C, Hedberg M, Nord CE. New findings in beta-lactam and metronidazole resistant Bacteroides fragilis group. Int. J. Antimicrob. Agents 2002; 19: 361–370.
Egidius E, Wiik K, Andersen K, Hoff KA, Hjeltnes B. Vibrio salmonicida spp. nov., a new fish pathogen. Int. J. Syst. Bacteriol. 1986; 36: 518–520.
Depaola A, Ulaszek J, Kaysner CA, Tenge BJ, Nordstrom JL, Wells J, Puhr N, Gendel SM. Molecular, serological and virulence characteristics of Vibrio parahaemolyticus isolated from environmental, food, and clinical sources in North America and asia. Appl. Environ. Microbiol. 2003; 69: 3999–4005.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Maki, T., Hasegawa, H., Kitami, H. et al. Bacterial degradation of antibiotic residues in marine fish farm sediments of Uranouchi Bay and phylogenetic analysis of antibiotic-degrading bacteria using 16S rDNA sequences. Fish Sci 72, 811–820 (2006). https://doi.org/10.1111/j.1444-2906.2006.01222.x
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1111/j.1444-2906.2006.01222.x